Abstract
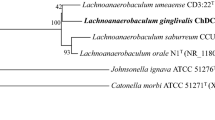
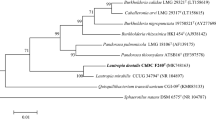
A novel Gram-negative, obligately anaerobic, non-motile, non-spore forming, and rod-shaped bacterium, designated strain JS262T, was isolated from human subgingival plaque of periodontitis lesion and was characterized by polyphasic taxonomic analysis. Comparison of 16S ribosomal RNA gene (16S rDNA) sequence revealed that the strain belonged to the genus Prevotella. The percent similarity of 16S rDNA of strain JS262T was closest to those of Prevotella buccae ATCC 33574T (89.1%) and Prevotella shahii JCM 12083T (88.9%). The major fatty acids of strain JS262T were C16:0 (29.2%), iso-C15:0 (19.2%), and anteiso-C15:0 (16.9%). Complete genome of strain JS262T was 2,691,540 bp in length and the G+C content was 43.9 mol%. Average nucleotide identity and genome-to-genome distance values between strain JS262T and P. buccae ATCC 33574T or P. loescheii DSM 19665T were > 70.4% and > 30.1%, respectively. On the basis of these data, a novel Prevotella species is proposed: Prevotella koreensis sp. nov. The type strain of P. koreensis is JS262T (= KCOM 3155T = JCM 33298T).

Similar content being viewed by others
References
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA, Pevzner PA (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477
Brondz I, Olsen I (1991) Multivariate analyses of cellular fatty acids in Bacteroides, Prevotella, Porphyromonas, Wolinella, and Campylobacter spp. J Clin Microbiol 29:183–189
Cho E, Park SN, Lim YK, Shin Y, Paek J, Hwang CH, Chang YH, Kook JK (2015) Fusobacterium hwasookii sp. nov., isolated from a human periodontitis lesion. Curr Microbiol 70:169–175. https://doi.org/10.1007/s00284-014-0692-7
De A, Varaiya A, Mathur M (2002) Anaerobes in pleuropulmonary infections. Indian J Med Microbiol. 20:150–152
Deng ZL, Szafrański SP, Jarek M, Bhuju S, Wagner-Döbler I (2017) Dysbiosis in chronic periodontitis: key microbial players and interactions with the human host. Sci Rep 7:3703. https://doi.org/10.1038/s41598-017-03804-8
Downes J, Sutcliffe IC, Tanner ACR, Wade WG (2005) Prevotella marshii sp. nov. and Prevotella baroniae sp. nov. isolated from the human oral cavity. Int J Syst Evol Microbiol 55:1551–1555. https://doi.org/10.1099/ijs.0.63634-0
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Kim HS, Lee DS, Chang YH, Kim MJ, Koh S, Kim J, Seong JH, Song SK, Shin HS, Son JB, Jung MY, Park SN, Yoo SY, Cho KW, Kim DK, Moon S, Kim D, Choi Y, Kim BO, Jang HS, Kim CS, Kim C, Choe SJ, Kook JK (2010) Application of rpoB and zinc protease gene for use in molecular discrimination of Fusobacterium nucleatum subspecies. J Clin Microbiol 48:545–553. https://doi.org/10.1128/JCM.01631-09
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Lee I, Kim YO, Park SC, Chun J (2016) OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103. https://doi.org/10.1099/ijsem.0.000760
Luo R, Liu B, Xie Y, Li Z, Huang W, Yuan J, He G, Chen Y, Pan Q, Liu Y, Tang J, Wu G, Zhang H, Shi Y, Liu Y, Yu C, Wang B, Lu Y, Han C, Cheung DW, Yiu SM, Peng S, Xiaoqian Z, Liu G, Liao X, Li Y, Yang H, Wang J, Lam TW, Wang J (2012) SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1:18. Erratum in: Gigascience (2015) 4:30
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:60. https://doi.org/10.1186/1471-2105-14-60
Rôças IN, Siqueira JF Jr (2018) Frequency and levels of candidate endodontic pathogens in acute apical abscesses as compared to asymptomatic apical periodontitis. PLoS ONE 13:e0190469. https://doi.org/10.1371/journal.pone.0190469
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sakamoto M, Suzuki M, Huang Y, Umeda M, Ishikawa I, Benno Y (2004) Prevotella shahii sp. nov. and Prevotella salivae sp. nov., isolated from the human oral cavity. Int J Syst Evol Microbiol 54:877–883. https://doi.org/10.1099/ijs.0.02876-0
Shah HN, Collins DM (1990) Prevotella, a new genus to include Bacteroides melaninogenicus and related species formerly classified in the genus Bacteroides. Int J Syst Bacteriol 40:205–208
Shah HN, Collins MD (1981) Bacteroides buccalis, sp. nov., Bacteroides denticola, sp. nov., and Bacteroides pentosaceus, sp. nov., new species of the genus Bacteroides from the oral cavity. Zbl Bakt Hyg I Abt Orig C 2:235–241
Shah HN, Gharbia SE, Duerden BI (1998) Bacteroides, Prevotella and Porphyromonas. In: Collier L, Balows A, Sussman M (eds) Topley & Wilson’s microbiology and microbial infections. Arnold, London, pp 1305–1330
Stackebrandt E, Goebel BM (2006) Taxonomic parameters revisited: tarnished gold standards. Microbiol Today 33:152–155
Subramaniam A, Kumar R, Cliver SP, Zhi D, Szychowski JM, Abramovici A, Biggio JR, Lefkowitz EJ, Morrow C, Edwards RK (2016) Vaginal microbiota in pregnancy: evaluation based on vaginal flora, birth outcome, and race. Am J Perinatol 33:401–408. https://doi.org/10.1055/s-0035-1565919
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Tatusova T, DiCuccio M, Badretdin A, Chetvernin V, Nawrocki EP, Zaslavsky L, Lomsadze A, Pruitt KD, Borodovsky M, Ostell J (2016) NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res 44:6614–6624. https://doi.org/10.1093/nar/gkw569
Usin MM, Menso J, Rodríguez VI, González A, Tabares S, Parodi R, Sembaj A (2016) Association between maternal periodontitis and preterm and/or low birth weight infants in normal pregnancies. J Matern Fetal Neonatal Med 29:115–119. https://doi.org/10.3109/14767058.2014.987751
Walker BJ, Abeel T, Shea T, Priest M, Abouelliel A, Sakthikumar S, Cuomo CA, Zeng Q, Wortman J, Young SK, Earl AM (2014) Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9:e112963. https://doi.org/10.1371/journal.pone.0112963
Williams A, Kerkering T (2017) Prevotella osteomyelitis after dental capping procedure. IDCases 8:32–33. https://doi.org/10.1016/j.idcr.2017.01.005
Wu PC, Tu MS, Lin PH, Chen YS, Tsai HC (2014) Prevotella brain abscesses and stroke following dental extraction in a young patient: a case report and review of the literature. Intern Med 53(16):1881–1887. https://doi.org/10.2169/internalmedicine.53.1299
Yamashita Y, Takeshita T (2017) The oral microbiome and human health. J Oral Sci 59:201–206. https://doi.org/10.2334/josnusd.16-0856
Acknowledgements
This research was supported by the Bio & Medical Technology Development Program of the National Research Foundation (NRF) funded by the Ministry of Science and ICT (Grant No. 2017M3A9B8065844), in part by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (Grant No. 2017R1A2B4004894), and in part the KRIBB Research Initiative Program funded by the Ministry of Science, ICT and Future Planning.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
DPD number: TA00867
GenBank accession number of 16S rRNA gene for strain JS262T: MK452274
GenBank accession number of genome for strain JS262T: RYYU00000000
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Park, SN., Lim, Y.K., Shin, J.H. et al. Prevotella koreensis sp. nov., Isolated from Human Subgingival Dental Plaque of Periodontitis Lesion. Curr Microbiol 76, 1055–1060 (2019). https://doi.org/10.1007/s00284-019-01720-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-019-01720-w