Abstract
Background
The involvement of ferroptosis in the pathogenesis and progression of various cancers has been well established. However, limited studies have investigated the role of ferroptosis-mediated tumor microenvironment (TME) in skin cutaneous melanoma (SKCM).
Methods
By leveraging single-cell RNA sequencing data, the nonnegative matrix factorization (NMF) approach was employed to comprehensively characterize and identify distinct gene signatures within ferroptosis-associated TME cell clusters. Prognostic and treatment response analyses were conducted using both bulk datasets and external cancer cohort to evaluate the clinical implications of TME clusters.
Results
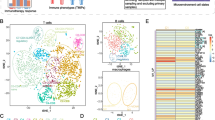
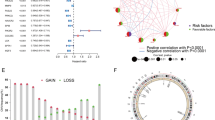
This NMF-based analysis successfully delineated fibroblasts, macrophages, T cells, and B cells into multiple clusters, enabling the identification of unique gene expression patterns and the annotation of distinct TME clusters. Furthermore, pseudotime trajectories, enrichment analysis, cellular communication analysis, and gene regulatory network analysis collectively demonstrated significant intercellular communication between key TME cell clusters, thereby influencing tumor cell development through diverse mechanisms. Importantly, our bulk RNA-seq analysis revealed the prognostic significance of ferroptosis-mediated TME cell clusters in SKCM patients. Moreover, our analysis of immune checkpoint blockade highlighted the crucial role of TME cell clusters in tumor immunotherapy, facilitating the discovery of potential immunotherapeutic targets.
Conclusions
In conclusion, this pioneering study employing NMF-based analysis unravels the intricate cellular communication mediated by ferroptosis within the TME and its profound implications for the pathogenesis and progression of SKCM. We provide compelling evidence for the prognostic value of ferroptosis-regulated TME cell clusters in SKCM, as well as their potential as targets for immunotherapy.








Similar content being viewed by others
Data availability
Publicly available datasets were analyzed in this study. These data can be found here: The datasets analyzed in the current study are available in the GEO and TCGA database. All raw data and original images can be found in the jianguoyun (https://www.jianguoyun.com/p/DYLfxGkQjpbCCxit5JAFIAA).
References
Garbe C, Amaral T, Peris K, Hauschild A, Arenberger P, Basset-Seguin N, Bastholt L, Bataille V, Del Marmol V, Dréno B et al (2022) European consensus-based interdisciplinary guideline for melanoma. Part 1: diagnostics update 2022. Eur J Cancer. 170:236–255. https://doi.org/10.1016/j.ejca.2022.03.008
Arnold M, Singh D, Laversanne M, Vignat J, Vaccarella S, Meheus F, Cust AE, de Vries E, Whiteman DC, Bray F (2022) Global burden of cutaneous melanoma in 2020 and projections to 2040. JAMA Dermatol. https://doi.org/10.1001/jamadermatol.2022.0160
Hassannia B, Vandenabeele P, Vanden BT (2019) Targeting ferroptosis to iron out cancer. Cancer Cell 35:830–849. https://doi.org/10.1016/j.ccell.2019.04.002
Koppula P, Zhuang L, Gan B (2021) Cystine transporter SLC7A11/xCT in cancer: ferroptosis, nutrient dependency, and cancer therapy. Protein Cell 12:599–620. https://doi.org/10.1007/s13238-020-00789-5
Zhang Y, Shi J, Liu X, Feng L, Gong Z, Koppula P, Sirohi K, Li X, Wei Y, Lee H et al (2018) BAP1 links metabolic regulation of ferroptosis to tumour suppression. Nat Cell Biol 20:1181–1192. https://doi.org/10.1038/s41556-018-0178-0
Jiang L, Kon N, Li T, Wang S-J, Su T, Hibshoosh H, Baer R, Gu W (2015) Ferroptosis as a p53-mediated activity during tumour suppression. Nature 520:57–62. https://doi.org/10.1038/nature14344
Talty R, Bosenberg M (2022) The role of ferroptosis in melanoma. Pigment Cell Melanoma Res 35:18–25. https://doi.org/10.1111/pcmr.13009
Liu C, Liu Y, Yu Y, Zhao Y, Yu A (2022) Comprehensive analysis of ferroptosis-related genes and prognosis of cutaneous melanoma. BMC Med Genomics 15:39. https://doi.org/10.1186/s12920-022-01194-z
Ping S, Wang S, Zhao Y, He J, Li G, Li D, Wei Z, Chen J (2022) Identification and validation of a ferroptosis-related gene signature for predicting survival in skin cutaneous melanoma. Cancer Med 11:3529–3541. https://doi.org/10.1002/cam4.4706
Duro-Sánchez S, Alonso MR, Arribas J (2023) Immunotherapies against HER2-positive breast cancer. Cancers (Basel) 15:1069. https://doi.org/10.3390/cancers15041069
Wang D, Han Y, Peng L, Huang T, He X, Wang J, Ou C (2023) Crosstalk between N6-methyladenosine (m6A) modification and noncoding RNA in tumor microenvironment. Int J Biol Sci 19:2198–2219. https://doi.org/10.7150/ijbs.79651
Biffi G, Tuveson DA (2021) Diversity and biology of cancer-associated fibroblasts. Physiol Rev 101:147–176. https://doi.org/10.1152/physrev.00048.2019
Xia Y, Rao L, Yao H, Wang Z, Ning P, Chen X (2020) Engineering macrophages for cancer immunotherapy and drug delivery. Adv Mater. https://doi.org/10.1002/adma.202002054
Oh DY, Fong L (2021) Cytotoxic CD4+ T cells in cancer: expanding the immune effector toolbox. Immunity 54:2701–2711. https://doi.org/10.1016/j.immuni.2021.11.015
Yazdanifar M, Barbarito G, Bertaina A, Airoldi I (2020) γδ T cells: the ideal tool for cancer immunotherapy. Cells 9:1305. https://doi.org/10.3390/cells9051305
Engelhard V, Conejo-Garcia JR, Ahmed R, Nelson BH, Willard-Gallo K, Bruno TC, Fridman WH (2021) B cells and cancer. Cancer Cell 39:1293–1296. https://doi.org/10.1016/j.ccell.2021.09.007
Michaud D, Steward CR, Mirlekar B, Pylayeva-Gupta Y (2021) Regulatory B cells in cancer. Immunol Rev 299:74–92. https://doi.org/10.1111/imr.12939
Mariathasan S, Turley SJ, Nickles D, Castiglioni A, Yuen K, Wang Y, Kadel EE, Koeppen H, Astarita JL, Cubas R et al (2018) TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554:544–548. https://doi.org/10.1038/nature25501
Galbo PM, Zang X, Zheng D (2021) Molecular features of cancer-associated fibroblast subtypes and their implication on cancer pathogenesis, prognosis, and immunotherapy resistance. Clin Cancer Res 27:2636–2647. https://doi.org/10.1158/1078-0432.CCR-20-4226
Zeng H, You C, Zhao L, Wang J, Ye X, Yang T, Wan C, Deng L (2021) Ferroptosis-associated classifier and indicator for prognostic prediction in cutaneous melanoma. J Oncol 2021:3658196. https://doi.org/10.1155/2021/3658196
Liu Y, Shou Y, Zhu R, Qiu Z, Zhang Q, Xu J (2022) Construction and validation of a ferroptosis-related prognostic signature for melanoma based on single-cell RNA sequencing. Front Cell Dev Biol. 10:818457. https://doi.org/10.3389/fcell.2022.818457
Chen Y, Zhu S, Liu T, Zhang S, Lu J, Fan W, Lin L, Xiang T, Yang J, Zhao X et al (2023) Epithelial cells activate fibroblasts to promote esophageal cancer development. Cancer Cell 41:903-918.e8. https://doi.org/10.1016/j.ccell.2023.03.001
Wei C, Yang C, Wang S, Shi D, Zhang C, Lin X, Liu Q, Dou R, Xiong B (2019) Crosstalk between cancer cells and tumor associated macrophages is required for mesenchymal circulating tumor cell-mediated colorectal cancer metastasis. Mol Cancer 18:64. https://doi.org/10.1186/s12943-019-0976-4
de Waal AM, Hiemstra PS, Ottenhoff TH, Joosten SA, van der Does AM (2022) Lung epithelial cells interact with immune cells and bacteria to shape the microenvironment in tuberculosis. Thorax 77:408–416. https://doi.org/10.1136/thoraxjnl-2021-217997
Angelopoulou E, Piperi C (2018) Emerging role of plexins signaling in glioma progression and therapy. Cancer Lett 414:81–87. https://doi.org/10.1016/j.canlet.2017.11.010
Toledano S, Nir-Zvi I, Engelman R, Kessler O, Neufeld G (2019) Class-3 semaphorins and their receptors: potent multifunctional modulators of tumor progression. Int J Mol Sci 20:556. https://doi.org/10.3390/ijms20030556
Tamagnone L (2012) Emerging role of semaphorins as major regulatory signals and potential therapeutic targets in cancer. Cancer Cell 22:145–152. https://doi.org/10.1016/j.ccr.2012.06.031
Worzfeld T, Offermanns S (2014) Semaphorins and plexins as therapeutic targets. Nat Rev Drug Discov 13:603–621. https://doi.org/10.1038/nrd4337
Zhang X, Klamer B, Li J, Fernandez S, Li L (2020) A pan-cancer study of class-3 semaphorins as therapeutic targets in cancer. BMC Med Genomics 13:45. https://doi.org/10.1186/s12920-020-0682-5
Nwabo Kamdje AH, Seke Etet PF, Kipanyula MJ, Vecchio L, Tagne Simo R, Njamnshi AK, Lukong KE, Mimche PN (2022) Insulin-like growth factor-1 signaling in the tumor microenvironment: carcinogenesis, cancer drug resistance, and therapeutic potential. Front Endocrinol (Lausanne). 13:927390. https://doi.org/10.3389/fendo.2022.927390
Fettig LM, Yee D (2020) Advances in insulin-like growth factor biology and -directed cancer therapeutics. Adv Cancer Res 147:229–257. https://doi.org/10.1016/bs.acr.2020.04.005
Parsons MJ, Tammela T, Dow LE (2021) WNT as a driver and dependency in cancer. Cancer Discov 11:2413–2429. https://doi.org/10.1158/2159-8290.CD-21-0190
Jian S, Luo D, Wang Y, Xu W, Zhang H, Zhang L, Zhou X (2021) MiR-337-3p confers protective effect on facet joint osteoarthritis by targeting SKP2 to inhibit DUSP1 ubiquitination and inactivate MAPK pathway. Cell Biol Toxicol. https://doi.org/10.1007/s10565-021-09665-2
Shen J, Xing W, Liu R, Zhang Y, Xie C, Gong F (2019) MiR-32-5p influences high glucose-induced cardiac fibroblast proliferation and phenotypic alteration by inhibiting DUSP1. BMC Mol Biol 20:21. https://doi.org/10.1186/s12867-019-0135-x
Li L, Yang M, Yu J, Cheng S, Ahmad M, Wu C, Wan X, Xu B, Ben-David Y, Luo H (2022) A novel L-phenylalanine dipeptide inhibits the growth and metastasis of prostate cancer cells via targeting DUSP1 and TNFSF9. Int J Mol Sci 23:10916. https://doi.org/10.3390/ijms231810916
Pan W, Han J, Wei N, Wu H, Wang Y, Sun J (2022) LINC00702-mediated DUSP1 transcription in the prevention of bladder cancer progression: Implications in cancer cell proliferation and tumor inflammatory microenvironment. Genomics. 114:110428. https://doi.org/10.1016/j.ygeno.2022.110428
Huang C, Zhan L (2022) Network pharmacology identifies therapeutic targets and the mechanisms of glutathione action in ferroptosis occurring in oral cancer. Front Pharmacol. 13:851540. https://doi.org/10.3389/fphar.2022.851540
Deng H, Lin Y, Gan F, Li B, Mou Z, Qin X, He X, Meng Y (2022) Prognostic model and immune infiltration of ferroptosis subcluster-related modular genes in gastric cancer. J Oncol 2022:5813522. https://doi.org/10.1155/2022/5813522
Chen X, Yu C, Kang R, Kroemer G, Tang D (2021) Cellular degradation systems in ferroptosis. Cell Death Differ 28:1135–1148. https://doi.org/10.1038/s41418-020-00728-1
Jin Q, Li R, Hu N, Xin T, Zhu P, Hu S, Ma S, Zhu H, Ren J, Zhou H (2018) DUSP1 alleviates cardiac ischemia/reperfusion injury by suppressing the Mff-required mitochondrial fission and Bnip3-related mitophagy via the JNK pathways. Redox Biol 14:576–587. https://doi.org/10.1016/j.redox.2017.11.004
Ji J, Liu Z, Hong X, Liu Z, Gao J, Liu J (2020) Protective effects of rolipram on endotoxic cardiac dysfunction via inhibition of the inflammatory response in cardiac fibroblasts. BMC Cardiovasc Disord 20:242. https://doi.org/10.1186/s12872-020-01529-7
Blumer S, Fang L, Chen W-C, Khan P, Hostettler K, Tamm M, Roth M, Lambers C (2021) IPF-fibroblast Erk1/2 activity is independent from microRNA cluster 17–92 but can be inhibited by treprostinil through DUSP1. Cells 10:2836. https://doi.org/10.3390/cells10112836
Peng H-Z, Yun Z, Wang W, Ma B-A (2017) Dual specificity phosphatase 1 has a protective role in osteoarthritis fibroblast-like synoviocytes via inhibition of the MAPK signaling pathway. Mol Med Rep 16:8441–8447. https://doi.org/10.3892/mmr.2017.7617
Ramkissoon A, Chaney KE, Milewski D, Williams KB, Williams RL, Choi K, Miller A, Kalin TV, Pressey JG, Szabo S et al (2019) Targeted inhibition of the dual specificity phosphatases DUSP1 and DUSP6 suppress MPNST growth via JNK. Clin Cancer Res 25:4117–4127. https://doi.org/10.1158/1078-0432.CCR-18-3224
Zhang L, Gao S, Shi X, Chen Y, Wei S, Mi Y, Zuo L, Qi C (2023) NUPR1 imparts oncogenic potential in bladder cancer. Cancer Med 12:7149–7163. https://doi.org/10.1002/cam4.5518
Fan T, Wang X, Zhang S, Deng P, Jiang Y, Liang Y, Jie S, Wang Q, Li C, Tian G et al (2022) NUPR1 promotes the proliferation and metastasis of oral squamous cell carcinoma cells by activating TFE3-dependent autophagy. Signal Transduct Target Ther 7:130. https://doi.org/10.1038/s41392-022-00939-7
Jiang L, Wang W, Li Z, Zhao Y, Qin Z (2021) NUPR1 participates in YAP-mediate gastric cancer malignancy and drug resistance via AKT and p21 activation. J Pharm Pharmacol 73:740–748. https://doi.org/10.1093/jpp/rgab010
Dai Q, Wu W, Amei A, Yan X, Lu L, Wang Z (2021) Regulation and characterization of tumor-infiltrating immune cells in breast cancer. Int Immunopharmacol. 90:107167. https://doi.org/10.1016/j.intimp.2020.107167
Liao P, Wang W, Wang W, Kryczek I, Li X, Bian Y, Sell A, Wei S, Grove S, Johnson JK et al (2022) CD8+ T cells and fatty acids orchestrate tumor ferroptosis and immunity via ACSL4. Cancer Cell 40:365-378.e6. https://doi.org/10.1016/j.ccell.2022.02.003
Wang W, Green M, Choi JE, Gijón M, Kennedy PD, Johnson JK, Liao P, Lang X, Kryczek I, Sell A et al (2019) CD8+ T cells regulate tumour ferroptosis during cancer immunotherapy. Nature 569:270–274. https://doi.org/10.1038/s41586-019-1170-y
Dolina JS, Van Braeckel-Budimir N, Thomas GD, Salek-Ardakani S (2021) CD8+ T cell exhaustion in cancer. Front Immunol. 12:715234. https://doi.org/10.3389/fimmu.2021.715234
Raskov H, Orhan A, Christensen JP, Gögenur I (2021) Cytotoxic CD8+ T cells in cancer and cancer immunotherapy. Br J Cancer 124:359–367. https://doi.org/10.1038/s41416-020-01048-4
Kanai Y (2022) Amino acid transporter LAT1 (SLC7A5) as a molecular target for cancer diagnosis and therapeutics. Pharmacol Ther. 230:107964. https://doi.org/10.1016/j.pharmthera.2021.107964
Wang L, Liu Y, Du T, Yang H, Lei L, Guo M, Ding H-F, Zhang J, Wang H, Chen X et al (2020) ATF3 promotes erastin-induced ferroptosis by suppressing system Xc. Cell Death Differ 27:662–675. https://doi.org/10.1038/s41418-019-0380-z
Ding Y, Chen X, Liu C, Ge W, Wang Q, Hao X, Wang M, Chen Y, Zhang Q (2021) Identification of a small molecule as inducer of ferroptosis and apoptosis through ubiquitination of GPX4 in triple negative breast cancer cells. J Hematol Oncol 14:19. https://doi.org/10.1186/s13045-020-01016-8
Liu Z, Xu Y, Liu X, Wang B (2022) PCDH7 knockdown potentiates colon cancer cells to chemotherapy via inducing ferroptosis and changes in autophagy through restraining MEK1/2/ERK/c-Fos axis. Biochem Cell Biol 100:445–457. https://doi.org/10.1139/bcb-2021-0513
Ishikawa C, Senba M, Mori N (2020) Evaluation of artesunate for the treatment of adult T-cell leukemia/lymphoma. Eur J Pharmacol. 872:172953. https://doi.org/10.1016/j.ejphar.2020.172953
Wei T-T, Zhang M-Y, Zheng X-H, Xie T-H, Wang W, Zou J, Li Y, Li H-Y, Cai J, Wang X et al (2022) Interferon-γ induces retinal pigment epithelial cell Ferroptosis by a JAK1-2/STAT1/SLC7A11 signaling pathway in age-related macular degeneration. FEBS J 289:1968–1983. https://doi.org/10.1111/febs.16272
Zhong Y, Zhang W, Yu H, Lin L, Gao X, He J, Li D, Chen Y, Zeng Z, Xu Y et al (2022) Multi-platform-based characterization of ferroptosis in human colorectal cancer. iScience. 25:104750. https://doi.org/10.1016/j.isci.2022.104750
Chen Z, Gu Q, Chen R (2023) Promotive role of IRF7 in ferroptosis of colonic epithelial cells in ulcerative colitis by the miR-375-3p/SLC11A2 axis. Biomol Biomed 23:437–449. https://doi.org/10.17305/bjbms.2022.8081
Liu J, Ren Z, Yang L, Zhu L, Li Y, Bie C, Liu H, Ji Y, Chen D, Zhu M et al (2022) The NSUN5-FTH1/FTL pathway mediates ferroptosis in bone marrow-derived mesenchymal stem cells. Cell Death Discov 8:99. https://doi.org/10.1038/s41420-022-00902-z
Ke S, Wang C, Su Z, Lin S, Wu G (2022) Integrated analysis reveals critical ferroptosis regulators and FTL contribute to cancer progression in hepatocellular carcinoma. Front Genet. 13:897683. https://doi.org/10.3389/fgene.2022.897683
Zhang L, Chen Z, Xu A (2017) FTL: a novel predictor in gastric cancer. Int J Clin Exp Pathol 10:7865–7872
Liu J, Gao L, Zhan N, Xu P, Yang J, Yuan F, Xu Y, Cai Q, Geng R, Chen Q (2020) Hypoxia induced ferritin light chain (FTL) promoted epithelia mesenchymal transition and chemoresistance of glioma. J Exp Clin Cancer Res 39:137. https://doi.org/10.1186/s13046-020-01641-8
Funding
This study was funded by the National Natural Science Foundation of China (82072182, 82002061).
Author information
Authors and Affiliations
Contributions
BYS, YZ, and YP conceptualized and designed the study; BYS, WL, and YZ contributed to data curation and methodology; BYS, YP, KW, and ZC analyzed and interpreted the data; BYS wrote the manuscript; and BQS reviewed the manuscript and took part in study supervision. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
262_2023_3504_MOESM1_ESM.pdf
Supplemental Figure 1 A–I Results of K–M survival analysis of NMF cells clusters in the external dataset of bladder cancer. The survival analysis of each high and low NMF cell cluster grouping in bladder cancer was significantly different (p < 0.05) (PDF 514 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Song, B., Peng, Y., Zheng, Y. et al. Role of single-cell ferroptosis regulation in intercellular communication and skin cutaneous melanoma progression and immunotherapy. Cancer Immunol Immunother 72, 3523–3541 (2023). https://doi.org/10.1007/s00262-023-03504-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-023-03504-5