Abstract
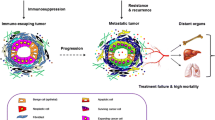
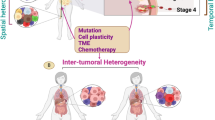
Tumors are highly heterogeneous tissues where malignant cells are surrounded by and interact with a complex tumor microenvironment (TME), notably composed of a wide variety of immune cells, as well as vessels and fibroblasts. As the dialectical influence between tumor cells and their TME is known to be clinically crucial, we need tools that allow us to study the cellular composition of the microenvironment. In this focused research review, we report MCP-counter, a methodology based on transcriptomic markers that assesses the proportion of several immune and stromal cell populations in the TME from transcriptomic data, and we highlight how it can provide a way to decipher the complex mechanisms at play in tumors. In several malignancies, MCP-counter scores have been used to show various prognostic impacts of the TME, which we also show to be linked with the mutational burden of tumors. We also compared established molecular classifications of colorectal cancer and clear-cell renal cell carcinoma with the output of MCP-counter, and show that molecular subgroups have different TME profiles, and that these profiles are consistent within a given subgroup. Finally, we provide insights as to how knowing the TME composition may shape patient care in the near future.



Similar content being viewed by others
Abbreviations
- ccRCC:
-
Clear-cell renal cell carcinoma
- CIBERSORT:
-
Cell-type identification by estimating relative subsets of RNA transcripts
- CMS:
-
Consensus molecular subgroups
- CRC:
-
Colorectal cancer
- GSEA:
-
Gene set enrichment analysis
- MCP-counter:
-
Microenvironment cell population-counter
- MSI:
-
Microsatellite instable
- NGS:
-
Next-generation sequencing
- NIBIT:
-
Network Italiano per la Bioterapia dei Tumori (the Italian network for cancer biotherapy)
- TCGA PANCAN:
-
The cancer genome atlas pan-cancer cohort
- Th17:
-
T helper lymphocyte producing interleukin 17
- TME:
-
Tumor microenvironment
- Treg:
-
Regulatory T cell
References
Becht E, Giraldo NA, Germain C et al (2016) Immune contexture, immunoscore, and malignant cell molecular subgroups for prognostic and theranostic classifications of cancers. Adv Immunol 130:95–190. doi:10.1016/bs.ai.2015.12.002
Fridman WH, Pagès F, Sautès-Fridman C, Galon J (2012) The immune contexture in human tumours: impact on clinical outcome. Nat Rev Cancer 12:298–306. doi:10.1038/nrc3245
Hanahan D, Coussens LM (2012) Accessories to the crime: functions of cells recruited to the tumor microenvironment. Cancer Cell 21:309–322. doi:10.1016/j.ccr.2012.02.022
Becht E, Giraldo NA, Dieu-Nosjean M-C et al (2016) Cancer immune contexture and immunotherapy. Curr Opin Immunol 39:7–13. doi:10.1016/j.coi.2015.11.009
Pagès F, Berger A, Camus M et al (2005) Effector memory T cells, early metastasis, and survival in colorectal cancer. N Engl J Med 353:2654–2666. doi:10.1056/NEJMoa051424
Remark R, Alifano M, Cremer I et al (2013) Characteristics and clinical impacts of the immune environments in colorectal and renal cell carcinoma lung metastases: influence of tumor origin. Clin Cancer Res 19:4079–4091
Postow MA, Callahan MK, Wolchok JD (2015) Immune checkpoint blockade in cancer therapy. J Clin Oncol 33:1974–1982. doi:10.1200/JCO.2014.59.4358
Kontermann RE, Brinkmann U (2015) Bispecific antibodies. Drug Discov Today 20:838–847. doi:10.1016/j.drudis.2015.02.008
Guo Y, Wang Y, Han W (2016) Chimeric antigen receptor-modified T cells for solid tumors: challenges and prospects. J Immunol Res 2016:3850839. doi:10.1155/2016/3850839
Galon J, Costes A, Sanchez-Cabo F et al (2006) Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science 313:1960–1964. doi:10.1126/science.1129139
Giraldo NA, Becht E, Pagès F et al (2015) Orchestration and prognostic significance of immune checkpoints in the microenvironment of primary and metastatic renal cell cancer. Clin Cancer Res 21:3031–3040. doi:10.1158/1078-0432.CCR-14-2926
Granier C, Dariane C, Combe P et al (2017) Tim-3 Expression on tumor-infiltrating PD-1(+)CD8(+) T cells correlates with poor clinical outcome in renal cell carcinoma. Cancer Res 77:1075–1082. doi:10.1158/0008-5472.CAN-16-0274
Shen-Orr SS, Gaujoux R (2013) Computational deconvolution: extracting cell type-specific information from heterogeneous samples. Curr Opin Immunol 25:571–578. doi:10.1016/j.coi.2013.09.015
Venet D, Pecasse F, Maenhaut C, Bersini H (2001) Separation of samples into their constituents using gene expression data. Bioinformatics 17:S279–S287. doi:10.1093/bioinformatics/17.suppl_1.S279
Abbas AR, Wolslegel K, Seshasayee D et al (2009) Deconvolution of blood microarray data identifies cellular activation patterns in systemic lupus erythematosus. PLoS One 4:e6098. doi:10.1371/journal.pone.0006098
Bindea G, Mlecnik B, Tosolini M et al (2013) Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity 39:782–795. doi:10.1016/j.immuni.2013.10.003
Şenbabaoğlu Y, Gejman RS, Winer AG et al (2016) Tumor immune microenvironment characterization in clear cell renal cell carcinoma identifies prognostic and immunotherapeutically relevant messenger RNA signatures. Genome Biol 17:231. doi:10.1186/s13059-016-1092-z
Charoentong P, Finotello F, Angelova M et al (2017) Pan-cancer immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep 18:248–262. doi:10.1016/j.celrep.2016.12.019
Newman AM, Liu CL, Green MR et al (2015) Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 12:453–457. doi:10.1038/nmeth.3337
Becht E, Giraldo NA, Lacroix L et al (2016) Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol 17:218. doi:10.1186/s13059-016-1070-5
Quon G, Haider S, Deshwar AG et al (2013) Computational purification of individual tumor gene expression profiles leads to significant improvements in prognostic prediction. Genome Med 5:29. doi:10.1186/gm433
Giraldo NA, Becht E, Vano Y et al (2017) Tumor-infiltrating and peripheral blood T cell immunophenotypes predict early relapse in localized clear cell renal cell carcinoma. Clin Cancer Res. doi:10.1158/1078-0432.CCR-16-2848
Choueiri TK, Motzer RJ (2017) Systemic therapy for metastatic renal-cell carcinoma. N Engl J Med 376:354–366. doi:10.1056/NEJMra1601333
Bauman TM, Huang W, Lee MH, Abel EJ (2016) Neovascularity as a prognostic marker in renal cell carcinoma. Hum Pathol 57:98–105. doi:10.1016/j.humpath.2016.07.005
Dieu-Nosjean M-C, Goc J, Giraldo NA et al (2014) Tertiary lymphoid structures in cancer and beyond. Trends Immunol 35:571–580. doi:10.1016/j.it.2014.09.006
Dieu-Nosjean M-C, Giraldo NA, Kaplon H et al (2016) Tertiary lymphoid structures, drivers of the anti-tumor responses in human cancers. Immunol Rev 271:260–275. doi:10.1111/imr.12405
Schumacher TN, Schreiber RD (2015) Neoantigens in cancer immunotherapy. Science 348:69–74. doi:10.1126/science.aaa4971
Alizadeh AA, Eisen MB, Davis RE et al (2000) Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 403:503–511. doi:10.1038/35000501
Ando T, Suguro M, Kobayashi T et al (2003) Multiple fuzzy neural network system for outcome prediction and classification of 220 lymphoma patients on the basis of molecular profiling. Cancer Sci 94:906–913
Guedj M, Marisa L, de Reynies A et al (2012) A refined molecular taxonomy of breast cancer. Oncogene 31:1196–1206. doi:10.1038/onc.2011.301
Perou CM, Sørlie T, Eisen MB et al (2000) Molecular portraits of human breast tumours. Nature 406:747–752. doi:10.1038/35021093
Guinney J, Dienstmann R, Wang X et al (2015) The consensus molecular subtypes of colorectal cancer. Nat Med 21:1350–1356. doi:10.1038/nm.3967
Becht E, de Reyniès A, Giraldo NA et al (2016) Immune and stromal classification of colorectal cancer is associated with molecular subtypes and relevant for precision immunotherapy. Clin Cancer Res 22:4057–4066. doi:10.1158/1078-0432.CCR-15-2879
Le DT, Uram JN, Wang H et al (2015) PD-1 blockade in tumors with mismatch-repair deficiency. N Engl J Med 372:2509–2520. doi:10.1056/NEJMoa1500596
Dienstmann R, Vermeulen L, Guinney J et al (2017) Consensus molecular subtypes and the evolution of precision medicine in colorectal cancer. Nat Rev Cancer 17:79–92. doi:10.1038/nrc.2016.126
Beuselinck B, Job S, Becht E et al (2015) Molecular subtypes of clear cell renal cell carcinoma are associated with sunitinib response in the metastatic setting. Clin Cancer Res 21:1329–1339. doi:10.1158/1078-0432.CCR-14-1128
Becht E, Giraldo NA, Beuselinck B et al (2015) Prognostic and theranostic impact of molecular subtypes and immune classifications in renal cell cancer (RCC) and colorectal cancer (CRC). Oncoimmunology 4:e1049804. doi:10.1080/2162402X.2015.1049804
Acknowledgements
We wish to thank Benoît Beuselinck, Laetitia Lacroix, Pierre Laurent-Puig, Stéphane Oudard, Jessica Zucman-Rossi, and all other members of the teams who contributed to the data.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This work was supported by the Institut National de la santé et de la Recherche Medicale (INSERM), University Paris-Descartes, University Pierre and Marie Curie, the Site de Recherche Integrée sur le Cancer (SIRIC) Cancer Research for Personalized Medicine (CARPEM) program, the LabEx Immuno-Oncology (LAXE62_9UMRS972 FRIDMAN), the Institut National Du Cancer (INCa), and the Cancéropôle Ile-de-France, O. Lecomte. Florent Petitprez is recipient of a CARPEM fellowship.
Conflict of interest
Yann A. Vano has received speaker honoraria from Novartis, Pfizer, Bristol-Myers-Squibb, and Astellas Sanofi, and has received financial support for attending symposia from Bristol-Myers-Squibb and Novartis. Aurélien de Reyniès holds intellectual property rights for patents related to immune cell population abundance estimation through transcriptomic analysis. Wolf H. Fridman is a consultant for Pierre Fabre Medicament, Sanofi, Bristol-Myers-Squibb, Novartis, Curetech, Servier, Efranet, Efralys, and Adaptimmune. All other authors declare no conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Petitprez, F., Vano, Y.A., Becht, E. et al. Transcriptomic analysis of the tumor microenvironment to guide prognosis and immunotherapies. Cancer Immunol Immunother 67, 981–988 (2018). https://doi.org/10.1007/s00262-017-2058-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-017-2058-z