Abstract
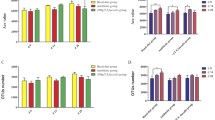
In-feed antibiotics can influence intestinal microbial structures in born and early-life within a period. However, the impact of antibiotics on gut microbiota during long-term antibiotic-free and antibiotic breeding at porcine-fattening phase have not been studied extensively so far. Here, we conducted a systematic 16S rRNA gene sequencing-based study combined with metagenomic analysis to reveal the variation of diversity and function of gut microbiota between antibiotic-free (treatment group, TG) and antibiotic (a mixture of flavomycin and enramycin, control group, CG) breeding at various stages of fattening pigs. In the present study, Bacteroidetes, Firmicutes, and Proteobacteria phyla were the core microbiomes in fattening pig gut microbiota. The ratio between Firmicutes and Bacteroidetes significantly increased with age (P = 0.03). TG showed significantly higher relative abundance of Proteobacteria and Fibrobacteres phyla than CG. The microbial community can be divided into several notably clustered blocks based on cooperative and competitive correlations. These blocks centered on numerous special genera, which play essential roles in body development and disease prevention. TG showed obviously higher proportions of metabolic pathways related to metabolism, endocrine system, nervous system and excretory system, but pathways included carbohydrate metabolism and immune system diseases in CG. Collectively, this study has comprehensively demonstrated microbial diversities, differences and correlations among gut microbiota, microbial metabolism and gene functions during long-term antibiotic-free breeding. This work provides a novel resource and information with positive implications for pig husbandry production and disease prevention.





Similar content being viewed by others
Data Availability
The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive (GSA; https://bigd.big.ac.cn/gsa) in BIG data Center, Beijing Institute of Genomics, Chinese Academy of Sciences, under accession number CRA002245.
References
Laxminarayan R, Duse A, Wattal C, Zaidi AK, Wertheim HF, Sumpradit N, Vlieghe E, Hara GL, Gould IM, Goossens H, Greko C, So AD, Bigdeli M, Tomson G, Woodhouse W, Ombaka E, Peralta AQ, Qamar FN, Mir F, Kariuki S, Bhutta ZA, Coates A, Bergstrom R, Wright GD, Brown ED, Cars O (2013) Antibiotic resistance-the need for global solutions. Lancet Infect Dis 13(12):1057–1098. https://doi.org/10.1016/S1473-3099(13)70318-9
Ventola CL (2015) The antibiotic resistance crisis: part 1: causes and threats. Pharm Ther 40(4):277–283
Prestinaci F, Pezzotti P, Pantosti A (2015) Antimicrobial resistance: a global multifaceted phenomenon. Pathog Glob Health 109(7):309–318. https://doi.org/10.1179/2047773215Y.0000000030
Kimman TG, Smits MA, Kemp B, Wever P, Verheijden J (2010) Banning antibiotics, reducing resistance, preventing and fighting infections: white paper on research enabling an 'antibiotic-free' animal husbandry. Wageningen UR etc., Wageningen [etc.]
Lange K, Buerger M, Stallmach A, Bruns T (2016) Effects of antibiotics on gut microbiota. Dig Dis 34(3):260–268. https://doi.org/10.1159/000443360
Sanchez B, Delgado S, Blanco-Miguez A, Lourenco A, Gueimonde M, Margolles A (2017) Probiotics, gut microbiota, and their influence on host health and disease. Mol Nutr Food Res. https://doi.org/10.1002/mnfr.201600240
Isaacson R, Kim HB (2012) The intestinal microbiome of the pig. Anim Health Res Rev 13(1):100–109. https://doi.org/10.1017/S1466252312000084
Liu Y, Zheng Z, Yu L, Wu S, Sun L, Wu S, Xu Q, Cai S, Qin N, Bao W (2019) Examination of the temporal and spatial dynamics of the gut microbiome in newborn piglets reveals distinct microbial communities in six intestinal segments. Sci Rep 9(1):3453. https://doi.org/10.1038/s41598-019-40235-z
Zhao W, Wang Y, Liu S, Huang J, Zhai Z, He C, Ding J, Wang J, Wang H, Fan W, Zhao J, Meng H (2015) The dynamic distribution of porcine microbiota across different ages and gastrointestinal tract segments. PLoS ONE 10(2):e0117441. https://doi.org/10.1371/journal.pone.0117441
Looft T, Allen HK, Cantarel BL, Levine UY, Bayles DO, Alt DP, Henrissat B, Stanton TB (2014) Bacteria, phages and pigs: the effects of in-feed antibiotics on the microbiome at different gut locations. ISME J 8(8):1566–1576. https://doi.org/10.1038/ismej.2014.12
Li N, Huang S, Jiang L, Dai Z, Li T, Han D, Wang J (2019) Characterization of the early life microbiota development and predominant Lactobacillus Species at distinct gut segments of low- and normal-birth-weight piglets. Front Microbiol 10:797. https://doi.org/10.3389/fmicb.2019.00797
Li N, Huang S, Jiang L, Wang W, Li T, Zuo B, Li Z, Wang J (2018) Differences in the gut microbiota establishment and metabolome characteristics between low- and normal-birth-weight piglets during early-Life. Front Microbiol 9:1798. https://doi.org/10.3389/fmicb.2018.01798
Mu C, Yang Y, Su Y, Zoetendal EG, Zhu W (2017) Differences in microbiota membership along the gastrointestinal tract of piglets and their differential alterations following an early-life antibiotic intervention. Front Microbiol 8:797. https://doi.org/10.3389/fmicb.2017.00797
Li H, Liang T, Chu Q, Xu F, Li Y, Fu L, Zhou B (2017) Effects of several in-feed antibiotic combinations on the abundance and diversity of fecal microbes in weaned pigs. Can J Microbiol 63(5):402–410. https://doi.org/10.1139/cjm-2016-0681
Schokker D, Zhang J, Zhang LL, Vastenhouw SA, Heilig HG, Smidt H, Rebel JM, Smits MA (2014) Early-life environmental variation affects intestinal microbiota and immune development in new-born piglets. PLoS ONE 9(6):e100040. https://doi.org/10.1371/journal.pone.0100040
Nair S, Farzan A, Weese JS, Poljak Z, Friendship RM (2020) Effect of flavophospholipol on fecal microbiota in weaned pigs challenged with Salmonella Typhimurium. Porcine Health Manag 6:14. https://doi.org/10.1186/s40813-020-00151-5
Fulde M, Sommer F, Chassaing B, van Vorst K, Dupont A, Hensel M, Basic M, Klopfleisch R, Rosenstiel P, Bleich A, Backhed F, Gewirtz AT, Hornef MW (2018) Neonatal selection by Toll-like receptor 5 influences long-term gut microbiota composition. Nature 560(7719):489–493. https://doi.org/10.1038/s41586-018-0395-5
Mach N, Berri M, Estelle J, Levenez F, Lemonnier G, Denis C, Leplat JJ, Chevaleyre C, Billon Y, Dore J, Rogel-Gaillard C, Lepage P (2015) Early-life establishment of the swine gut microbiome and impact on host phenotypes. Environ Microbiol Rep 7(3):554–569. https://doi.org/10.1111/1758-2229.12285
Massacci FR, Berri M, Lemonnier G, Guettier E, Blanc F, Jardet D, Rossignol MN, Mercat M-J, Doré J, Lepage P, Rogel-Gaillard C, Estellé J (2020) Late weaning is associated with increased microbial diversity and Faecalibacterium prausnitzii abundance in the fecal microbiota of piglets. Anim Microbiome 2(1):2. https://doi.org/10.1186/s42523-020-0020-4
Hjorth MF, Blaedel T, Bendtsen LQ, Lorenzen JK, Holm JB, Kiilerich P, Roager HM, Kristiansen K, Larsen LH, Astrup A (2019) Prevotella-to-Bacteroides ratio predicts body weight and fat loss success on 24-week diets varying in macronutrient composition and dietary fiber: results from a post-hoc analysis. Int J Obes (Lond) 43(1):149–157. https://doi.org/10.1038/s41366-018-0093-2
Ramayo-Caldas Y, Mach N, Lepage P, Levenez F, Denis C, Lemonnier G, Leplat JJ, Billon Y, Berri M, Dore J, Rogel-Gaillard C, Estelle J (2016) Phylogenetic network analysis applied to pig gut microbiota identifies an ecosystem structure linked with growth traits. ISME J 10(12):2973–2977. https://doi.org/10.1038/ismej.2016.77
Liang H, Dai Z, Liu N, Ji Y, Chen J, Zhang Y, Yang Y, Li J, Wu Z, Wu G (2018) Dietary L-tryptophan modulates the structural and functional composition of the intestinal microbiome in weaned piglets. Front Microbiol 9:1736. https://doi.org/10.3389/fmicb.2018.01736
Holman DB, Brunelle BW, Trachsel J, Allen HK (2017) Meta-analysis to define a core microbiota in the swine gut. mSystems. https://doi.org/10.1128/mSystems.00004-17
Adhikari B, Kim SW, Kwon YM (2019) Characterization of microbiota associated with digesta and mucosa in different regions of gastrointestinal tract of nursery pigs. Int J Mol Sci. https://doi.org/10.3390/ijms20071630
Mariat D, Firmesse O, Levenez F, Guimaraes V, Sokol H, Dore J, Corthier G, Furet JP (2009) The Firmicutes/bacteroidetes ratio of the human microbiota changes with age. BMC Microbiol 9:123. https://doi.org/10.1186/1471-2180-9-123
Ley RE, Turnbaugh PJ, Klein S, Gordon JI (2006) Microbial ecology: human gut microbes associated with obesity. Nature 444(7122):1022–1023. https://doi.org/10.1038/4441022a
Guo X, Xia X, Tang R, Zhou J, Zhao H, Wang K (2008) Development of a real-time PCR method for Firmicutes and Bacteroidetes in faeces and its application to quantify intestinal population of obese and lean pigs. Lett Appl Microbiol 47(5):367–373. https://doi.org/10.1111/j.1472-765X.2008.02408.x
Furet JP, Kong LC, Tap J, Poitou C, Basdevant A, Bouillot JL, Mariat D, Corthier G, Dore J, Henegar C, Rizkalla S, Clement K (2010) Differential adaptation of human gut microbiota to bariatric surgery-induced weight loss: links with metabolic and low-grade inflammation markers. Diabetes 59(12):3049–3057. https://doi.org/10.2337/db10-0253
Naito E, Yoshida Y, Makino K, Kounoshi Y, Kunihiro S, Takahashi R, Matsuzaki T, Miyazaki K, Ishikawa F (2011) Beneficial effect of oral administration of Lactobacillus casei strain Shirota on insulin resistance in diet-induced obesity mice. J Appl Microbiol 110(3):650–657. https://doi.org/10.1111/j.1365-2672.2010.04922.x
Wang Q, Dong J, Zhu Y (2012) Probiotic supplement reduces risk of necrotizing enterocolitis and mortality in preterm very low-birth-weight infants: an updated meta-analysis of 20 randomized, controlled trials. J Pediatr Surg 47(1):241–248. https://doi.org/10.1016/j.jpedsurg.2011.09.064
Neuman H, Forsythe P, Uzan A, Avni O, Koren O (2018) Antibiotics in early life: dysbiosis and the damage done. FEMS Microbiol Rev 42(4):489–499. https://doi.org/10.1093/femsre/fuy018
Zhernakova A, Kurilshikov A, Bonder MJ, Tigchelaar EF, Schirmer M, Vatanen T, Mujagic Z, Vila AV, Falony G, Vieira-Silva S, Wang J, Imhann F, Brandsma E, Jankipersadsing SA, Joossens M, Cenit MC, Deelen P, Swertz MA, Life Lines cohort study, Weersma RK, Feskens EJ, Netea MG, Gevers D, Jonkers D, Franke L, Aulchenko YS, Huttenhower C, Raes J, Hofker MH, Xavier RJ, Wijmenga C, Fu J (2016) Population-based metagenomics analysis reveals markers for gut microbiome composition and diversity. Science 352(6285):565–569. https://doi.org/10.1126/science.aad3369
Ferrer M, Mendez-Garcia C, Rojo D, Barbas C, Moya A (2017) Antibiotic use and microbiome function. Biochem Pharmacol 134:114–126. https://doi.org/10.1016/j.bcp.2016.09.007
Quan LH, Zhang C, Dong M, Jiang J, Xu H, Yan C, Liu X, Zhou H, Zhang H, Chen L, Zhong FL, Luo ZB, Lam SM, Shui G, Li D, Jin W (2019) Myristoleic acid produced by enterococci reduces obesity through brown adipose tissue activation. Gut. https://doi.org/10.1136/gutjnl-2019-319114
Niu Q, Li P, Hao S, Zhang Y, Kim SW, Li H, Ma X, Gao S, He L, Wu W, Huang X, Hua J, Zhou B, Huang R (2015) Dynamic distribution of the gut microbiota and the relationship with apparent crude fiber digestibility and growth stages in pigs. Sci Rep 5:9938. https://doi.org/10.1038/srep09938
Muniz Pedrogo DA, Jensen MD, Van Dyke CT, Murray JA, Woods JA, Chen J, Kashyap PC, Nehra V (2018) Gut microbial carbohydrate metabolism hinders weight loss in overweight adults undergoing lifestyle intervention with a volumetric diet. Mayo Clin Proc 93(8):1104–1110. https://doi.org/10.1016/j.mayocp.2018.02.019
Soeiro C, Quilici IR, Legoff A, Oussalah MB, Morin M, Alauzet C, Charmillon A (2019) Hepatic abscess due to Dialister pneumosintes—a case report. Anaerobe 59:35–37. https://doi.org/10.1016/j.anaerobe.2019.05.006
Patil Y, Gooneratne R, Ju XH (2019) Interactions between host and gut microbiota in domestic pigs: a review. Gut Microbes. https://doi.org/10.1080/19490976.2019.1690363
Yu T, Wang Y, Chen S, Hu M, Wang Z, Wu G, Ma X, Chen Z, Zheng C (2017) Low-molecular-weight chitosan supplementation increases the population of prevotella in the cecal contents of weanling pigs. Front Microbiol 8:2182. https://doi.org/10.3389/fmicb.2017.02182
Stamm LV (2010) Global challenge of antibiotic-resistant Treponema pallidum. Antimicrob Agents Chemother 54(2):583–589. https://doi.org/10.1128/AAC.01095-09
Giannenas I, Doukas D, Karamoutsios A, Tzora A, Bonos E, Skoufos I, Tsinas A, Christaki E, Tontis D, Florou-Paneri P (2016) Effects of Enterococcus faecium, mannan oligosaccharide, benzoic acid and their mixture on growth performance, intestinal microbiota, intestinal morphology and blood lymphocyte subpopulations of fattening pigs. Anim Feed Sci Technol 220:159–167. https://doi.org/10.1016/j.anifeedsci.2016.08.003
Adamowicz EM, Flynn J, Hunter RC, Harcombe WR (2018) Cross-feeding modulates antibiotic tolerance in bacterial communities. ISME J 12(11):2723–2735. https://doi.org/10.1038/s41396-018-0212-z
Wang Y, Cao P, Wang L, Zhao Z, Chen Y, Yang Y (2017) Bacterial community diversity associated with different levels of dietary nutrition in the rumen of sheep. Appl Microbiol Biotechnol 101(9):3717–3728. https://doi.org/10.1007/s00253-017-8144-5
Mills EL, Pierce KA, Jedrychowski MP, Garrity R, Winther S, Vidoni S, Yoneshiro T, Spinelli JB, Lu GZ, Kazak L, Banks AS, Haigis MC, Kajimura S, Murphy MP, Gygi SP, Clish CB, Chouchani ET (2018) Accumulation of succinate controls activation of adipose tissue thermogenesis. Nature 560(7716):102–106. https://doi.org/10.1038/s41586-018-0353-2
Cani PD, Delzenne NM (2009) The role of the gut microbiota in energy metabolism and metabolic disease. Curr Pharm Des 15(13):1546–1558. https://doi.org/10.2174/138161209788168164
Neis EP, Dejong CH, Rensen SS (2015) The role of microbial amino acid metabolism in host metabolism. Nutrients 7(4):2930–2946. https://doi.org/10.3390/nu7042930
Barko PC, McMichael MA, Swanson KS, Williams DA (2018) The Gastrointestinal Microbiome: a review. J Vet Intern Med 32(1):9–25. https://doi.org/10.1111/jvim.14875
LeBlanc JG, Milani C, de Giori GS, Sesma F, van Sinderen D, Ventura M (2013) Bacteria as vitamin suppliers to their host: a gut microbiota perspective. Curr Opin Biotechnol 24(2):160–168. https://doi.org/10.1016/j.copbio.2012.08.005
Koh A, De Vadder F, Kovatcheva-Datchary P, Backhed F (2016) From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell 165(6):1332–1345. https://doi.org/10.1016/j.cell.2016.05.041
Zhang L, Wu W, Lee YK, Xie J, Zhang H (2018) Spatial heterogeneity and co-occurrence of mucosal and luminal microbiome across swine intestinal tract. Front Microbiol 9:48. https://doi.org/10.3389/fmicb.2018.00048
Kim HB, Isaacson RE (2015) The pig gut microbial diversity: Understanding the pig gut microbial ecology through the next generation high throughput sequencing. Vet Microbiol 177(3–4):242–251. https://doi.org/10.1016/j.vetmic.2015.03.014
Acknowledgements
Many thanks to Shenzhen Kingsino Technology Co., Ltd. (www.kingsino.cn) for providing experimental samples. Thanks to Realbio Technology Co., Ltd (https://www.realbio.cn/) for providing sequencing platform and technical assistance in data analyses. We would like to thank EssayStar (https://essaystar.com) for English language editing.
Funding
This research was funded by the National Key Research and Development Program of China (2017YFD0501000, 2018YFD0500600), Shenzhen Key Technology Projects (JSGG20180507182028625), and the National Natural Science Foundation of China (81770434).
Author information
Authors and Affiliations
Contributions
Conceptualization, YC and HS; formal analysis, YL; resources, YZ and HW; writing—original draft preparation, YL; writing—review and editing, YZ, HW, YC and HS. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical Approval
The animal experiment was reviewed and approved by the Institutional Animal Care and Use Committee of Sun Yat-Sen University (Approval No. SYSU-IACUC-2019-B563) in accordance with the animal ethics guidelines set by Laboratory Animal Center (SYXK [Yue] 2016–0112).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, Y., Zhu, Y., Wei, H. et al. Study on the Diversity and Function of Gut Microbiota in Pigs Following Long-Term Antibiotic and Antibiotic-Free Breeding. Curr Microbiol 77, 4114–4128 (2020). https://doi.org/10.1007/s00284-020-02240-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-020-02240-8