Abstract
Purpose
Tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) binds to death receptor (DR) 4 and DR5 and induces tumor-selective apoptosis. The fusion proteins NCTR25–TRAIL and NCTR25–TGF3L–TRAIL self-assembled into polymers and triggered super-active cancer cell killing. The role of TGF3L in self-assembly and super-activity was unclear. These multivalent TRAILs elicited apoptosis with great potency, but their specificity towards receptors and subsequent efficacy in signal activation were unclear.
Methods
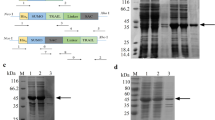
NCTR25–TRAIL fusion was constructed and prokaryotically expressed. The size of fusion protein polymers was estimated. Their cytotoxicity was assessed in eight cancer cell lines and two noncancerous cell lines. Receptor binding and activation specificity were determined by antibody blockade. Apoptosis was evaluated, and the associated pathway was verified by quantifying caspase activity. The NF-κB signaling pathway was assessed by dual-luciferase assay. The in vivo antitumor activity was also evaluated in nude mice.
Results
NCTR25 fusion to TRAIL promoted its self-assembly into polymers and showed similar super-cytotoxicity to NCTR25–TGF3L–TRAIL in vitro. The multivalent TRAILs exclusively activated both DR4 and DR5 and showed a bias towards DR4 in mediating cytotoxicity in NCI-H460 cells. They activated caspase pathway and induced apoptosis with higher potency but in similar efficacy than TRAIL. A higher potency and a greater efficacy were observed in activating NF-κB pathway by NCTR25–TRAIL comparing to TRAIL. Both the polymers showed better in vivo antitumor activity than TRAIL.
Conclusions
NCTR25 fusion alone facilitates the formation of TRAIL polymers. Multivalent TRAIL polymers bind and activate DR4 and DR5 specifically and exclusively, triggering the signaling pathways with higher potency, and greater efficacy than TRAIL.









Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information file. The data are also available from the corresponding author on reasonable request.
References
Pitti RM, Marsters SA, Ruppert S, Donahue CJ, Moore A, Ashkenazi A (1996) Induction of apoptosis by Apo-2 ligand, a new member of the tumor necrosis factor cytokine family. J Biol Chem 271(22):12687–12690
Ashkenazi A, Dixit VM (1998) Death receptors: signaling and modulation. Science 281(5381):1305–1308
Sprick MR, Weigand MA, Rieser E, Rauch CT, Juo P, Blenis J, Krammer PH, Walczak H (2000) FADD/MORT1 and caspase-8 are recruited to TRAIL receptors 1 and 2 and are essential for apoptosis mediated by TRAIL receptor 2. Immunity 12(6):599–609
Degli-Esposti MA, Dougall WC, Smolak PJ, Waugh JY, Smith CA, Goodwin RG (1997) The novel receptor TRAIL-R4 induces NF-κB and protects against TRAIL-mediated apoptosis, yet retains an incomplete death domain. Immunity 7(6):813–820. https://doi.org/10.1016/S1074-7613(00)80399-4
Degli-Esposti MA, Smolak PJ, Walczak H, Waugh J, Huang CP, DuBose RF, Goodwin RG, Smith CA (1997) Cloning and characterization of TRAIL-R3, a novel member of the emerging TRAIL receptor family. J Exp Med 186(7):1165–1170
Marsters SA, Sheridan JP, Pitti RM, Huang A, Skubatch M, Baldwin D, Yuan J, Gurney A, Goddard AD, Godowski P, Ashkenazi A (1997) A novel receptor for Apo2L/TRAIL contains a truncated death domain. Curr Biol 7(12):1003–1006
Emery JG, McDonnell P, Burke MB, Deen KC, Lyn S, Silverman C, Dul E, Appelbaum ER, Eichman C, DiPrinzio R, Dodds RA, James IE, Rosenberg M, Lee JC, Young PR (1998) Osteoprotegerin is a receptor for the cytotoxic ligand TRAIL. J Biol Chem 273(23):14363–14367
Clancy L, Mruk K, Archer K, Woelfel M, Mongkolsapaya J, Screaton G, Lenardo MJ, Chan FK (2005) Preligand assembly domain-mediated ligand-independent association between TRAIL receptor 4 (TR4) and TR2 regulates TRAIL-induced apoptosis. Proc Natl Acad Sci U S A 102(50):18099–18104. https://doi.org/10.1073/pnas.0507329102
Neumann S, Hasenauer J, Pollak N, Scheurich P (2014) Dominant negative effects of tumor necrosis factor (TNF)-related apoptosis-inducing ligand (TRAIL) receptor 4 on TRAIL receptor 1 signaling by formation of heteromeric complexes. J Biol Chem 289(23):16576–16587. https://doi.org/10.1074/jbc.M114.559468
Smulski CR, Decossas M, Chekkat N, Beyrath J, Willen L, Guichard G, Lorenzetti R, Rizzi M, Eibel H, Schneider P, Fournel S (2017) Hetero-oligomerization between the TNF receptor superfamily members CD40, Fas and TRAILR2 modulate CD40 signalling. Cell Death Dis 8(2):e2601. https://doi.org/10.1038/cddis.2017.22
Vanamee ES, Faustman DL (2018) Structural principles of tumor necrosis factor superfamily signaling. Sci Signal. https://doi.org/10.1126/scisignal.aao4910
Wilson NS, Yang B, Yang A, Loeser S, Marsters S, Lawrence D, Li Y, Pitti R, Totpal K, Yee S, Ross S, Vernes JM, Lu Y, Adams C, Offringa R, Kelley B, Hymowitz S, Daniel D, Meng G, Ashkenazi A (2011) An Fcgamma receptor-dependent mechanism drives antibody-mediated target-receptor signaling in cancer cells. Cancer Cell 19(1):101–113. https://doi.org/10.1016/j.ccr.2010.11.012
Muhlenbeck F, Schneider P, Bodmer JL, Schwenzer R, Hauser A, Schubert G, Scheurich P, Moosmayer D, Tschopp J, Wajant H (2000) The tumor necrosis factor-related apoptosis-inducing ligand receptors TRAIL-R1 and TRAIL-R2 have distinct cross-linking requirements for initiation of apoptosis and are non-redundant in JNK activation. J Biol Chem 275(41):32208–32213. https://doi.org/10.1074/jbc.M000482200
Graves JD, Kordich JJ, Huang TH, Piasecki J, Bush TL, Sullivan T, Foltz IN, Chang W, Douangpanya H, Dang T, O’Neill JW, Mallari R, Zhao X, Branstetter DG, Rossi JM, Long AM, Huang X, Holland PM (2014) Apo2L/TRAIL and the death receptor 5 agonist antibody AMG 655 cooperate to promote receptor clustering and antitumor activity. Cancer Cell 26(2):177–189. https://doi.org/10.1016/j.ccr.2014.04.028
Gieffers C, Kluge M, Merz C, Sykora J, Thiemann M, Schaal R, Fischer C, Branschadel M, Abhari BA, Hohenberger P, Fulda S, Fricke H, Hill O (2013) APG350 induces superior clustering of TRAIL receptors and shows therapeutic antitumor efficacy independent of cross-linking via Fcgamma receptors. Mol Cancer Ther 12(12):2735–2747. https://doi.org/10.1158/1535-7163.MCT-13-0323
De Miguel D, Gallego-Lleyda A, Ayuso JM, Pejenaute-Ochoa D, Jarauta V, Marzo I, Fernandez LJ, Ochoa I, Conde B, Anel A, Martinez-Lostao L (2016) High-order TRAIL oligomer formation in TRAIL-coated lipid nanoparticles enhances DR5 cross-linking and increases antitumour effect against colon cancer. Cancer Lett 383(2):250–260. https://doi.org/10.1016/j.canlet.2016.10.005
Wang Y, Lei Q, Yan Z, Shen C, Wang N (2018) TGF3L fusion enhances the antitumor activity of TRAIL by promoting assembly into polymers. Biochem Pharmacol 155:510–523. https://doi.org/10.1016/j.bcp.2018.07.035
Nestor JJ Jr, Newman SR, DeLustro B, Todaro GJ, Schreiber AB (1985) A synthetic fragment of rat transforming growth factor alpha with receptor binding and antigenic properties. Biochem Biophys Res Commun 129(1):226–232
Eppstein DA, Marsh YV, Schryver BB, Bertics PJ (1989) Inhibition of epidermal growth factor/transforming growth factor-alpha-stimulated cell growth by a synthetic peptide. J Cell Physiol 141(2):420–430. https://doi.org/10.1002/jcp.1041410224
Kih M, Lee EJ, Lee NK, Kim YK, Lee KE, Jeong C, Yang Y, Kim DH, Kim IS (2018) Designed trimer-mimetic TNF superfamily ligands on self-assembling nanocages. Biomaterials 180:67–77. https://doi.org/10.1016/j.biomaterials.2018.07.009
Liu H, Su D, Zhang J, Ge S, Li Y, Wang F, Gravel M, Roulston A, Song Q, Xu W, Liang JG, Shore G, Wang X, Liang P (2017) Improvement of Pharmacokinetic Profile of TRAIL via Trimer-Tag Enhances its Antitumor Activity in vivo. Sci Rep 7(1):8953. https://doi.org/10.1038/s41598-017-09518-1
Nussinov R, Ma B, Tsai CJ (2014) Multiple conformational selection and induced fit events take place in allosteric propagation. Biophys Chem 186:22–30. https://doi.org/10.1016/j.bpc.2013.10.002
Pan W, Wang Y, Wang N (2019) A new metal affinity NCTR25 tag as a better alternative to the His-tag for the expression of recombinant fused proteins. Protein Expr Purif 164:105477. https://doi.org/10.1016/j.pep.2019.105477
Sambrook J, Fritsch EF, Maniatis T (2001) Molecular cloning: a laboratory manual. CSH
Chennamsetty N, Voynov V, Kayser V, Helk B, Trout BL (2009) Design of therapeutic proteins with enhanced stability. Proc Natl Acad Sci U S A 106(29):11937–11942. https://doi.org/10.1073/pnas.0904191106
Mosmann T (1983) Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays. J Immunol Methods 65(1–2):55–63
Anoopkumar-Dukie S, Carey JB, Conere T, O’Sullivan E, van Pelt FN, Allshire A (2005) Resazurin assay of radiation response in cultured cells. Br J Radiol 78(934):945–947. https://doi.org/10.1259/bjr/54004230
Lazareno S, Birdsall NJ (1993) Estimation of competitive antagonist affinity from functional inhibition curves using the Gaddum, Schild and Cheng-Prusoff equations. Br J Pharmacol 109(4):1110–1119. https://doi.org/10.1111/j.1476-5381.1993.tb13737.x
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227(5259):680–685
Long J, Zhao J, Yan Z, Liu Z, Wang N (2009) Antitumor effects of a novel sulfur-containing hydroxamate histone deacetylase inhibitor H40. Int J Cancer 124(5):1235–1244. https://doi.org/10.1002/ijc.24074
Crowley LC, Marfell BJ, Scott AP (2016) Waterhouse NJ (2016) Quantitation of Apoptosis and Necrosis by Annexin V Binding, Propidium Iodide Uptake, and Flow Cytometry. Cold Spring Harb Protoc 11:pdbprot087288. https://doi.org/10.1101/pdb.prot087288
Kajstura M, Halicka HD, Pryjma J, Darzynkiewicz Z (2007) Discontinuous fragmentation of nuclear DNA during apoptosis revealed by discrete “sub-G1” peaks on DNA content histograms. Cytometry A 71(3):125–131. https://doi.org/10.1002/cyto.a.20357
De Feo CJ, Aller SG, Siluvai GS, Blackburn NJ, Unger VM (2009) Three-dimensional structure of the human copper transporter hCTR1. J Proc Natl Acad Sci 106(11):4237–4242. https://doi.org/10.1073/pnas.0810286106
Wagner KW, Punnoose EA, Januario T, Lawrence DA, Pitti RM, Lancaster K, Lee D, von Goetz M, Yee SF, Totpal K, Huw L, Katta V, Cavet G, Hymowitz SG, Amler L, Ashkenazi A (2007) Death-receptor O-glycosylation controls tumor-cell sensitivity to the proapoptotic ligand Apo2L/TRAIL. Nat Med 13(9):1070–1077. https://doi.org/10.1038/nm1627
Jeon YJ, Middleton J, Kim T, Lagana A, Piovan C, Secchiero P, Nuovo GJ, Cui R, Joshi P, Romano G, Di Leva G, Lee BK, Sun HL, Kim Y, Fadda P, Alder H, Garofalo M, Croce CM (2017) Correction for Jeon et al., A set of NF-kappaB-regulated microRNAs induces acquired TRAIL resistance in Lung cancer. Proc Natl Acad Sci USA 114(12):2542. https://doi.org/10.1073/pnas.1701795114
Hu WH, Johnson H, Shu HB (1999) Tumor necrosis factor-related apoptosis-inducing ligand receptors signal NF-kappaB and JNK activation and apoptosis through distinct pathways. J Biol Chem 274(43):30603–30610
Varewijck AJ, Janssen JA (2012) Insulin and its analogues and their affinities for the IGF1 receptor. Endocr Relat Cancer 19(5):F63-75. https://doi.org/10.1530/erc-12-0026
De Miguel D, Gallego-Lleyda A, Anel A, Martinez-Lostao L (2015) Liposome-bound TRAIL induces superior DR5 clustering and enhanced DISC recruitment in histiocytic lymphoma U937 cells. Leuk Res 39(6):657–666. https://doi.org/10.1016/j.leukres.2015.03.019
Wajant H, Moosmayer D, Wuest T, Bartke T, Gerlach E, Schonherr U, Peters N, Scheurich P, Pfizenmaier K (2001) Differential activation of TRAIL-R1 and -2 by soluble and membrane TRAIL allows selective surface antigen-directed activation of TRAIL-R2 by a soluble TRAIL derivative. Oncogene 20(30):4101–4106. https://doi.org/10.1038/sj.onc.1204558
Carlo-Stella C, Lavazza C, Di Nicola M, Cleris L, Longoni P, Milanesi M, Magni M, Morelli D, Gloghini A, Carbone A, Gianni AM (2006) Antitumor activity of human CD34+ cells expressing membrane-bound tumor necrosis factor-related apoptosis-inducing ligand. Hum Gene Ther 17(12):1225–1240. https://doi.org/10.1089/hum.2006.17.1225
Dufour F, Rattier T, Constantinescu AA, Zischler L, Morle A, Ben Mabrouk H, Humblin E, Jacquemin G, Szegezdi E, Delacote F, Marrakchi N, Guichard G, Pellat-Deceunynck C, Vacher P, Legembre P, Garrido C, Micheau O (2017) TRAIL receptor gene editing unveils TRAIL-R1 as a master player of apoptosis induced by TRAIL and ER stress. Oncotarget 8(6):9974–9985. https://doi.org/10.18632/oncotarget.14285
Lawrence D, Shahrokh Z, Marsters S, Achilles K, Shih D, Mounho B, Hillan K, Totpal K, DeForge L, Schow P, Hooley J, Sherwood S, Pai R, Leung S, Khan L, Gliniak B, Bussiere J, Smith CA, Strom SS, Kelley S, Fox JA, Thomas D, Ashkenazi A (2001) Differential hepatocyte toxicity of recombinant Apo2L/TRAIL versions. Nat Med 7(4):383–385. https://doi.org/10.1038/86397
Acknowledgements
This work was supported by a grant from the Beijing Municipal Natural Science Foundation (Grant No. 7162134) and the Drug Innovation Major Project (No. 2013HXW-20, 2018ZX09711001-003-001).
Author information
Authors and Affiliations
Contributions
Conceptualization, NW; data curation, YW; formal analysis, YW; funding acquisition, NW; investigation, YW, QL, and CS; methodology, YW and NW; project administration, YW and NW; supervision, NW; validation, YW, QL, and CS; visualization, YW; writing—original draft, YW; writing—review & editing, YW and NW.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, Y., Lei, Q., Shen, C. et al. NCTR25 fusion facilitates the formation of TRAIL polymers that selectively activate TRAIL receptors with higher potency and efficacy than TRAIL. Cancer Chemother Pharmacol 88, 289–306 (2021). https://doi.org/10.1007/s00280-021-04283-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00280-021-04283-5