Abstract
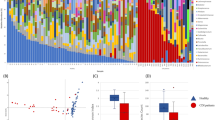
Clostridioides difficile infection (CDI) is one of the most common causes of nosocomial diarrhea in developed countries and the main cause in healthcare settings. This case–control study was designed to evaluate the composition of the gut microbiota dominant bacterial groups in patients with CDI compared to the healthy control subjects. A total of 100 adult subjects involving 50 inpatients with CDI and 50 healthy persons were enrolled in the study. C. difficile isolates were characterized according to the anaerobic culture and presence of toxin genes with multiplex PCR. An ecological analysis was performed real-time quantitative PCR for bacterial elements. The abundances of Enterococcus spp., Lactobacillus spp., Escherichia coli, C. difficile, and Akkermansia muciniphila were higher in group CDI compared with group HC (P < 0.05). The abundances of Bacteroides spp., Bifidobacterium spp., and Faecalibacterium prausnitzii were lower in group CDI than in group HC (P < 0.05). No significant difference was observed in the copy number of Prevotella genus between the CDI and HC subjects (P-value = 0.087). We observed that economic status and income levels were reduced at patients with CDI, however, there was no significant difference between CDI and HC group results and other variables, such as age, BMI, and educational level. These findings showed a reduction in butyrate-producing bacteria and increase in lactic acid-producing bacteria was seen in CDI status. Overrepresentation of Akkermansia may be a predictive marker for the development of nosocomial diarrhea can result in a worse CDI prognosis.

Similar content being viewed by others
References
Schneider D, Thürmer A, Gollnow K, Lugert R, Gunka K, Groß U et al (2017) Gut bacterial communities of diarrheic patients with indications of Clostridioides difficile infection. Sci Data 4:170152. https://doi.org/10.1038/sdata.2017.152
Hernandez M, de Frutos M, Rodriguez-Lazaro D, López-Urrutia L, Quijada NM, Eiros JM (2018) Fecal microbiota of toxigenic Clostridioides difficile-associated diarrhea. Front Microbiol 9:3331. https://doi.org/10.3389/fmicb.2018.03331
Shoaei P, Shojaei H, Khorvash F, Ataei B, Esfandiari Z, Vakili B et al (2019) Clostridium difficile infection in cancer patients with hospital-acquired diarrhea at the teaching hospitals in Iran: multilocus sequence typing analysis (MLST) and antimicrobial resistance pattern. Anna Ig iene 31(4):365–373. https://doi.org/10.7416/ai.2019.2298
Milani C, Ticinesi A, Gerritsen J, Nouvenne A, Lugli GA, Mancabelli L et al (2016) Gut microbiota composition and Clostridium difficile infection in hospitalized elderly individuals: a metagenomic study. Sci Rep 6:25945. https://doi.org/10.1038/srep25945
Shoaei P, Shojaei H, Khorvash F, Hosseini SM, Ataei B, Tavakoli H et al (2019) Molecular epidemiology of Clostridium difficile infection in Iranian hospitals. Antimicrob Resist Infect Control 8(1):12. https://doi.org/10.1186/s13756-018-0454-6
Han S-H, Yi J, Kim J-H, Lee S, Moon H-W (2019) Composition of gut microbiota in patients with toxigenic Clostridioides (Clostridium) difficile: Comparison between subgroups according to clinical criteria and toxin gene load. PLoS ONE 14(2):e0212626. https://doi.org/10.1371/journal.pone.0212626
Shoaei P, Shojaei H, Jalali M, Khorvash F, Hosseini SM, Ataei B et al (2019) Clostridium difficile isolated from faecal samples in patients with ulcerative colitis. BMC Infect Dis 19(1):361. https://doi.org/10.1186/s12879-019-3965-8
Rea MC, O'Sullivan O, Shanahan F, O'Toole PW, Stanton C, Ross RP et al (2012) Clostridium difficile carriage in elderly subjects and associated changes in the intestinal microbiota. J Clin Microbiol 50(3):867–875. https://doi.org/10.1128/JCM.05176-11
Skraban J, Dzeroski S, Zenko B, Mongus D, Gangl S, Rupnik M (2013) Gut microbiota patterns associated with colonization of different Clostridium difficile ribotypes. PLoS ONE 8(2):e58005. https://doi.org/10.1371/journal.pone.0058005
Sangster W, Hegarty JP, Schieffer KM, Wright JR, Hackman J, Toole DR et al (2016) Bacterial and fungal microbiota changes distinguish C. difficile infection from other forms of diarrhea: results of a prospective inpatient study. Front Microbiol 7:789. https://doi.org/10.3389/fmicb.2016.00789
Samarkos M, Mastrogianni E, Kampouropoulou O (2018) The role of gut microbiota in Clostridium difficile infection. Eur J Intern Med 50:28–32. https://doi.org/10.1016/j.ejim.2018.02.006
Gu S, Chen Y, Zhang X, Lu H, Lv T, Shen P et al (2016) Identification of key taxa that favor intestinal colonization of Clostridium difficile in an adult Chinese population. Microbes Infect 18(1):30–38. https://doi.org/10.1016/j.micinf.2015.09.008
Schubert AM, Rogers MA, Ring C, Mogle J, Petrosino JP, Young VB et al (2014) Microbiome data distinguish patients with Clostridium difficile infection and non-C. difficile-associated diarrhea from healthy controls. MBio 5(3):e01021–e1114. https://doi.org/10.1128/mBio.01021-14
Fraher MH, O’toole PW, Quigley EM (2012) Techniques used to characterize the gut microbiota: a guide for the clinician. Nat Rev Gastroenterol 9(6):312. https://doi.org/10.1038/nrgastro.2012.44
Eckburg PB, Bik EM, Bernstein CN, Purdom E, Dethlefsen L, Sargent M et al (2005) Diversity of the human intestinal microbial flora. Science 308(5728):1635–1638. https://doi.org/10.1126/science.1110591
Jimeno R, Brailey PM, Barral P (2018) Quantitative polymerase chain reaction-based analyses of murine intestinal microbiota after oral antibiotic treatment. JoVE 141:e58481. https://doi.org/10.3791/58481
Lemee L, Dhalluin A, Pestel-Caron M, Lemeland J-F, Pons J-L (2004) Multilocus sequence typing analysis of human and animal Clostridium difficile isolates of various toxigenic types. J Clin Microbiol 42(6):2609–2617
Rodriguez C, Taminiau B, Avesani V, Van Broeck J, Delmée M, Daube G (2014) Multilocus sequence typing analysis and antibiotic resistance of Clostridium difficile strains isolated from retail meat and humans in Belgium. Food Microbiol 42:166–171. https://doi.org/10.1016/j.fm.2014.03.021
Pitcher D, Saunders N, Owen R (1989) Rapid extraction of bacterial genomic DNA with guanidium thiocyanate. Lett Appl Microbiol 8(4):151–156. https://doi.org/10.1111/j.1472-765x.1989.tb00262x
Stubbs S, Rupnik M, Gibert M, Brazier J, Duerden B, Popoff M (2000) Production of actin-specific ADP-ribosyltransferase (binary toxin) by strains of Clostridium difficile. FEMS Microbiol Lett 186(2):307–312
Carroll IM, Chang Y-H, Park J, Sartor RB, Ringel Y (2010) Luminal and mucosal-associated intestinal microbiota in patients with diarrhea-predominant irritable bowel syndrome. Gut Pathog 2(1):19. https://doi.org/10.1186/1757-4749-2-19
Mello CS, do Carmo RMS, de Araújo FHB, Melli LCFL, Tahan S, Pignatari ACC et al (2018) Fecal microbiota analysis of children with small intestinal bacterial overgrowth among residents of an urban slum in Brazil. J Pediatr (Rio J) 94(5):483–490. https://doi.org/10.1016/j.jped.2017.09.003
Nabizadeh E, Jazani NH, Bagheri M, Shahabi S (2017) Association of altered gut microbiota composition with chronic urticaria. Ann Allergy Asthma Immunol 119(1):48–53. https://doi.org/10.1016/j.anai.2017.05.006
Sun F, Wu D, Qiu Z, Jin M, Li J (2010) Development of real-time PCR systems based on SYBR Green for the specific detection and quantification of Klebsiella pneumoniae in infant formula. Food Control 21(4):487–491. https://doi.org/10.1016/j.foodcont.2009.07.014
Ling Z, Liu X, Jia X, Cheng Y, Luo Y, Yuan L et al (2014) Impacts of infection with different toxigenic Clostridium difficile strains on faecal microbiota in children. Sci Rep 4:7485. https://doi.org/10.1038/srep07485
Buffie CG, Pamer EG (2013) Microbiota-mediated colonization resistance against intestinal pathogens. Nat Rev Immunol 13(11):790. https://doi.org/10.1038/nri3535
Lopez Siles M, Enrich-Capó N, Aldeguer X, Sabat-Mir M, Duncan S, Garcia-Gil J et al (2018) Alterations in the abundance and co-occurrence of Akkermansia muciniphila and Faecalibacterium prausnitzii in the colonic mucosa of inflammatory bowel disease subjects. Front Cell Infect Microbiol 8:281. https://doi.org/10.3389/fcimb.2018.00281
Punni E, Pula JL, Asslo F, Baddoura W, DeBari VA (2015) Is obesity a risk factor for Clostridium difficile infection? Obes Res Clin Pract 9(1):50–54. https://doi.org/10.1016/j.orcp.2013.12.007
Chandradas S, Khalili H, Ananthakrishnan A, Wayman C, Reidel W, Waalen J et al (2018) Does obesity influence the risk of Clostridium difficile infection among patients with ulcerative colitis? Digest Dis Sci 63(9):2445–2450. https://doi.org/10.1007/s10620-018-5108-2
Acknowledgements
The authors would like to express their gratitude to Dr. Parisa Shoaei and Dr. Bahram Bagherpour for their special cooperation in our study. We are thankful to the staff of Isfahan University Training Hospitals and Infectious Disease Research center, Isfahan, University of medical sciences, Isfahan, Iran for their great help in performing tests.
Funding
A part of this article (PCR analysis and sample collection) is performed from research (Grant No. 291235) that performed in the Infectious Disease Research center, Isfahan University of medical sciences, Isfahan, Iran.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical Approval
The Ethics Committee on Clinical Investigation of the Isfahan University of Medical Sciences, Isfahan, Iran, approved this presented research.
Informed Consent
Written consent informed was obtained from all participants.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Vakili, B., Fateh, A., Asadzadeh Aghdaei, H. et al. Characterization of Gut Microbiota in Hospitalized Patients with Clostridioides difficile Infection. Curr Microbiol 77, 1673–1680 (2020). https://doi.org/10.1007/s00284-020-01980-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-020-01980-x