Abstract
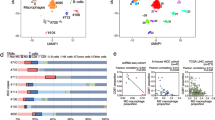
An important mechanism of oncolytic virotherapy in ameliorating cancer immunotherapy is by inducing significant changes in the immune landscape in the tumor microenvironment (TME). Despite this notion and the potential therapeutic implications, a comprehensive analysis of the immune changes in carcinomas induced by virotherapy has not yet been elucidated. We conducted single-cell RNA sequencing analysis on carcinomas treated with an HSV-2-based oncolytic virus to characterize the immunogenic changes in the TME. We specifically analyzed and compared the immune cell composition between viral treated and untreated tumors. We also applied CellChat to analyze the complex interactions among the infiltrated immune cells. Our data revealed significant infiltration of B cells in addition to other important immune cells, including CD4+, CD8+, and NK cells following virotherapy. Further analysis identified distinct subset compositions of the infiltrated immune cells and their activation status upon virotherapy. The intensive interactions among the infiltrated immune cells as revealed by CellChat analysis may further shape the immune landscape in favor of generating antitumor immunity. Our findings will facilitate the design of new strategies in incorporating immunotherapy into virotherapy for clinical translation. Moreover, the significant infiltration of B cells makes it suitable for combining virotherapy with immune checkpoint inhibitors.







Similar content being viewed by others
Data availability
All data needed to evaluate the conclusions in the paper are present in the paper and/or the Supplementary Materials. The raw scRNA-seq data is available on the GEO database (GEO accession number GSE186548). The materials will be available upon request with a simple MTA.
References
Conry RM, Westbrook B, McKee S, Norwood TG (2018) Talimogene laherparepvec: First in class oncolytic virotherapy. Hum Vaccin Immunother 14:839–846. https://doi.org/10.1080/21645515.2017.1412896
Hwang JK, Hong J, Yun C-O (2020) Oncolytic viruses and immune checkpoint inhibitors: preclinical developments to clinical trials. Int J Molecular Sciences 21:8627–8651. https://doi.org/10.3390/ijms21228627
Ma J, Ramachandran M, Jin C, Quijano-Rubio C, Martikainen M, Yu D, Essand M (2020) Characterization of virus-mediated immunogenic cancer cell death and the consequences for oncolytic virus-based immunotherapy of cancer. Cell Death Dis 11:48. https://doi.org/10.1038/s41419-020-2236-3
Li H, Dutuor A, Fu X, Zhang X (2007) Induction of strong antitumor immunity by an HSV-2-based oncolytic virus in a murine mammary tumor model. J Gene Med 9:161–169. https://doi.org/10.1002/jgm.1005
Li H, Dutuor A, Tao L, Fu X, Zhang X (2007) Virotherapy with a type 2 herpes simplex virus-derived oncolytic virus induces potent antitumor immunity against neuroblastoma. Clin Cancer Res 13:316–322
Pearl TM, Markert JM, Cassady KA, Ghonime MG (2019) Oncolytic Virus-based cytokine expression to improve immune activity in brain and solid tumors. Molecular Therapy Oncolytics 13:14–21. https://doi.org/10.1016/j.omto.2019.03.001
Pol JG, Workenhe ST, Konda P, Gujar S, Kroemer G (2020) Cytokines in oncolytic virotherapy. Cytokine Growth Factor Rev 56:4–27. https://doi.org/10.1016/2020.10.007
Berkey SE, Thorne SH, Bartlett DL (2017) Oncolytic virotherapy and the tumor microenvironment. Adv Exp Med Biol 1036:157–172. https://doi.org/10.1007/978-3-319-67577-0_11
Thorsson V, Gibbs DL, Brown SD et al (2018) The immune landscape of cancer. Immunity 48(812–30):e14. https://doi.org/10.1016/j.immuni.2018.03.023
Chon HJ, Lee WS, Yang H et al (2019) Tumor microenvironment remodeling by intratumoral oncolytic vaccinia virus enhances the efficacy of immune-checkpoint blockade. Clin Cancer Res 25:1612–1623. https://doi.org/10.1158/1078-0432.Ccr-18-1932
Linette GP, Carreno BM (2019) Tumor-infiltrating lymphocytes in the checkpoint inhibitor era. Curr Hematol Malig Rep 14:286–291. https://doi.org/10.1007/s11899-019-00523-x
Uryvaev A, Passhak M, Hershkovits D, Sabo E, Bar-Sela G (2018) The role of tumor-infiltrating lymphocytes (TILs) as a predictive biomarker of response to anti-PD1 therapy in patients with metastatic non-small cell lung cancer or metastatic melanoma. Med Oncol 35:25. https://doi.org/10.1007/s12032-018-1080-0
Hendry S, Salgado R, Gevaert T et al (2017) Assessing tumor-infiltrating lymphocytes in solid tumors: a practical review for pathologists and proposal for a standardized method from the international immuno-oncology biomarkers working group: Part 2: TILs in melanoma, gastrointestinal tract carcinomas, non-small cell lung carcinoma and mesothelioma, endometrial and ovarian carcinomas, squamous cell carcinoma of the head and neck, genitourinary carcinomas, and primary brain tumors. Adv Anat Pathol 24:311–335. https://doi.org/10.1097/PAP.0000000000000161
Fu X, Rivera A, Tao L, Zhang X (2015) An HSV-2 based oncolytic virus can function as an attractant to guide migration of adoptively transferred T cells to tumor sites. Oncotarget 6:902–914
Ribas A, Dummer R, Puzanov I et al (2017) Oncolytic virotherapy promotes intratumoral T cell infiltration and improves Anti-PD-1 immunotherapy. Cell 170(1109–19):e10. https://doi.org/10.1016/j.cell.2017.08.027
Koske I, Rössler A, Pipperger L et al (2019) Oncolytic virotherapy enhances the efficacy of a cancer vaccine by modulating the tumor microenvironment. Int J Cancer 145:1958–1969. https://doi.org/10.1002/ijc.32325
Fu X, Tao L, Wu W, Zhang X (2020) Arming HSV-based oncolytic viruses with the ability to redirect the host’s innate antiviral immunity to attack tumor cells. Mol Ther Oncolytics 19:33–46. https://doi.org/10.1016/j.omto.2020.09.002
Fu X, Tao L, Cai R, Prigge J, Zhang X (2006) A mutant Type 2 Herpes Simplex Virus deleted for the protein kinase domain of the ICP10 gene is a potent oncolytic virus. Mol Ther 13:882–890
Fu X, Tao L, Li M, Fisher WE, Zhang X (2006) Effective treatment of pancreatic cancer xenografts with a conditionally replicating virus derived from type 2 herpes simplex virus. Clin Cancer Res 12:3152–3157
Du Y, Huang Q, Arisdakessian C, Garmire LX (2020) Evaluation of STAR and Kallisto on single cell RNA-Seq data alignment. G3: Genes|Genomes|Genetics. 10: 1775–83. https://doi.org/10.1534/g3.120.401160
Butler A, Hoffman P, Smibert P, Papalexi E, Satija R (2018) Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol 36:411–420. https://doi.org/10.1038/nbt.4096
Pei G, Yan F, Simon LM, Dai Y, Jia P, Zhao Z (2021) deCS: a tool for systematic cell type annotations of single-cell RNA sequencing data among human tissues. bioRxiv. 2021.09.19.460993. https://doi.org/10.1101/2021.09.19.460993
Aran D, Looney AP, Liu L et al (2019) Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat Immunol 20:163–172. https://doi.org/10.1038/s41590-018-0276-y
Liao M, Liu Y, Yuan J et al (2020) Single-cell landscape of bronchoalveolar immune cells in patients with COVID-19. Nat Med 26:842–844. https://doi.org/10.1038/s41591-020-0901-9
Vieth B, Parekh S, Ziegenhain C, Enard W, Hellmann I (2019) A systematic evaluation of single cell RNA-seq analysis pipelines. Nat Commun 10:4667. https://doi.org/10.1038/s41467-019-12266-7
Stuart T, Butler A, Hoffman P et al (2019) Comprehensive integration of single-cell data. Cell 177(1888–902):e21. https://doi.org/10.1016/j.cell.2019.05.031
Nakanishi Y, Lu B, Gerard C, Iwasaki A (2009) CD8(+) T lymphocyte mobilization to virus-infected tissue requires CD4(+) T-cell help. Nature 462:510–513. https://doi.org/10.1038/nature08511
Bengsch B, Seigel B, Flecken T, Wolanski J, Blum HE, Thimme R (2012) Human Th17 cells express high levels of enzymatically active dipeptidylpeptidase IV (CD26). J Immunol 188:5438–5447. https://doi.org/10.4049/jimmunol.1103801
Wakita D, Sumida K, Iwakura Y, Nishikawa H, Ohkuri T, Chamoto K, Kitamura H, Nishimura T (2010) Tumor-infiltrating IL-17-producing γδ T cells support the progression of tumor by promoting angiogenesis. Eur J Immunol 40:1927–1937. https://doi.org/10.1002/eji.200940157
Ma Y, Aymeric L, Locher C et al (2011) Contribution of IL-17-producing gamma delta T cells to the efficacy of anticancer chemotherapy. J Exp Med 208:491–503. https://doi.org/10.1084/jem.20100269
Petitprez F, de Reyniès A, Keung EZ et al (2020) B cells are associated with survival and immunotherapy response in sarcoma. Nature 577:556–560. https://doi.org/10.1038/s41586-019-1906-8
Helmink BA, Reddy SM, Gao J et al (2020) B cells and tertiary lymphoid structures promote immunotherapy response. Nature 577:549–555. https://doi.org/10.1038/s41586-019-1922-8
Cabrita R, Lauss M, Sanna A et al (2020) Tertiary lymphoid structures improve immunotherapy and survival in melanoma. Nature 577:561–565. https://doi.org/10.1038/s41586-019-1914-8
Hollern DP, Xu N, Thennavan A et al (2019) B Cells and T follicular helper cells mediate response to checkpoint inhibitors in high mutation burden mouse models of breast cancer. Cell 179:1191–206.e21. https://doi.org/10.1016/j.cell.2019.10.028
Shen Y, Iqbal J, Xiao L et al (2004) Distinct gene expression profiles in different B-cell compartments in human peripheral lymphoid organs. BMC Immunol 5:20. https://doi.org/10.1186/1471-2172-5-20
DeNardo DG, Ruffell B (2019) Macrophages as regulators of tumour immunity and immunotherapy. Nat Rev Immunol 19:369–382. https://doi.org/10.1038/s41577-019-0127-6
Jin S, Guerrero-Juarez CF, Zhang L, Chang I, Ramos R, Kuan C-H, Myung P, Plikus MV, Nie Q (2021) Inference and analysis of cell-cell communication using Cell Chat. Nat Commun 12:1088. https://doi.org/10.1038/s41467-021-21246-9
Janssen LME, Ramsay EE, Logsdon CD, Overwijk WW (2017) The immune system in cancer metastasis: friend or foe? J Immunother Cancer 5:79. https://doi.org/10.1186/s40425-017-0283-9
Brucher BL, Jamall IS (2014) Cell-cell communication in the tumor microenvironment, carcinogenesis, and anticancer treatment. Cell Physiol Biochem 34:213–243. https://doi.org/10.1159/000362978
Calandra T, Roger T (2003) Macrophage migration inhibitory factor: a regulator of innate immunity. Nat Rev Immunol 3:791–800. https://doi.org/10.1038/nri1200
Russell L, Swanner J, Jaime-Ramirez AC et al (2018) PTEN expression by an oncolytic herpesvirus directs T-cell mediated tumor clearance. Nat Commun 9:5006. https://doi.org/10.1038/s41467-018-07344-1
Samson A, Scott KJ, Taggart D et al. (2018) Intravenous delivery of oncolytic reovirus to brain tumor patients immunologically primes for subsequent checkpoint blockade. Sci Transl Med. 10: eaam7577. https://doi.org/10.1126/scitranslmed.aam7577
Burger JA, Quiroga MP, Hartmann E, Bürkle A, Wierda WG, Keating MJ, Rosenwald A (2009) High-level expression of the T-cell chemokines CCL3 and CCL4 by chronic lymphocytic leukemia B cells in nurselike cell cocultures and after BCR stimulation. Blood 113:3050–3058. https://doi.org/10.1182/blood-2008-07-170415
Tsai S-C, Lin S-J, Lin C-J et al (2013) Autocrine CCL3 and CCL4 Induced by the Oncoprotein LMP1 Promote Epstein-Barr Virus-Triggered B Cell proliferation. J Virol 87:9041–9052. https://doi.org/10.1128/jvi.00541-13
Takahashi K, Sivina M, Hoellenriegel J et al (2015) CCL3 and CCL4 are biomarkers for B cell receptor pathway activation and prognostic serum markers in diffuse large B cell lymphoma. Br J Haematol 171:726–735. https://doi.org/10.1111/bjh.13659
Wang Q, Ren J, Morgan S, Liu Z, Dou C, Liu B (2014) Monocyte chemoattractant protein-1 (MCP-1) regulates macrophage cytotoxicity in abdominal aortic aneurysm. PLoS ONE 9:e92053. https://doi.org/10.1371/journal.pone.0092053
Ramelyte E, Tastanova A, Balazs Z et al (2021) Oncolytic virotherapy-mediated anti-tumor response: a single-cell perspective. Cancer Cell 39:394-406.e4. https://doi.org/10.1016/j.ccell.2020.12.022
Petitprez F, Meylan M, de Reyniès A, Sautès-Fridman C, Fridman WH (2020) The Tumor microenvironment in the response to immune checkpoint blockade therapies. Front Immunol 11:784. https://doi.org/10.3389/fimmu.2020.00784
Oh JE, Iijima N, Song E, Lu P, Klein J, Jiang R, Kleinstein SH, Iwasaki A (2019) Migrant memory B cells secrete luminal antibody in the vagina. Nature 571:122–126. https://doi.org/10.1038/s41586-019-1285-1
Ford ES, Sholukh AM, Boytz R et al (2021) B cells, antibody-secreting cells, and virus-specific antibodies respond to herpes simplex virus 2 reactivation in skin. J Clin Invest 131:e142088. https://doi.org/10.1172/JCI142088
Acknowledgements
This work was supported by the Cancer Prevention and Research Institute of Texas (CPRIT) grant RP200464 and a grant from the William and Ella Owens Medical Research Foundation (to X.Z.), the National Institutes of Health grant R01LM012806 (to Z.Z), and the CPRIT grant RP180734 (to Z.Z.). We thank the University of Houston sequencing core for library construction and sequencing, and Dr. Brandon Mistretta for his assistance with pre-processing and upstream analysis of the scRNA-seq data. We thank Dr. Xinli Liu for assistance with the cell sorting, and Dr. Wanfu Wu for assistance with IHC staining. We also thank Drs. Weiyi Peng and Navin Varadarajan for advice, and Dr. Tho Tran for careful reading of the manuscript.
Author information
Authors and Affiliations
Contributions
D.R and X.Z. conceived the project and designed the experiments. D.R. executed all the experiments in the study. G.P. and D.R. performed bioinformatics analysis. D.R. and X.Z. wrote the manuscript draft and G.P. and Z.Z. helped the finalization of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ravirala, D., Pei, G., Zhao, Z. et al. Comprehensive characterization of tumor immune landscape following oncolytic virotherapy by single-cell RNA sequencing. Cancer Immunol Immunother 71, 1479–1495 (2022). https://doi.org/10.1007/s00262-021-03084-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-021-03084-2