Abstract
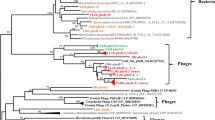
Although bacteriophages are the most abundant biological entities on the planet, their genetic diversity, especially in natural wetlands, is poorly understood. In this study, the genetic diversity of cyanopodoviruses in sediments of two coastal wetlands in Northeast China was investigated by targeting the DNA polymerase (pol) gene. A total of 66 DNA pol clones were obtained. A BLAST search at the amino acid level showed that the obtained sequences had the highest identity ranged from 83 to 99% to the known sequences. A phylogenetic tree showed that the distribution patterns of DNA pol sequence were different between two wetland soils, and 29 clones of this study formed four wetland-specific groups, which suggested that unrevealed novel groups of cyanopodovirus inhabited in wetlands. In addition, nonmetric multidimensional scaling (NMDS) analysis of all DNA pol sequences from various environments showed that cyanopodovirus communities of coastal wetlands are in the intermediate position between marine water environments and terrestrial freshwater environments, which highlights that the coastal wetlands as transitional zones between inland freshwater environments and marine environments.



Similar content being viewed by others
References
Adriaenssens EM, Cowan DA (2014) Using signature genes as tools to assess environmental viral ecology and diversity. Appl Environ Microb 80:4470–4480. https://doi.org/10.1128/AEM.00878-14
Balasooriya WK, Denef K, Peters J, Verhoest NEC, Boeckx P (2007) Vegetation composition and soil microbial community structural changes along a wetland hydrological gradient. Hydrol Earth Syst Sci 12:277–291. https://doi.org/10.5194/hess-12-277-2008
Bergh Ø, BØrsheim KY, Bratbak G, Heldal M (1989) High abundance of viruses found in aquatic environments. Nature 340:467–468. https://doi.org/10.1038/340467a0
Chen F, Wang K, Huang SJ, Cai HY, Zhao MR, Jiao NZ, Wommack KE (2009) Diverse and dynamic populations of cyanobacterial podoviruses in the Chesapeake Bay unveiled through DNA polymerase gene sequences. Environ Microbiol 11:2884–2892. https://doi.org/10.1111/j.1462-2920.2009.02033.x
Davidsson TE, Stepanauskas R, Leonardson L (1997) Vertical patterns of nitrogen transformations during infiltration in two wetland soils. Appl Environ Microb 63:3648–3656. https://doi.org/10.1016/S0027-5107(97)00112-7
Dekel-Bird NP, Avrani S, Sabehi G, Pekarsky I, Marston MF, Kirzner S, Lindell D (2013) Diversity and evolutionary relationships of T7-like podoviruses infecting marine cyanobacteria. Environ Microbiol 15:1476–1491. https://doi.org/10.1111/1462-2920.12103
Dorigo U, Jacquet S, Humbert JF (2004) Cyanophage diversity, inferred from g20 gene analyses, in the largest natural lake in France, Lake Bourget. Appl Environ Microb 70:1017–1022
Filée J, Tétart F, Suttle CA, Krisch HM (2005) Marine T4-type bacteriophages, a ubiquitous component of the dark matter of the biosphere. PNAS 102:12471–12476. https://doi.org/10.1128/AEM.70.2.1017-1022.2004
Fuhrman JA (1999) Marine viruses and their biogeochemical and ecological effects. Nature 399:541–548. https://doi.org/10.1038/21119
Güemes AGC, Youle M, Cantú VA, Felts B, Nulton J, Rohwer F (2016) Viruses as winners in the game of life. Annu Rev Virol 3:197–214. https://doi.org/10.1146/annurev-virology-100114-054952
Gutknecht JLM, Goodman RM, Balser TC (2006) Linking soil process and microbial ecology in freshwater wetland ecosystems. Plant Soil 289:17–34. https://doi.org/10.1007/s11104-006-9105-4
Huang SJ, Wilhelm SW, Jiao NZ, Chen F (2010) Ubiquitous cyanobacterial podoviruses in the global oceans unveiled through viral DNA polymerase gene sequences. ISME J 4:1243–1251. https://doi.org/10.1038/ismej.2010.56
Ji XL, Zhang CJ, Kuang AX, Li JK, Cui YS, Qin KH, Lin LB, Cheng BX, Zhang Q, Wei YL (2015) Morphological diversity of cultured cold-active lytic bacteriophages isolated from the Napahai plateau wetland in China. Virol Sin 30:457–459. https://doi.org/10.1007/s12250-015-3674-4
Jia ZJ, Ishihara R, Nakajima Y, Asakawa S, Kimura M (2007) Molecular characterization of T4-type bacteriophages in a rice field. Environ Microbiol 9:1091–1096. https://doi.org/10.1111/j.1462-2920.2006.01207.x
Jing RY, Liu JJ, Yu ZH, Liu XB, Wang GH (2014) Phylogenetic distribution of the capsid assembly protein gene (g20) of cyanophages in paddy floodwaters in Northeast China. PLoS ONE 9:e88634. https://doi.org/10.1111/j.1462-2920.2006.01207.x
Labrie SJ, Frois-Moniz K, Osburne MS, Kelly L, Roggensack SE, Sullivan MB, Gearin G, Zeng Q, Fitzgerald M, Henn MR, Chisholm SW (2013) Genomes of marine cyanopodoviruses reveal multiple origins of diversity. Environ Microbiol 15:1356–1376. https://doi.org/10.1111/1462-2920.12053
Liu JJ, Wang GH, Zheng CY, Yuan XH, Jin J, Liu XB (2011) Specific assemblages of major capsid genes (g23) of T4-type bacteriophages isolated from upland black soils in Northeast China. Soil Biol Biochem 43:1980–1984. https://doi.org/10.1016/j.soilbio.2011.05.005
Millard AD, Mann NH (2006) A temporal and spatial investigation of cyanophage abundance in the Gulf of Aqaba, Red Sea. J Mar Biol Assoc UK 86:507–515. https://doi.org/10.1017/S0025315406013415
Mühling M, Fuller NJ, Millard A, Somerfield PJ, Marie D, Wilson WH, Scanlan DJ, Post AF, Joint I, Mann NH (2005) Genetic diversity of marine Synechococcus and co-occurring cyanophage communities: evidence for viral control of phytoplankton. Environ Microbiol 7:499–508. https://doi.org/10.1111/j.1462-2920.2004.00713.x
Partensky F, Hess WR, Vaulot D (1999) Prochlorococcus, a marine photosynthetic prokaryote of global significance. Microbiol Mol Biol R 63:106–127. https://doi.org/10.0000/PMID10066832
Rohwer F, Edwards R (2002) The phage proteomic tree: a genome-based taxonomy for phage. J Bacteriol 184:4529–4535. https://doi.org/10.1128/jb.184.16.4529-4535.2002
Rohwer F, Thurber RV (2009) Viruses manipulate the marine environment. Nature 459:207–212. https://doi.org/10.1038/nature08060
Roux S, Hallam SJ, Woyke T, Sullivan MB (2015) Viral dark matter and virus-host interactions resolved from publicly available microbial genomes. eLife 4:e08490. https://doi.org/10.7554/elife.08490
Sabehi G, Shaulov L, Silver DH, Yanai I, Harel A, Lindell D (2012) A novel lineage of myoviruses infecting cyanobacteria is widespread in the oceans. PNAS 109:2037–2042. https://doi.org/10.1073/pnas.1115467109
Short CM, Suttle CA (2005) Nearly identical bacteriophage structural gene sequences are widely distributed in both marine and freshwater environments. Appl Environ Microb 71:480–486. https://doi.org/10.1128/AEM.71.1.480-486.2005
Sullivan MB, Waterbury JB, Chisholm SW (2003) Cyanophages infecting the oceanic cyanobacterium Prochlorococcus. Nature 424:1047–1051. https://doi.org/10.1038/nature01929
Suttle CA (2000) Cyanophages and their role in the ecology of cyanobacteria. In: Brian AW, Malcolm P (eds) the ecology of cyanobacteria. Springer, Dordrecht, pp 563–589
Suttle CA (2005) Viruses in the sea. Nature 437:365–369. https://doi.org/10.1038/nature04160
Suttle CA, Chan AM (1993) Marine cyanophages infecting oceanic and coastal strains of Synechococcus: abundance, morphology, cross-reactivity and growth characteristics. Mar Ecol Prog Ser 92:99–109. https://doi.org/10.3354/meps092099
Tamura K, Stecher G, Peterson D, Filipski Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Team RC (2013) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882. https://doi.org/10.1093/nar/25.24.4876
Wang GH, Asakawa S, Kimura M (2011) Spatial and temporal changes of cyanophage communities in paddy field soils as revealed by the capsid assembly protein gene g20. FEMS Microb Ecol 76:352–359. https://doi.org/10.1111/j.1574-6941.2011.01052.x
Wang GH, Liu JJ, Yu ZH, Jin J, Liu XB (2014) Unique distribution of cyanobacterial podoviruses and their potential hosts in a paddy field of northeast China. FEMS Microb Ecol 90:331–334. https://doi.org/10.1111/1574-6941.12401
Wang K, Chen F (2008) Prevalence of highly host-specific cyanophages in the estuarine environment. Environ Microbiol 10:300–312. https://doi.org/10.1111/j.1462-2920.2007.01452.x
Wang MN, Ge XY, Wu YQ, Yang XL, Tan B, Zhang YJ, Shi ZL (2015) Genetic diversity and temporal dynamics of phytoplankton viruses in East Lake, China. Virol Sin 30:290–300. https://doi.org/10.1007/s12250-015-3603-6
Wang XZ, Liu JJ, Yu ZH, Jin J, Liu XB, Wang GH (2016) Novel groups of cyanobacterial podovirus DNA polymerase (pol) genes exist in paddy waters in northeast China. FEMS Microb Ecol 92:192. https://doi.org/10.1093/femsec/fiw192
Waterbury JB, Watson SW, Valois FW, Franks DG (1986) Biological and ecological characterization of the marine unicellular cyanobacterium Synechococcus. Can Bull Fish Aquat Sci 214:71–120
Weigele PR, Pope WH, Pedulla ML, Houtz JM, Smith AL, Conway JF, King J, Hatfull GF, Lawrence JG, Hendrix RW (2007) Genomic and structural analysis of Syn9, a cyanophage infecting marine Prochlorococcus and Synechococcus. Environ Microbiol 9:1675–1695. https://doi.org/10.1111/j.1462-2920.2007.01285.x
Weinbauer MG (2004) Ecology of prokaryotic viruses. FEMS Microb Rev 28:127–181. https://doi.org/10.1016/j.femsre.2003.08.001
Wilhelm SW, Carberry MJ, Eldridge ML, Poorvin L, Saxton MA, Doblin MA (2006) Marine and freshwater cyanophages in a Laurentian Great Lake: evidence from infectivity assays and molecular analyses of g20 genes. Appl Environ Microb 72(7):4957–4963. https://doi.org/10.1128/AEM.00349-06
Williamson KE, Wommack KE, Radosevich M (2003) Sampling natural viral communities from soil for culture-independent analyses. Appl Environ Microb 69(11):6628–6633. https://doi.org/10.1128/AEM.69.11.6628-6633.2003
Youle M, Haynes M, Rohwer F (2012) Scratching the surface of biology’s dark matter. In: Witzany G (ed) Viruses: essential agents of life. Springer, Dordrecht, pp 61–80
Zheng CY, Wang GH, Liu JJ, Song CC, Gao HX, Liu XB (2013) Characterization of the major capsid genes (g23) of T4-type bacteriophages in the wetlands of Northeast China. Microb Ecol 65:616–625. https://doi.org/10.1007/s00248-012-0158-z
Zhong X, Jacquet S (2013) Prevalence of viral photosynthetic and capsid protein genes from cyanophages in two large and deep perialpine lakes. Appl Environ Microb 79:7169–7178. https://doi.org/10.1128/AEM.01914-13
Zhong Y, Chen F, Wilhelm SW, Poorvin L, Hodson RE (2002) Phylogenetic diversity of marine cyanophage isolates and natural virus communities as revealed by sequences of viral capsid assembly protein gene g20. Appl Environ Microb 68(4):1576–1584. https://doi.org/10.1128/aem.68.4.1576-1584.2002
Acknowledgements
This study was financially supported by the National Nature Science Foundation of China (41271262). The authors thank two anonymous reviewers for their valuable comments on the revision of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, X., Sun, Y., Liu, J. et al. Molecular Diversity of Cyanopodoviruses in Two Coastal Wetlands in Northeast China. Curr Microbiol 76, 863–871 (2019). https://doi.org/10.1007/s00284-019-01700-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-019-01700-0