Abstract
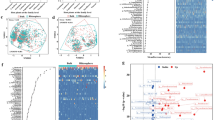
Climate change causes an unprecedented increase in glacial retreats. The melting ice exposes land for colonization and diversification of bacterial communities leading to soil development, changes in plant community composition, and ecosystem functioning. Although a few studies have focused on macro-level deglaciation impacts, little is known about such effects on the bacterial community succession. Here, we provide meta-barcoding-based insight into the ecological attributes of bacterial community across different retreating periods of the Gangotri glacier, western Himalaya. We selected three sites along a terminal moraine representing recent (~ 20 yrs), intermediate (~ 100 yrs), and late (~ 300 yrs) deglaciation periods. Results showed that the genus Mycobacterium belonging to phylum Actinobacteria dominated recently deglaciated land. Relative abundance of these pioneer bacterial taxa decreased by 20–50% in the later stages with the emergence of new and rising of the less abundant members of the phyla Proteobacteria, Firmicutes, Planctomycetes, Acidobacteria, Verrucomicrobia, Candidatus TM6, and Chloroflexi. The community in the recent stage was less rich and harbored competitive interactions, while the later stages experienced a surge in bacterial diversity with cooperative interactions. The shift in α-diversity and composition was strongly influenced by soil organic carbon, carbon to nitrogen ratio, and soil moisture content. The functional analyses revealed a progression from a metabolism focused to a functionally progressive community required for bacterial co-existence and succession in plant communities. Overall, the findings indicate that the bacterial communities inhabit, diversify, and develop specialized functions post-deglaciation leading to nutrient inputs to soil and vegetation development, which may provide feedback to climate change.





Similar content being viewed by others
Data Availability
The DNA sequence dataset has been deposited to the National Centre for Biotechnology Information (NCBI) Short Read Archive (https://www.ncbi.nlm.nih.gov/sra). The BioProject accession number is PRJNA754406.
Code Availability
Not applicable.
References
Cramer W, Bondeau A, Woodward FI et al (2001) Global response of terrestrial ecosystem structure and function to CO 2 and climate change: results from six dynamic global vegetation models: ecosystem dynamics, CO 2 and climate change. Glob Change Biol 7:357–373. https://doi.org/10.1046/j.1365-2486.2001.00383.x
Venkatachalam S, Kannan VM, Saritha VN et al (2021) Bacterial diversity and community structure along the glacier foreland of Midtre Lovénbreen, Svalbard. Arct Ecol Indic 126:107704. https://doi.org/10.1016/j.ecolind.2021.107704
Wu X, Zhang W, Liu G et al (2012) Bacterial diversity in the foreland of the Tianshan No. 1 glacier China. Environ Res Lett 7:014038. https://doi.org/10.1088/1748-9326/7/1/014038
Hotaling S, Hood E, Hamilton TL (2017) Microbial ecology of mountain glacier ecosystems: biodiversity, ecological connections and implications of a warming climate. Environ Microbiol 19:2935–2948. https://doi.org/10.1111/1462-2920.13766
Alfaro FD, Salazar-Burrows A, Bañales-Seguel C et al (2020) Soil microbial abundance and activity across forefield glacier chronosequence in the northern Patagonian Ice Field, Chile. Arct Antarct Alp Res 52:553–562. https://doi.org/10.1080/15230430.2020.1820124
Frkova Z, Pistocchi C, Vystavna Y, et al (2021) Phosphorus dynamics during early soil development in extreme environment. Soils and biogeochemical cycling. Preprint.
Bradley JA, Anesio AM, Arndt S (2017) Microbial and biogeochemical dynamics in glacier forefields are sensitive to century-scale climate and anthropogenic change. Front Earth Sci. https://doi.org/10.3389/feart.2017.00026
Gilbert JA, Dupont CL (2011) Microbial metagenomics: beyond the genome. Annu Rev Mar Sci 3:347–371. https://doi.org/10.1146/annurev-marine-120709-142811
Feng W, Zhang Y, Yan R et al (2020) Dominant soil bacteria and their ecological attributes across the deserts in northern China. Eur J Soil Sci 71:524–535. https://doi.org/10.1111/ejss.12866
Ortiz-Estrada ÁM, Gollas-Galván T, Martínez-Córdova LR, Martínez-Porchas M (2019) Predictive functional profiles using metagenomic 16S rRNA data: a novel approach to understanding the microbial ecology of aquaculture systems. Rev Aquac 11:234–245. https://doi.org/10.1111/raq.12237
Langille MGI, Zaneveld J, Caporaso JG et al (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31:814–821. https://doi.org/10.1038/nbt.2676
Sun S, Jones RB, Fodor AA (2020) Inference-based accuracy of metagenome prediction tools varies across sample types and functional categories. Microbiome 8:46. https://doi.org/10.1186/s40168-020-00815-y
Shrestha UB, Gautam S, Bawa KS (2012) Widespread climate change in the Himalayas and associated changes in local ecosystems. PLoS ONE 7:e36741. https://doi.org/10.1371/journal.pone.0036741
Singh RK (2018) Impact of Climate Change on the Retreat of Himalayan Glaciers and Its Impact on Major River Hydrology: Himalayan Glacier Hydrology. In Climate Change and Environmental Concerns: Breakthroughs in Research and Practice, IGI Global, pp. 681–694
Pusalkar PK, Singh DK (2012) Flora of Gangotri national park, western Himalaya. Botanical Survey of India, India
Tiwari P, Bhattacharya P, Rawat GS, Talukdar G (2021) Equilibrium in soil respiration across a climosequence indicates its resilience to climate change in a glaciated valley, western Himalaya. Sci Rep. https://doi.org/10.1038/s41598-021-02199-x
Sanyal AK, Uniyal VP, Chandra K, Bhardwaj M (2013) Diversity, distribution pattern and seasonal variation in moth assemblages along altitudinal gradient in Gangotri landscape area, Western Himalaya, Uttarakhand, India. J Threat Taxa 5:3646–3653. https://doi.org/10.11609/JoTT.o2597.3646-53
Tiwari P, Bhattacharya P, Rawat GS et al (2021) Experimental warming increases ecosystem respiration by increasing above-ground respiration in alpine meadows of Western Himalaya. Sci Rep. https://doi.org/10.1038/s41598-021-82065-y
Lanzén A, Epelde L, Blanco F et al (2016) Multi-targeted metagenetic analysis of the influence of climate and environmental parameters on soil microbial communities along an elevational gradient. Sci Rep. https://doi.org/10.1038/srep28257
Rayment GE, Lyons DJ (2011) Soil chemical methods: Australasia. CSIRO Publishing, Collingwood, Vic
Blakemore LC, Searle PL, Daly BK (1987) Methods for chemical analysis of soils. NZ Soil Bureau, Dept. of Scientific and Industrial Research, Lower Hutt, N.Z.
Walkley A, Black IA (1934) An examination of the Degtjareff method for determining soil organic matter, and a proposed modification of the chromic acid titration method. Soil Sci 37:29–38. https://doi.org/10.1097/00010694-193401000-00003
Bremner JM (2018) Nitrogen-Total. In: Sparks DL, Page AL, Helmke PA et al (eds) SSSA book series. Soil Science Society of America, American Society of Agronomy, Madison, WI, USA, pp 1085–1121
Luo Z, Liu J, Zhao P et al (2019) Biogeographic patterns and assembly mechanisms of bacterial communities differ between habitat generalists and specialists across elevational gradients. Front Microbiol. https://doi.org/10.3389/fmicb.2019.00169
Schloss PD, Westcott SL, Ryabin T et al (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. https://doi.org/10.1128/AEM.01541-09
Edgar RC, Haas BJ, Clemente JC et al (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Pedrós-Alió C (2006) Marine microbial diversity: can it be determined? Trends Microbiol 14:257–263. https://doi.org/10.1016/j.tim.2006.04.007
Deng Y, Jiang Y-H, Yang Y et al (2012) Molecular ecological network analyses. BMC Bioinformatics 13:113. https://doi.org/10.1186/1471-2105-13-113
Shannon P, Markiel A, Ozier O et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Spellerberg IF, Fedor PJ (2003) A tribute to Claude Shannon (1916–2001) and a plea for more rigorous use of species richness, species diversity and the ‘Shannon-Wiener’ Index: on species richness and diversity. Glob Ecol Biogeogr 12:177–179. https://doi.org/10.1046/j.1466-822X.2003.00015.x
Oksanen J, Guillaume Blanchet F, Friendly M Kindt R, Legendre P, McGlinn D, Minchin PR, O'Hara RB, Simpson GL, Solymos P, Henry M, Stevens M, Szoecs E, Wagner H (2020) Vegan: community ecology package. R package version 2.5-7. https://CRAN.R-project.org/package=vegan.
Johnson JB, Omland KS (2004) Model selection in ecology and evolution. Trends Ecol Evol 19:101–108. https://doi.org/10.1016/j.tree.2003.10.013
Li J, Ma Y-B, Hu H-W et al (2015) Field-based evidence for consistent responses of bacterial communities to copper contamination in two contrasting agricultural soils. Front Microbiol. https://doi.org/10.3389/fmicb.2015.00031
Grace JB (2007) Structural equation modeling and natural systems. Biometrics 63:977–977. https://doi.org/10.1111/j.1541-0420.2007.00856_13.x
R Core Team (2020) R: A language and environment for statistical computing. R Found Stat Comput, Vienna, Austria
Ciccazzo S, Esposito A, Borruso L, Brusetti L (2016) Microbial communities and primary succession in high altitude mountain environments. Ann Microbiol 66:43–60. https://doi.org/10.1007/s13213-015-1130-1
Ji M, Greening C, Vanwonterghem I et al (2017) Atmospheric trace gases support primary production in Antarctic desert surface soil. Nature 552:400–403. https://doi.org/10.1038/nature25014
Jangid K, Whitman WB, Condron LM et al (2013) Soil bacterial community succession during long-term ecosystem development. Mol Ecol 22:3415–3424. https://doi.org/10.1111/mec.12325
Bhattacharya P, Tiwari P, Rai ID et al (2022) Edaphic factors override temperature in shaping soil bacterial diversity across an elevation-vegetation gradient in Himalaya. Appl Soil Ecol 170:104306. https://doi.org/10.1016/j.apsoil.2021.104306
Brady NC, Weil RR (2002) The nature and properties of soils, 13th edn. Prentice Hall, New Jersey, p 249
Qiang W, He L, Zhang Y et al (2021) Aboveground vegetation and soil physicochemical properties jointly drive the shift of soil microbial community during subalpine secondary succession in southwest China. CATENA 202:105251. https://doi.org/10.1016/j.catena.2021.105251
Zhu B, Li C, Wang J et al (2020) Elevation rather than season determines the assembly and co-occurrence patterns of soil bacterial communities in forest ecosystems of Mount Gongga. Appl Microbiol Biotechnol 104:7589–7602. https://doi.org/10.1007/s00253-020-10783-w
Santos R, de Carvalho CCCR, Stevenson A et al (2015) Extraordinary solute-stress tolerance contributes to the environmental tenacity of mycobacteria: extraordinary stress-tolerance of mycobacteria. Environ Microbiol Rep 7:746–764. https://doi.org/10.1111/1758-2229.12306
Miteva VI, Sheridan PP, Brenchley JE (2004) Phylogenetic and physiological diversity of microorganisms isolated from a deep greenland glacier ice core. Appl Environ Microbiol 70:202–213. https://doi.org/10.1128/AEM.70.1.202-213.2004
Johnson SS, Hebsgaard MB, Christensen TR et al (2007) Ancient bacteria show evidence of DNA repair. Proc Natl Acad Sci 104:14401–14405. https://doi.org/10.1073/pnas.0706787104
Gupta P, Sangwan N, Lal R, Vakhlu J (2015) Bacterial diversity of drass, cold desert in Western Himalaya, and its comparison with Antarctic and Arctic. Arch Microbiol 197:851–860. https://doi.org/10.1007/s00203-015-1121-4
Shao K, Bai C, Cai J et al (2019) Illumina sequencing revealed soil microbial communities in a Chinese alpine grassland. Geomicrobiol J 36:204–211. https://doi.org/10.1080/01490451.2018.1534902
Edwards A, Pachebat JA, Swain M et al (2013) A metagenomic snapshot of taxonomic and functional diversity in an alpine glacier cryoconite ecosystem. Environ Res Lett 8:035003. https://doi.org/10.1088/1748-9326/8/3/035003
Ahmad T, Gupta G, Sharma A et al (2021) Metagenomic analysis exploring taxonomic and functional diversity of bacterial communities of a Himalayan urban fresh water lake. PloS One 16:e0248116
Dube JP, Valverde A, Steyn JM et al (2019) Differences in bacterial diversity, composition and function due to long-term agriculture in soils in the eastern free state of South Africa. Diversity 11:61. https://doi.org/10.3390/d11040061
Kazemi S, Hatam I, Lanoil B (2016) Bacterial community succession in a high-altitude subarctic glacier foreland is a three-stage process. Mol Ecol 25:5557–5567. https://doi.org/10.1111/mec.13835
Koch AL (2001) Oligotrophs versus copiotrophs. BioEssays 23:657–661
Ortiz-Álvarez R, FiererRíos NA et al (2018) Consistent changes in the taxonomic structure and functional attributes of bacterial communities during primary succession. ISME J 12:1658–1667. https://doi.org/10.1038/s41396-018-0076-2
Drake HL, Gößner AS, Daniel SL (2008) Old acetogens, new light. Ann N Y Acad Sci 1125:100–128. https://doi.org/10.1196/annals.1419.016
Kuypers MMM, Marchant HK, Kartal B (2018) The microbial nitrogen-cycling network. Nat Rev Microbiol 16:263–276. https://doi.org/10.1038/nrmicro.2018.9
Wang J, Wu Y, Li J et al (2021) Energetic supply regulates heterotrophic nitrogen fixation along a glacial chronosequence. Soil Biol Biochem 154:108150. https://doi.org/10.1016/j.soilbio.2021.108150
Tiwari P, Singh JS (2017) A plant growth promoting rhizospheric Pseudomonas aeruginosa strain inhibits seed germination in Triticum aestivum (L) and Zea mays (L). Microbiol Res. https://doi.org/10.4081/mr.2017.7233
Florentino AP, Weijma J, Stams AJM, Sánchez-Andrea I (2016) Ecophysiology and application of acidophilic sulfur-reducing microorganisms. In: Rampelotto PH (ed) Biotechnology of extremophiles. Springer International Publishing, Cham, pp 141–175
Acknowledgements
Forest Department of Uttarakhand provided necessary permits to research in Gangotri National Park. We acknowledge help from Dr. Devendra Kumar and Umed S Rana (field assistance), Arun Kumar (laboratory work), Dr. Punyasloke Bhadury (DNA sequencing), Dr. Awadhesh Pandit and Tejali Naik (NGS work), Sitendu Goswami, and Dr. Raman Kumar (Data analysis). We thank Dr. Samrat Mondol, Dr. Sathyakumar (Nodal Scientist, NMSHE), Dr. Dhananjay Mohan (Director), Dr. Y.V. Jhala (Dean), Dr. Bitapi Sinha (Research Coordinator), and Nodal Officer of Wildlife Forensics and Conservation Genetics Cell of Wildlife Institute of India for facilitating this work.
Funding
This research is part of the project National Mission for Sustaining the Himalayan Ecosystem (NMSHE) funded by the Department of Science and Technology, Government of India (Grant No. DST/SPLICE/CCP/NMSHE/TF-2/WII/2014[G]). Partial funding for the bacterial Next Generation Sequencing was provided by the United Nations Development Programme and the Ministry of Environment, Forest and Climate Change Government of India through the Third National Communication project (Grant No. 7/2/2015-CC). Pamela Bhattacharya was supported by the Council of Scientific and Industrial Research, Government of India (Award no. 09/668(0012)/2019– EMR–I).
Author information
Authors and Affiliations
Contributions
Conceptualization: PT and PB; Funding acquisition: GSR, GT, and PB; Resources: GSR and GT; Supervision: GSR and GT; Methodology: PT and PB; Data curation: PB; Formal analysis and investigation: PB and PT; Writing—original draft preparation: PB and PT; Writing—review and editing: GSR, GT, PB, and PT.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical Approval
Not applicable.
Consent to Participate
All authors gave consent to participate and publish all information provided in the manuscript.
Consent to Publish
All authors gave consent to participate and publish all information provided in the manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bhattacharya, P., Tiwari, P., Talukdar, G. et al. Shifts in Bacterial Community Composition and Functional Traits at Different Time Periods Post-deglaciation of Gangotri Glacier, Himalaya. Curr Microbiol 79, 91 (2022). https://doi.org/10.1007/s00284-022-02779-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-022-02779-8