Abstract
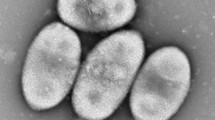
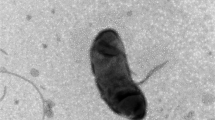
A novel strain KS-M25T was isolated from estuary water in South Korea. Strain KS-M25T was Gram-staining-negative, strictly aerobic, motile rods-shaped bacterium and showed oxidase- and catalase-positive reactions. Growth of strain KS-M25T was observed at 10–25 °C (optimum, 20 °C), at pH 5.5–9.0 (optimum, pH 7.5), and with 0–6.0% (w/v) NaCl (optimum, 1%). Ubiquinone-8 was identified as the sole isoprenoid quinone and the major fatty acids were C16:0, cyclo–C17:0 and sum in feature 3 (comprising C16:1 ω7c and/or C16:1 ω6c). The G+C content values based on genome sequences was 62.3 mol%. Phylogenetic analysis based on 16S rRNA gene sequences showed that strain KS-M25T formed a clear cluster within the genus Achromobacter. Strain KS-M25T was most closely related to Achromobacter anxifer LMG 26857T, Achromobacter dolens LMG 26840T, and Achromobacter xylosoxidans NBRC 15126T with 97.8%, 97.8%, and 97.7% 16S rRNA gene sequence similarities, respectively. Based on phenotypic, chemotaxonomic, and phylogenetic analysis, strain KS-M25T represents a novel species of the genes Achromobacter, for which the name Achromobacter aestuarii sp. nov. is proposed. The type strain is KS-M25T (= KACC 21219T = JCM 33329T).

Similar content being viewed by others
References
Bergey DH, Harrison FC, Breed RS, Hammer BW, Huntton FM (1923) Bergey’s manual of determinative bacteriology. The Williams & Wilkins Co., Baltimore, USA, Baltimore
Skerman VBD, McGowan V, Sneath PHA (1980) Approved lists of bacterial names. Int J Syst Evol Microbiol 30:225–420
Yabuuchi E, Yano I (1981) Achromobacter gen. nov. and Achromobacter xylosoxidans (ex Yabuuchi and Ohyama 1971) nom. rev. Int J Syst Evol Microbiol 31:477–478
Yabuuchi E, Kawamura Y, Kosako Y, Ezaki T (1998) Emendation of genus Achromobacter and Achromobacter xylosoxidans (Yabuuchi and Yano) and proposal of Achromobacter ruhlandii (Packer and Vishniac) comb. nov., Achromobacter piechaudii (Kiredjian et al.) comb. nov., and Achromobacter xylosoxidans subsp. denitrificans (Ru¨ger and Tan) comb. nov. Microbiol Immunol 42:429–438
Packer L, Vishniac W (1955) Chemosynthetic fixation of carbon dioxide and characteristics of hydrogenase in resting cell suspensions of Hydrogenomonas ruhlandii nov. spec. J Bacteriol 70:216–223
Kiredjian M, Holmes B, Kersters K, Guilvout I, Deley J (1986) Alcaligenes piechaudii, a new species from human clinical specimens and the environment. Int J Syst Evol Microbiol 36:282–287
Rüger HJ, Tan TL (1983) Separation of Alcaligenes denitrificans sp. nov., nom. rev. from Alcaligenes faecalis on the basis of DNA base composition, DNA homology, and nitrate reduction. Int J Syst Evol Microbiol 33:85–89
Kim JM, Le NT, Chung BS, Park JH, Bae JW, Madsen EL, Jeon CO (2008) Influence of soil components on the biodegradation of benzene, toluene, ethylbenzene, and o-, m-, and p-xylenes by the newly isolated bacterium Pseudoxanthomonas spadix BD-a59. Appl Environ Microbiol 74:7313–7320
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: A taxonomically united database of 16S rRNA and whole genome assemblies. Int J Syst Evol Microbiol 67:1613–1617
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacterial 173:697–703
Nawrocki EP, Eddy SR (2007) Query-dependent banding (QDB) for faster RNA similarity searches. PLoS Comput Biol 3:e56
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Luo R, Liu B, Xie Y, Li Z, Huang W, Yuan J, He G, Chen Y, Pan Q, Lu Y, Tang J, Wu G, Zhang H, Shi Y, Liu Y, Yu C, Wang B, Lu Y, Han C, Cheung DW, Yiu S, Peng S, Xiaoqian Z, Liu G, Liao X, Li Y, Yang H, Wang J, Lam T, Wang J (2012) SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1:18
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:24690–32461
Richter M, Rosselló-Móra R, Oliver Glöckner F, Peplies J (2016) JSpeciesWS: a web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics 32:929–931
Meier-Kolthof JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:60
Gomori G (1955) Preparation of buffers for use in enzyme studies. Methods Enzymol 1:138–146
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington, DC, pp 607–654
Lányi B (1987) Classical and rapid identification methods for medically important bacteria. Methods Microbiol 19:1–67
Minnikin DE, O’donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Komagata K, Suzuki KI (1988) 4 Lipid and cell-wall analysis in bacterial systematics. Methods Microbiol 19:161–208
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids, MIDI Technical Note 101. MIDI Inc, Newark
Bossemeyer D, Borchard A, Dosch DC, Helmer GC, Epstein W, Booth IR, Bakker EP (1989) K+-transport protein TrkA of Escherichia coli is a peripheral membrane protein that requires other trk gene products for attachment to the cytoplasmic membrane. J Biol Chem 264:16403–16410
Dosch DC, Helmer GL, Sutton SH, Salvacion FF, Epstein W (1991) Genetic analysis of potassium transport loci in Escherichia coli: evidence for three constitutive systems mediating uptake potassium. J Bacteriol 173:687–696
Matsuda N, Kobayashi H, Katoh H, Ogawa T, Futatsugi L, Nakamura T, Bakker EP, Uozumi N (2004) Na+ dependent K+ uptake Ktr system from the cyanobacterium Synechocystis sp. PCC 6803 and its role in the early phases of cell adaptation to hyperosmotic shock. J Biol Chem 279:54952–54962
Ali MK, Li XF, Tang Q, Liu XY, Chen F, Xiao JF, Ali M, Chou SH, He J (2017) Regulation of inducible potassium transporter KdpFABC by the KdpD/KdpE two-component system in Mycobacterium smegmatis. Front Microbiol 8:570
Ness LS, Booth IR (1999) Different foci for the regulation of the activity of the KefB and KefC glutathione-gated K+ efflux systems. J Biol Chem 274:9524–9530
Kung C, Martinac B, Sukharev S (2010) Mechanosensitive channels in microbes. Annu Rev Microbiol 64:313–329
Vimont S, Berche P (2000) NhaA, an Na+/H+ antiporter involved in environmental survival of Vibrio cholerae. J Bacteriol 182:2937–2944
Kosono S, Haga K, Tomizawa R, Kajiyama Y, Hatano K, Takeda S, Wakai Y, Hino M, Kudo T (2005) Characterization of a multigene-encoded sodium/ hydrogen antiporter (Sha) from Pseudomonas aeruginosa: its involvement in pathogenesis. J Bacteriol 187:5242–5248
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA 106:19126–19131
Coenye T, Vancanneyt M, Falsen E, Swings J, Vandamme P (2003) Achromobacter insolitus sp. nov. and Achromobacter spanius sp. nov., from human clinical samples. Int J Syst Evol Microbiol 53:1819–1824
Vandamme P, Moore ER, Cnockaert M, Peeters C, Svensson-Stadler L, Houf K, Spilker T, LiPuma JJ (2013) Classification of Achromobacter genogroups 2, 5, 7 and 14 as Achromobacter insuavis sp. nov., Achromobacter aegrifaciens sp. nov., Achromobacter anxifer sp. nov. and Achromobacter dolens sp. nov., respectively. Syst Appl Microbiol 36:474–482
Acknowledgements
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (No. 2017R1C1B5077091) and the National Institute of Fisheries Science (NIFS, R2020051).
Author information
Authors and Affiliations
Contributions
SCK isolated the type strain from an estuary and performed the experiments. SOC analyzed the genome. HJL supervised all works and wrote the manuscript. The manuscript has been reviewed and edited by all authors.
Corresponding author
Ethics declarations
Conflicts of interests
The authors declare no competing financial conflicts of interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kim, S.C., Chung, S.O. & Lee, H.J. Achromobacter aestuarii sp. nov., Isolated from an Estuary. Curr Microbiol 78, 411–416 (2021). https://doi.org/10.1007/s00284-020-02231-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-020-02231-9