Abstract
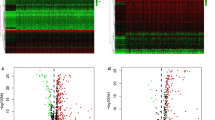
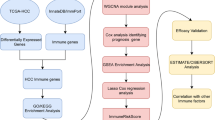
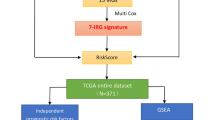
The immune microenvironment plays a vital role in the progression of hepatocellular carcinoma (HCC). Thousands of immune-related genes (IRGs) have been identified, but their effects on HCC are not fully understood. In this study, we identified the differentially expressed IRGs and analyzed their functions in HCC in a systematic way. Furthermore, we constructed a diagnostic and a prognostic model using multiple statistical methods, and both models had good distinguishing performance, which we verified in several independent datasets. This diagnostic model was also adaptable to proteomic data. The combination of a prognostic risk model and classic clinical staging can effectively distinguish patients in high- and low-risk groups. Furthermore, we systematically explore the differences in the immune microenvironment between the high-risk group and the low-risk group to help clinical decision-making. In summary, we systematically analyzed immune-related genes in HCC, explored their functions, constructed a diagnostic and a prognostic model and investigated potential therapeutic schedules in high-risk patients. The model performance was verified in multiple databases. Our findings can provide directions for future research.








Similar content being viewed by others
References
Yang JD, Hainaut P, Gores GJ, Amadou A, Plymoth A, Roberts LR (2019) A global view of hepatocellular carcinoma: trends, risk, prevention and management. Nat Rev Gastroenterol Hepatol 16(10):589–604. https://doi.org/10.1038/s41575-019-0186-y
Turato C, Balasso A, Carloni V, Tiribelli C, Mastrotto F, Mazzocca A, Pontisso P (2017) New molecular targets for functionalized nanosized drug delivery systems in personalized therapy for hepatocellular carcinoma. J Control Release 268:184–197. https://doi.org/10.1016/j.jconrel.2017.10.027
Fu Y, Liu S, Zeng S, Shen H (2019) From bench to bed: the tumor immune microenvironment and current immunotherapeutic strategies for hepatocellular carcinoma. J Exp Clin Cancer Res 38(1):396. https://doi.org/10.1186/s13046-019-1396-4
Cheng AL, Hsu C, Chan SL, Choo SP, Kudo M (2020) Challenges of combination therapy with immune checkpoint inhibitors for hepatocellular carcinoma. J Hepatol 72(2):307–319. https://doi.org/10.1016/j.jhep.2019.09.025
GTEx Consortium (2015) Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348(6235):648–660. https://doi.org/10.1126/science.1262110
Hutter C, Zenklusen JC (2018) The cancer genome atlas: creating lasting value beyond its data. Cell 173(2):283–285. https://doi.org/10.1016/j.cell.2018.03.042
Jiang P, Liu XS (2015) Big data mining yields novel insights on cancer. Nat Genet 47(2):103–104. https://doi.org/10.1038/ng.3205
Zhang J, Baran J, Cros A, Guberman JM, Haider S, Hsu J, Liang Y, Rivkin E, Wang J, Whitty B, Wong-Erasmus M, Yao L, Kasprzyk A (2011) International Cancer Genome Consortium Data Portal–a one-stop shop for cancer genomics data. Database (Oxford). https://doi.org/10.1093/database/bar026
Edwards NJ, Oberti M, Thangudu RR, Cai S, McGarvey PB, Jacob S, Madhavan S, Ketchum KA (2015) The CPTAC data portal: a resource for cancer proteomics research. J Proteome Res 14(6):2707–2713. https://doi.org/10.1021/pr501254j
Uhlén M, Fagerberg L, Hallström BM, Lindskog C, Oksvold P, Mardinoglu A, Sivertsson Å, Kampf C, Sjöstedt E, Asplund A, Olsson I, Edlund K, Lundberg E, Navani S, Szigyarto CA, Odeberg J, Djureinovic D, Takanen JO, Hober S, Alm T, Edqvist PH, Berling H, Tegel H, Mulder J, Rockberg J, Nilsson P, Schwenk JM, Hamsten M, von Feilitzen K, Forsberg M, Persson L, Johansson F, Zwahlen M, von Heijne G, Nielsen J, Pontén F (2015) Proteomics Tissue-based map of the human proteome. Science 347(6220):1260419. https://doi.org/10.1126/science.1260419
Bhattacharya S, Dunn P, Thomas CG et al (2018) ImmPort, toward repurposing of open access immunological assay data for translational and clinical research. Sci Data 5:180015. https://doi.org/10.1038/sdata.2018.15
Breuer K, Foroushani AK, Laird MR, Chen C, Sribnaia A, Lo R, Winsor GL, Hancock RE, Brinkman FS, Lynn DJ (2013) InnateDB: systems biology of innate immunity and beyond–recent updates and continuing curation. Nucl Acids Res 41:D1228–1233. https://doi.org/10.1093/nar/gks1147
Pei G, Chen L, Zhang W (2017) WGCNA application to proteomic and metabolomic data analysis. Methods Enzymol 585:135–158. https://doi.org/10.1016/bs.mie.2016.09.016
Gujar H, Weisenberger DJ, Liang G (2019) The roles of human DNA methyltransferases and their isoforms in shaping the epigenome. Genes (Basel). https://doi.org/10.3390/genes10020172
Khare SP, Habib F, Sharma R, Gadewal N, Gupta S, Galande S (2012) HIstome: a relational knowledgebase of human histone proteins and histone modifying enzymes. Nucl Acids Res 40:D337–342. https://doi.org/10.1093/nar/gkr1125
Liu T, Ortiz JA, Taing L, Meyer CA, Lee B, Zhang Y, Shin H, Wong SS, Ma J, Lei Y, Pape UJ, Poidinger M, Chen Y, Yeung K, Brown M, Turpaz Y, Liu XS (2011) Cistrome: an integrative platform for transcriptional regulation studies. Genome Biol 12(8):R83. https://doi.org/10.1186/gb-2011-12-8-r83
Chen M, Wong CM (2020) The emerging roles of N6-methyladenosine (m6A) deregulation in liver carcinogenesis. Mol Cancer 19(1):44. https://doi.org/10.1186/s12943-020-01172-y
Williams K, Christensen J, Helin K (2011) DNA methylation: TET proteins-guardians of CpG islands. EMBO Rep 13(1):28–35. https://doi.org/10.1038/embor.2011.233
Li K, Guo ZW, Zhai XM, Yang XX, Wu YS, Liu TC (2020) RBPTD: a database of cancer-related RNA-binding proteins in humans. Database (Oxford). https://doi.org/10.1093/database/baz156
Subramanian A, Kuehn H, Gould J, Tamayo P, Mesirov JP (2007) GSEA-P: a desktop application for Gene Set Enrichment Analysis. Bioinformatics 23(23):3251–3253. https://doi.org/10.1093/bioinformatics/btm369
Hänzelmann S, Castelo R, Guinney J (2013) GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform 14:7. https://doi.org/10.1186/1471-2105-14-7
Racle J, Gfeller D (2020) EPIC: a tool to estimate the proportions of different cell types from bulk gene expression data. Methods Mol Biol 2120:233–248. https://doi.org/10.1007/978-1-0716-0327-7_17
Liberzon A, Birger C, Thorvaldsdóttir H, Ghandi M, Mesirov JP, Tamayo P (2015) The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst 1(6):417–425. https://doi.org/10.1016/j.cels.2015.12.004
Yoshihara K, Shahmoradgoli M, Martínez E, Vegesna R, Kim H, Torres-Garcia W, Treviño V, Shen H, Laird PW, Levine DA, Carter SL, Getz G, Stemke-Hale K, Mills GB, Verhaak RG (2013) Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun 4:2612. https://doi.org/10.1038/ncomms3612
Bertolazzi P, Bock ME, Guerra C (2013) On the functional and structural characterization of hubs in protein-protein interaction networks. Biotechnol Adv 31(2):274–286. https://doi.org/10.1016/j.biotechadv.2012.12.002
Aubé C, Bazeries P, Lebigot J, Cartier V, Boursier J (2017) Liver fibrosis, cirrhosis, and cirrhosis-related nodules: Imaging diagnosis and surveillance. Diagn Interv Imaging 98(6):455–468. https://doi.org/10.1016/j.diii.2017.03.003
Zeng D, Li M, Zhou R, Zhang J, Sun H, Shi M, Bin J, Liao Y, Rao J, Liao W (2019) Tumor microenvironment characterization in gastric cancer identifies prognostic and immunotherapeutically relevant gene signatures. Cancer Immunol Res 7(5):737–750. https://doi.org/10.1158/2326-6066.CIR-18-0436
Watson MM, Lea D, Gudlaugsson E, Skaland I, Hagland HR, Søreide K (2020) Prevalence of PD-L1 expression is associated with EMAST, density of peritumoral T-cells and recurrence-free survival in operable non-metastatic colorectal cancer. Cancer Immunol Immunother. https://doi.org/10.1007/s00262-020-02573-0
Hollern DP, Xu N, Thennavan A, Glodowski C, Garcia-Recio S, Mott KR, He X, Garay JP, Carey-Ewend K, Marron D, Ford J, Liu S, Vick SC, Martin M, Parker JS, Vincent BG, Serody JS, Perou CM (2019) B cells and T follicular helper cells mediate response to checkpoint inhibitors in high mutation burden mouse models of breast cancer. Cell 179(5):1191–1206.e21. https://doi.org/10.1016/j.cell.2019.10.028
Luchini C, Bibeau F, Ligtenberg M, Singh N, Nottegar A, Bosse T, Miller R, Riaz N, Douillard JY, Andre F, Scarpa A (2019) ESMO recommendations on microsatellite instability testing for immunotherapy in cancer, and its relationship with PD-1/PD-L1 expression and tumour mutational burden: a systematic review-based approach. Ann Oncol 30(8):1232–1243. https://doi.org/10.1093/annonc/mdz116
Garrido F (2019) HLA class-I expression and cancer immunotherapy. Adv Exp Med Biol 1151:79–90. https://doi.org/10.1007/978-3-030-17864-2_3
Burke JD, Young HA (2019) IFN-γ: a cytokine at the right time, is in the right place. Semin Immunol 43:101280. https://doi.org/10.1016/j.smim.2019.05.002
Ungefroren H (2019) Blockade of TGF-β signaling: a potential target for cancer immunotherapy. Expert Opin Ther Targets 23(8):679–693. https://doi.org/10.1080/14728222.2019.1636034
Lu C, Rong D, Zhang B, Zheng W, Wang X, Chen Z, Tang W (2019) Current perspectives on the immunosuppressive tumor microenvironment in hepatocellular carcinoma: challenges and opportunities. Mol Cancer 18(1):130. https://doi.org/10.1186/s12943-019-1047-6
Refolo MG, Lotesoriere C, Messa C, Caruso MG, D'Alessandro R (2020) Integrated immune gene expression signature and molecular classification in gastric cancer: new insights. J Leukoc Biol. https://doi.org/10.1002/JLB.4MR0120-221R
Liu J, Lichtenberg T, Hoadley KA, Poisson LM, Lazar AJ, Cherniack AD, Kovatich AJ, Benz CC, Levine DA, Lee AV, Omberg L, Wolf DM, Shriver CD, Thorsson V, Cancer Genome Atlas Research Network, Hu H (2018) An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell 173(2):400–416.e11. https://doi.org/10.1016/j.cell.2018.02.052
Chen M, Wei L, Law CT, Tsang FH, Shen J, Cheng CL, Tsang LH, Ho DW, Chiu DK, Lee JM, Wong CC, Ng IO, Wong CM (2018) RNA N6-methyladenosine methyltransferase-like 3 promotes liver cancer progression through YTHDF2-dependent posttranscriptional silencing of SOCS2. Hepatology 67(6):2254–2270. https://doi.org/10.1002/hep.29683
Di Tommaso L, Spadaccini M, Donadon M, Personeni N, Elamin A, Aghemo A, Lleo A (2019) Role of liver biopsy in hepatocellular carcinoma. World J Gastroenterol 25(40):6041–6052. https://doi.org/10.3748/wjg.v25.i40.6041
Schwabe RF, Greten TF (2020) Gut microbiome in HCC. Mechanisms, diagnosis and therapy. J Hepatol 72(2):230–238. https://doi.org/10.1016/j.jhep.2019.08.016
Jihye C, Jinsil S (2012) Application of radiotherapeutic strategies in the BCLC-defined stages of hepatocellular carcinoma. Liver Cancer 1(3–4):216–225. https://doi.org/10.1159/000343836
Gentile D, Donadon M, Lleo A, Aghemo A, Roncalli M, di Tommaso L, Torzilli G (2020) Surgical treatment of hepatocholangiocarcinoma: a systematic review. Liver Cancer 9(1):15–27. https://doi.org/10.1159/000503719
Prieto J, Melero I, Sangro B (2015) Immunological landscape and immunotherapy of hepatocellular carcinoma. Nat Rev Gastroenterol Hepatol 12(12):681–700. https://doi.org/10.1038/nrgastro.2015.173
Rhodes DR, Kalyana-Sundaram S, Mahavisno V, Barrette TR, Ghosh D, Chinnaiyan AM (2005) Mining for regulatory programs in the cancer transcriptome. Nat Genet 37(6):579–583. https://doi.org/10.1038/ng1578
Long J, Wang A, Bai Y et al (2019) Development and validation of a TP53-associated immune prognostic model for hepatocellular carcinoma. EBioMedicine 42:363–374. https://doi.org/10.1016/j.ebiom.2019.03.022
Funding
This project was supported by the National Science Foundation of China (Nos. 81170442, 81470899), National Scholarship Foundation (No. 201208505116), and Outstanding young talent fund of the second hospital of CQMU (2011).
Author information
Authors and Affiliations
Contributions
WDG, ZXP and LZJ performed research and wrote the first draft. WDG and LZJ collected and analyzed the data. WDG, YY and LZJ participated in manuscript revision. All authors contributed to the design and interpretation of the study and to further drafts. LZJ is the guarantor.
Corresponding author
Ethics declarations
Conflict of interest
No benefits in any form have been received or will be received from a commercial party related directly or indirectly to the subject of this article.
Ethical approval
This article does not contain any studies with animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wen, Dg., Zhao, Xp., You, Y. et al. Systematic analysis of immune-related genes based on a combination of multiple databases to build a diagnostic and a prognostic risk model for hepatocellular carcinoma. Cancer Immunol Immunother 70, 773–786 (2021). https://doi.org/10.1007/s00262-020-02733-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-020-02733-2