Abstract
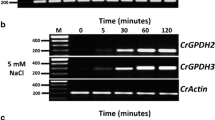
Glycerol-3-phosphate dehydrogenase (GPDH) catalyzes the conversion of dihydroxyacetone phosphate (DHAP) and NADH to glycerol-3-phosphate (G3P) and NAD+. G3P is important as a precursor for glycerol and glycerolipid synthesis in microalgae. A GPDH enzyme has been previously purified from the green microalga Chlamydomonas reinhardtii, however, no genes coding for GPDH have been characterized before. In this study, we report the in silico characterization of three putative GPDH genes from C. reinhardtii: CrGPDH1, CrGPDH2, and CrGPDH3. These sequences showed a significant similarity to characterized GPDH genes from the microalgae Dunaliella salina and Dunaliella viridis. The prediction of the three-dimensional structure of the proteins showed the characteristic fold topology of GPDH enzymes. Furthermore, the phylogenetic analysis showed that the three CrGPDHs share the same clade with characterized GPDHs from Dunaliella suggesting a common evolutionary origin and a similar catalytic function. In addition, the K a/K s ratios of these sequences suggested that they are under purifying selection. Moreover, the expression analysis showed a constitutive expression of CrGPDH1, while CrGPDH2 and CrGPDH3 were induced in response to osmotic stress, suggesting a possible role for these two sequences in the synthesis of glycerol as a compatible solute in osmoregulation, and perhaps also in lipid synthesis in C. reinhardtii. This study has provided a foundation for further biochemical and genetic studies of the GPDH family in this model microalga, and also opportunities to assess the potential of these genes to enhance the synthesis of TAGs for biodiesel production.




Similar content being viewed by others
References
Albertyn J, Hohmann S, Thevelein JM, Prior BA (1994) GPD1, which encodes glycerol-3-phosphate dehydrogenase, is essential for growth under osmotic stress in Saccharomyces cerevisiae, and its expression is regulated by the high-osmolarity glycerol response pathway. Mol Cell Biol 14:4135–4144
Altschul SF, Madden TL, Schäffer AA et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Ansell R, Granath K, Hohmann S, Thevelein JM, Adler L (1997) The two isoenzymes for yeast NAD+-dependent glycerol 3-phosphate dehydrogenase encoded by GPD1 and GPD2 have distinct roles in osmoadaptation and redox regulation. EMBO J 16:2179–2187
Bordoli L, Kiefer F, Arnold K et al (2009) Protein structure homology modeling using SWISS-MODEL workspace. Nature Protoc 4:1–13
Chen H, Lao Y-M, Jiang J-G (2011) Effects of salinities on the gene expression of a (NAD+)-dependent glycerol-3-phosphate dehydrogenase in Dunaliella salina. Sci Total Environ 409:1291–1297
Chen X, Fang H, Rao Z et al (2008) Cloning and characterization of a NAD+-dependent glycerol-3-phosphate dehydrogenase gene from Candida glycerinogenes, an industrial glycerol producer. FEMS Yeast Res 8:725–734
Fan J, Andre C, Xu C (2011) A chloroplast pathway for the de novo biosynthesis of triacylglycerol in Chlamydomonas reinhardtii. FEBS Lett 585:1985–1991
Gee R, Goyal A, Byerrum RU, Tolbert NE (1993) Two isoforms of dihydroxyacetone phosphate reductase from the chloroplast of Dunaliella tertiolecta. Plant Physiol 103:243–249
Goshal D, Mach D, Agarwal M et al (2002) Osmoregulatory isoform of dihydroxyacetone phosphate reductase from Dunaliella tertiolecta: purification and characterization. Protein Expr Purif 24:404–411
Harris EH (1989) The Chlamydomonas sourcebook, a comprehensive guide to biology and laboratory use. Academic Press, Inc, San Diego
He Q, Qiao D, Bai L et al (2007) Cloning and characterization of a plastidic glycerol 3-phosphate dehydrogenase cDNA from Dunaliella salina. J Plant Physiol 164:214–220
He Y, Meng X, Fan Q et al (2009) Cloning and characterization of two novel chloroplastic glycerol-3-phosphate dehydrogenases from Dunaliella viridis. Plant Mol Biol 71:193–205
Heckman DS, Geiser DM, Eidell BR et al (2001) Molecular evidence for the early colonization of land by fungi and plants. Science 293:1129–1133
Herrera-Valencia VA, Contreras-Pool PY, López-Adrián SJ et al (2011) The green microalga Chlorella saccharophila as a suitable source of oil for biodiesel production. Curr Microbiol 63:151–157
Hu Q, Sommerfeld M, Jarvis E et al (2008) Microalgal triacylglycerols as feedstocks for biofuel production: perspectives and advances. Plant J 54:621–639
Husic HD, Tolbert NE (1986) Effect of osmotic stress on carbon metabolism in Chlamydomonas reinhardtii. Plant Physiol 82:594–596
Klöck G, Kreuzberg K (1989) Kinetic properties of a sn-glycerol-3-phosphate dehydrogenase purified from the unicellular alga Chlamydomonas reinhardtii. Biochim Biophys Acta 991:347–352
Larkin M, Blackshields N, Brown R et al (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Larsson C, Pahlman I, Ansell R et al (1998) The importance of the glycerol 3-phosphate shuttle during aerobic growth of Saccharomyces cerevisiae. Yeast 14:347–357
Lee D, Kim M, Ryu Y, Seo J (2008) Cloning and characterization of CmGPD1, the Candida magnoliae homologue of glycerol-3-phosphate dehydrogenase. FEMS Yeast Res 8:1324–1333
León R, Galván F (1994) Halotolerance studies on Chlamydomonas reinhardtii: glycerol excretion by free and immobilized cells. J Appl Phycol 6:13–20
Letunic I, Doerks T, Bork P (2009) SMART 6: recent updates and new developments. Nucleic Acids Res 37:D229–D232
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Miller R, Wu G, Deshpande RR et al (2010) Changes in transcript abundance in Chlamydomonas reinhardtii following nitrogen deprivation predict diversion of metabolism. Plant Physiol 154:1737–1752
Ou X, Ji C, Han X et al (2006) Crystal structures of human glycerol-3-phosphate dehydrogenase 1 (GPD1). J Mol Biol 357:858–869
Peng F, Li G, Wang X et al (2010) Cloning and characterization of a glycerol-3-phosphate dehydrogenase (NAD+) gene from halotolerant yeast Pichia farinosa. Yeast 27:115–121
Rao AR, Dayananda C, Sarada R et al (2007) Effect of salinity on growth of green alga Botryococcus braunii and its constituents. Bioresour Technol 98:560–564
Riekhof WR, Sears BB, Benning C (2005) Annotation of genes involved in glycerolipid biosynthesis in Chlamydomonas reinhardtii: discovery of betaine lipid synthase BTA1Cr. Eukaryot Cell 4:242–252
Ronnow B, Kielland-Brandt MC (1993) GUT2, a gene for mitochondrial glycerol-3-phosphate dehydrogenase of Saccharomyces cerevisiae. Yeast 9:1121–1130
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Shen W, Wei Y, Dauk M et al (2006) Involvement of a glycerol-3-phosphate dehydrogenase in modulating the NADH/NAD+ ratio provides evidence of a mitochondrial glycerol-3-phosphate shuttle in Arabidopsis. Plant Cell 18:422–441
Siaut M, Cuiné S, Cagnon C et al (2011) Oil accumulation in the model green alga Chlamydomonas reinhardtii: characterization, variability between common laboratory strains and relationship with starch reserves. BMC Biotechnol 11:1–15
Sugase Y, Hirono M, Kindle K, Kamiya R (1996) Cloning and characterization of the actin-encoding gene of Chlamydomonas reinhardtii. Gene 168:117–121
Takagi M, Karseno, Yoshida T (2006) Effect of salt concentration on intracellular accumulation of lipids and triacylglyceride in marine microalgae Dunalliela cells. J Biosci Bioeng 101:223–226
Tamura K, Peterson D, Peterson N et al. (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol. doi:10.1093/molbev/msr121
Thomé P (2004) Isolation of a GPD gene from Debaryomyces hansenii encoding a glycerol-3-phosphate dehydrogenase (NAD+). Yeast 21:119–126
Vigeolas H, Geigenberger P (2004) Increased levels of glycerol-3-phosphate lead to a stimulation of flux into tryacylglycerol synthesis after supplying glycerol to developing seeds of Brassica napus L. in planta. Planta 219:827–835
Vigeolas H, Waldeck P, Zank T, Geigenberger P (2007) Increasing seed oil content in oil-seed rape (Brassica napus L.) by over-expression of a yeast glycerol-3-phosphate dehydrogenase under the control of a seed-specific promoter. Plant Biotechnol J 5:431–441
Watanabe Y, Tsuchimoto S, Tamai Y (2004) Heterologous expression of Zygosaccharomyces rouxii glycerol 3-phosphate dehydrogenase gene (ZrGPD1) and glycerol dehydrogenase gene (ZrGCY1) in Saccharomyces cerevisiae. FEMS Yeast Res 4:505–510
Yang ZH, Bielawski JP (2000) Statistical methods for detecting molecular adaptation. Trends Ecol Evol 15:496–503
Yang W, Cao Y, Sun X et al (2007) Isolation of a FAD-GPDH gene encoding a mitochondrial FAD-dependent glycerol-3-phosphate dehydrogenase from Dunaliella salina. J Basic Microbiol 47:266–274
Acknowledgments
This research was supported by the Consejo Nacional de Ciencia y Tecnología (CONACYT, México) Project No. CB-169217 and Centro de Investigación Científica de Yucatán (CICY, Mexico) Project FB0054. Laura Anahi Macario-González, Melissa Lessen Casais-Molina and Anayeli Beltrán Aguilar are grateful to Consejo Nacional de Ciencia y Tecnología (CONACYT, Mexico) for the Scholarship Nos. 235997, 228280 and 264772, respectively. Authors are grateful to Ileana C. Borges Argáez for technical support and Miguel Ángel Vallejo Reyna for assistance on the protein modeling.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Herrera-Valencia, V.A., Macario-González, L.A., Casais-Molina, M.L. et al. In Silico Cloning and Characterization of the Glycerol-3-Phosphate Dehydrogenase (GPDH) Gene Family in the Green Microalga Chlamydomonas reinhardtii . Curr Microbiol 64, 477–485 (2012). https://doi.org/10.1007/s00284-012-0095-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-012-0095-6