Abstract
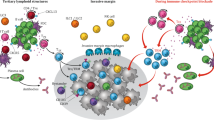
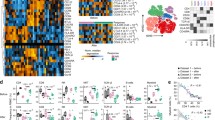
Adoptive cell transfer (ACT) using autologous tumor infiltrating lymphocytes (TILs) was previously shown to yield clinical response in metastatic melanoma patients as an advanced line. Unfortunately, there is no reliable marker for predicting who will benefit from the treatment. We analyzed TIL samples from the infusion bags used for treatment of 57 metastatic melanoma patients and compared their microRNA profiles. The discovery cohort included six responding patients and seven patients with progressive disease, as defined by RECIST1.1. High throughput analysis with NanoString nCounter demonstrated significantly higher levels of miR-34a-5p and miR-22-3p among TIL from non-responders. These results were validated in TIL infusion bag samples from an independent cohort of 44 patients, using qRT-PCR of the individual microRNAs. Using classification trees, a data-driven predictive model for response was built, based on the level of expression of these microRNAs. Patients that achieved stable disease were classified with responders, setting apart the patients with progressive disease. Moreover, the expression levels of miR-34a-5p in the infused TIL created distinct survival groups, which strongly supports its role as a potential biomarker for TIL-ACT therapy. Indeed, when tested against autologous melanoma cells, miRLow TIL cultures exhibited significantly higher cytotoxic activity than miRHigh TIL cultures, and expressed features of terminally exhausted effectors. Finally, overexpression of miR-34a-5p or miR-22-3p in TIL inhibited their cytotoxic ability in vitro. Overall, we show that a two-microRNA signature correlates with failure of TIL-ACT therapy and survival in melanoma patients.






Similar content being viewed by others
References
Sang M, Wang L, Ding C, Zhou X, Wang B, Lian Y, Shan B (2011) Melanoma-associated antigen genes - an update. Cancer Lett 302:85–90. (S0304-3835(10)00506-9)
Lawrence MS, Stojanov P, Polak P et al (2013) Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499:214–218. https://doi.org/10.1038/nature12213
Zikich D, Schachter J, Besser MJ (2013) Immunotherapy for the management of advanced melanoma: the next steps. Am J Clin Dermatol 14:261–272. https://doi.org/10.1007/s40257-013-0013-0
Lee C, Collichio F, Ollila D, Moschos S (2013) Historical review of melanoma treatment and outcomes. Clin Dermatol 31:141–147. https://doi.org/10.1016/j.clindermatol.2012.08.015s0738-081x
Hodi FS, O’Day SJ, McDermott DF et al (2010) Improved survival with ipilimumab in patients with metastatic melanoma. N Engl J Med 363:711–723. https://doi.org/10.1056/nejmoa1003466
Robert C, Thomas L, Bondarenko I et al (2011) Ipilimumab plus dacarbazine for previously untreated metastatic melanoma. N Engl J Med 364:2517–2526. https://doi.org/10.1056/nejmoa1104621
Robert C, Long GV, Brady B et al (2014) Nivolumab in Previously Untreated Melanoma without BRAF Mutation. N Engl J Med. https://doi.org/10.1056/nejmoa1412082
Robert C, Schachter J, Long GV et al (2015) Pembrolizumab versus Ipilimumab in Advanced Melanoma. N Engl J Med 372:2521–2532. https://doi.org/10.1056/nejmoa1503093
Larkin J, Chiarion-Sileni V, Gonzalez R et al (2015) Combined Nivolumab and Ipilimumab or Monotherapy in Untreated Melanoma. N Engl J Med 373:23–34. https://doi.org/10.1056/nejmoa1504030
Margolis N, Markovits E, Markel G (2019) Reprogramming lymphocytes for the treatment of melanoma: from biology to therapy. Adv Drug Deliv Rev 141:104–124
Markel G, Seidman R, Stern N et al (2006) Inhibition of human tumor-infiltrating lymphocyte effector functions by the homophilic carcinoembryonic cell adhesion molecule 1 interactions. J Immunol 177:6062–6071. (177/9/6062)
Ortenberg R, Sapir Y, Raz L et al (2012) Novel Immunotherapy for Malignant Melanoma with a Monoclonal Antibody That Blocks CEACAM1 Homophilic Interactions. Mol Cancer Ther 11:1300–1310. https://doi.org/10.1158/1535-7163
Rosenberg SA, Dudley ME (2009) Adoptive cell therapy for the treatment of patients with metastatic melanoma. Curr Opin Immunol 21:233–240. (S0952-7915(09)00025-9)
Besser MJ, Shapira-Frommer R, Treves AJ et al (2009) Minimally cultured or selected autologous tumor-infiltrating lymphocytes after a lympho-depleting chemotherapy regimen in metastatic melanoma patients. J Immunother 32:415–423. https://doi.org/10.1097/cji.0b013e31819c8bda00002371
Dudley ME, Wunderlich JR, Shelton TE, Even J, Rosenberg SA (2003) Generation of tumor-infiltrating lymphocyte cultures for use in adoptive transfer therapy for melanoma patients. J Immunother 26:332–342
Tran KQ, Zhou J, Durflinger KH, Langhan MM, Shelton TE, Wunderlich JR, Robbins PF, Rosenberg SA, Dudley ME (2008) Minimally cultured tumor-infiltrating lymphocytes display optimal characteristics for adoptive cell therapy. J Immunother 31:742–751. https://doi.org/10.1097/cji.0b013e31818403d500002371
Besser MJ, Shapira-Frommer R, Itzhaki O et al (2013) Adoptive Transfer of Tumor Infiltrating Lymphocytes in Metastatic Melanoma Patients: intent-to-Treat Analysis and Efficacy after Failure to Prior Immunotherapies. Clin Cancer Res 19:4792–4800. (1078-0432.CCR-13-0380)
Rosenberg SA, Yang JC, Sherry RM et al (2011) Durable complete responses in heavily pretreated patients with metastatic melanoma using T-cell transfer immunotherapy. Clin Cancer Res 17:4550–4557. https://doi.org/10.1158/1078-0432.ccr-11-01161078-0432
Radvanyi LG, Bernatchez C, Zhang M et al (2012) Specific lymphocyte subsets predict response to adoptive cell therapy using expanded autologous tumor-infiltrating lymphocytes in metastatic melanoma patients. Clin Cancer Res 18:6758–6770. https://doi.org/10.1158/1078-0432.ccr-12-11771078-0432
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297. (S0092867404000455)
Kim VN, Nam JW (2006) Genomics of microRNA. Trends Genet 22:165–173. (S0168-9525(06)00021-7)
Wojcicka A, de la Chapelle A, Jazdzewski K (2013) MicroRNA-related sequence variations in human cancers. Hum Genet. https://doi.org/10.1007/s00439-013-1397-x
Friedman RC, Farh KK, Burge CB, Bartel DP (2009) Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 19:92–105. https://doi.org/10.1101/gr.082701.108
O’Connell RM, Rao DS, Chaudhuri AA, Baltimore D (2010) Physiological and pathological roles for microRNAs in the immune system. Nat Rev Immunol 10:111–122. https://doi.org/10.1038/nri2708
Basak I, Patil KS, Alves G, Larsen JP, Moller SG (2015) microRNAs as neuroregulators, biomarkers and therapeutic agents in neurodegenerative diseases. Cell Mol Life Sci. https://doi.org/10.1007/s00018-015-2093
Pencheva N, Tavazoie SF (2013) Control of metastatic progression by microRNA regulatory networks. Nat Cell Biol 15:546–554. https://doi.org/10.1038/ncb2769ncb2769
Di Leva G, Croce CM (2013) miRNA profiling of cancer. Curr Opin Genet Dev 23:3–11. https://doi.org/10.1016/j.gde.2013.01.004s0959-437x
Iorio MV, Croce CM (2012) microRNA involvement in human cancer. Carcinogenesis 33:1126–1133. https://doi.org/10.1093/carcin/bgs140bgs140
Hayes J, Peruzzi PP, Lawler S (2014) MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol Med. 20:460–469. https://doi.org/10.1016/j.molmed.2014.06.005
Nana-Sinkam SP, Croce CM (2013) Clinical applications for microRNAs in cancer. Clin Pharmacol Ther 93:98–104. https://doi.org/10.1038/clpt.2012.192
Itzhaki O, Hovav E, Ziporen Y et al (2011) Establishment and large-scale expansion of minimally cultured “young” tumor infiltrating lymphocytes for adoptive transfer therapy. J Immunother 34:212–220. https://doi.org/10.1097/cji.0b013e318209c94c
Besser MJ, Shapira-Frommer R, Treves AJ et al (2010) Clinical responses in a phase II study using adoptive transfer of short-term cultured tumor infiltration lymphocytes in metastatic melanoma patients. Clin Cancer Res 16:2646–2655. https://doi.org/10.1158/1078-0432.ccr-10-00411078-0432
Galore-Haskel G, Nemlich Y, Greenberg E et al. (2015) A novel immune resistance mechanism of melanoma cells controlled by the ADAR1 enzyme. Oncotarget. 6: 28999-9015. https://doi.org/10.18632/oncotarget.4905
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29:15–21. https://doi.org/10.1093/bioinformatics/bts635
Liao Y, Smyth GK, Shi W (2014) FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30:923–930
Law CW, Chen Y, Shi W, Smyth GK (2014) Voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol 15:R29. https://doi.org/10.1186/gb-2014-15-2-r29
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43:e47. https://doi.org/10.1093/nar/gkv007
Rooney MS, Shukla SA, Wu CJ, Getz G, Hacohen N (2015) Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160:48–61. https://doi.org/10.1016/j.cell.2014.12.033
Hothorn T, Hornik K, Zeileis A (2006) Unbiased recursive partitioning: a conditional inference framework. Journal of Computational and Graphical statistics. 15:651–674
Shmueli G, Bruce P, Yahav I, Petal R, Lichtendahl KC (2017) Data mining for business intelligence: Concepts, techniques, and applications in R.. 1st ed. John Wiley and Sons
Zikich D, Schachter J, Besser MJ (2016) Predictors of tumor-infiltrating lymphocyte efficacy in melanoma. Immunotherapy. 8:35–43. https://doi.org/10.2217/imt.15.99
Ji Y, Hocker JD, Gattinoni L (2015) Enhancing adoptive T cell immunotherapy with microRNA therapeutics. Semin Immunol. (S1044-5323(15)00100-1)
Jeker LT, Bluestone JA (2013) MicroRNA regulation of T-cell differentiation and function. Immunol Rev 253:65–81. https://doi.org/10.1111/imr.12061
Paley MA, Kroy DC, Odorizzi PM et al (2012) Progenitor and terminal subsets of CD8 + T cells cooperate to contain chronic viral infection. Science 338:1220–1225. https://doi.org/10.1126/science.1229620
Thommen DS, Koelzer VH, Herzig P et al (2018) A transcriptionally and functionally distinct PD-1 + CD8 + T cell pool with predictive potential in non-small-cell lung cancer treated with PD-1 blockade. Nat Med 24:994–1004. https://doi.org/10.1038/s41591-018-0057-z
LaFleur MW, Nguyen TH, Coxe MA et al (2019) PTPN2 regulates the generation of exhausted CD8 + T cell subpopulations and restrains tumor immunity. Nat Immunol 20:1335–1347. https://doi.org/10.1038/s41590-019-0480-4
Agostini M, Knight RA (2014) miR-34: from bench to bedside. Oncotarget. 5: 872-81. doi: 1825 [pii] https://doi.org/10.18632/oncotarget.1825
Xiong J (2012) Emerging roles of microRNA-22 in human disease and normal physiology. Curr Mol Med 12:247–258. (CMM-EPub-201201190201-005)
Cobb BS, Hertweck A, Smith J et al (2006) A role for Dicer in immune regulation. J Exp Med 203:2519–2527. jem.20061692
Kuijk LM, Verstege MI, Rekers NV, Bruijns SC, Hooijberg E, Roep BO, de Gruijl TD, van Kooyk Y, Unger WW (2013) Notch controls generation and function of human effector CD8 + T cells. Blood 121:2638–2646. https://doi.org/10.1182/blood-2012-07-442962
Sierra RA, Thevenot P, Raber PL, Cui Y, Parsons C, Ochoa AC, Trillo-Tinoco J, Del Valle L, Rodriguez PC (2014) Rescue of notch-1 signaling in antigen-specific CD8 + T cells overcomes tumor-induced T-cell suppression and enhances immunotherapy in cancer. Cancer Immunol Res 2:800–811. https://doi.org/10.1158/2326-6066
Yang L, Pang Y, Moses HL (2010) TGF-beta and immune cells: an important regulatory axis in the tumor microenvironment and progression. Trends Immunol 31:220–227. https://doi.org/10.1016/j.it.2010.04.002
Acknowledgements
The authors would like to thank the Aronson Fund and the Lemelbaum family fund for their generous support. G.M is supported by the Melanoma Research Alliance Saban Family Team Sciences Award, Samueli Foundation Grant for Integrative Immuno-Oncology and Israel Science Foundation Grant 15/1925.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors state no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Galore-Haskel, G., Greenberg, E., Yahav, I. et al. microRNA expression patterns in tumor infiltrating lymphocytes are strongly associated with response to adoptive cell transfer therapy. Cancer Immunol Immunother 70, 1541–1555 (2021). https://doi.org/10.1007/s00262-020-02782-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-020-02782-7