Abstract
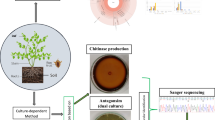
A rapid molecular identification technique was applied on microbial microflora isolated from Brazilian cassava roots given a yeast profile presented in the samples analyzed. A total of 24 strain isolated from cassava were initially grouped and identified in five groups using restriction-fragment length polymorphism (RFLPs) of 5.8S-ITS rDNA region. Sequencing analysis of the domains D1 and D2 of the 26S rRNA gene or 5.8S rRNA-ITS region were used to identify different groups of yeasts. Representative colonies of yeasts of each group were isolated and identified as Debaromyces hansenii, Kodamaea ohmeri, Candida glabrata, C. haemulonii, and Pichia gullhermondii. It is hoped that these results will contribute toward selecting yeast from this microflora capable to degrade cassava starch in the near future.
Similar content being viewed by others
References
Achi OK, Akubor PI (2000) Microbiological characterization of yam fermentation for ‘elubo’ (yam flour) production. World J Microb Biot 16:3–7
Alves AAC (2002) Cassava botany and physiology. In: Hillocks RJ, Thresh JM, Bellotti AC (eds) Cassava: biology, production and utilization. CABI Publishing, New York
Dixon AGO, Nukenine EN (2000) Genotype X environment interaction and optimum resource allocation for yield and yield components of cassava. Afr Crop Sci J 8(1):1–10
El-Sharkawy MA, Cock JH (1987) Response of cassava to water stress. Plant Soil 100:345–360
Esteve-Zarzoso B, Belloch C, Uruburu F, Querol A (1999) Identification of yeast by RFPL analysis of the 5.8S rRNA gene and the two ribosomal internal transcribed spacers. Int J Syst Bacteriol 49:329–337
Fredricks DN, Smith C, Meier A (2005) Comparison of six DNA extraction methods for recovery of fungal DNA as assessed by quantitative PCR. J Clin Microbiol 43(10):5122–5128
Kurtzman CP, Robnett CJ (1997) Identification of clinically important ascomycetous yeasts based on nucleotide divergence in the 5′ end of the large-subunit (26 s) ribosomal DNA gene. J Clin Microbiol 35(5):1216–1223
Kurtzman CP, Robnett CJ (1998) Identification and phylogeny of ascomycetous yeasts from analysis of nuclear subunit (26S) ribosomal DNA sequences. Anton van Leeuw 73:331–371
Leaw SN, Chang HC, Sun HF, Barton R, Bouchara JP, Chang CC (2006) Identification of medically important yeast species by sequence analysis of the internal transcribed spacer regions. J Clin Microbiol 44(3):693–699
Liebmann B, Marengo JA (2001) Interannual variability of the rainy season and rainfall in the Brazilian Amazon Basin. J Climate 14(22):4308–4318
Nogueira CAM, Momesso CAS, Machado RLD, Almeida MTG, Rossit ARB (2004) Performance of commercial kits and laboratory protocols for bacterial genomic DNA extraction. Rev Panam Infectol 6(2):35–38
Nout MJR, Essers AJA (1989) The safety of dark, moulded cassava flour compared with white: a comparison of traditionally dried cassava pieces in North East Mozambique. Trop Sci 29:261–268
Oates CG (1997) Towards an understanding of starch granule structure and hydrolysis. Trends Food Sci Technol 8:375–382
Oguntoyinbo FA (2008) Evaluation of diversity of Candida species isolated from fermented cassava during traditional small scale gari production in Nigeria. Food Control 19:465–469
Querol A, Barrio E, Huerta T, Ramón D (1992) Molecular monitoring of wine conducted by active dry yeast strains. Appl Environ Microb 58(9):2948–2953
Sarikaya E, Higasa T, Adachi M, Mikami B (2000) Comparison of degradation abilities of a- and b-amylases on raw starch granules. Process Biochem 35:711–715
Smit E, Leeflang P, Glandorf B, Elsas JDV, Wernars K (1999) Analysis of fungal diversity in the wheat rhizosphere by sequencing of cloned PCR-amplified genes encoding 18S rRNA and temperature gradient gel electrophoresis. Appl Environ Microb 65(6):2614–2621
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tonukari NJ (2004) Cassava and the future of starch. Electron J Biotechnol 7(1):5–8
Wydra K, Verdier V (2002) Occurrence of cassava diseases in relation to environmental, agronomic and plant characteristics. Agr Ecosyst Environ 93(1–3):211–226
Yamada Y, Suzuki T, Matsuda M, Mikata K (1995) The phylogeny of Yamadazyma ohmeri (Etchells et Bell) Billon-Grand based on the partial sequences of 18S and 26S ribosomal RNAs: the proposal of Kodamaea Gen. Nov. (Saccharomycetaceae). Biosci Biotech Biochem 59(6):1172–1174
Acknowledgment
This work was supported by CSIC/IATA and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Ferreira, N., Belloch, C., Querol, A. et al. Yeast Microflora Isolated From Brazilian Cassava Roots: Taxonomical Classification Based on Molecular Identification. Curr Microbiol 60, 287–293 (2010). https://doi.org/10.1007/s00284-009-9539-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-009-9539-z