Abstract
Purpose
Alzheimer’s disease (AD) is a heterogeneous disease that presents a broad spectrum of clinicopathologic profiles. To date, objective subtyping of AD independent of disease progression using brain imaging has been required. Our study aimed to extract representations of unique brain metabolism patterns different from disease progression to identify objective subtypes of AD.
Methods
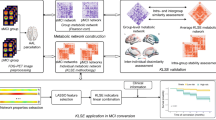
A total of 3620 FDG brain PET images with AD, mild cognitive impairment (MCI), and cognitively normal (CN) were obtained from the ADNI database from 1607 participants at enrollment and follow-up visits. A conditional variational autoencoder model was trained on FDG brain PET images of AD patients with the corresponding condition of AD severity score. The k-means algorithm was applied to generate clusters from the encoded representations. The trained deep learning-based cluster model was also transferred to FDG PET of MCI patients and predicted the prognosis of subtypes for conversion from MCI to AD. Spatial metabolism patterns, clinical and biological characteristics, and conversion rate from MCI to AD were compared across the subtypes.
Results
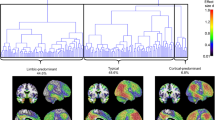
Four distinct subtypes of spatial metabolism patterns in AD with different brain pathologies and clinical profiles were identified: (i) angular, (ii) occipital, (iii) orbitofrontal, and (iv) minimal hypometabolic patterns. The deep learning model was also successfully transferred for subtyping MCI, and significant differences in frequency (P < 0.001) and risk of conversion (log-rank P < 0.0001) from MCI to AD were observed across the subtypes, highest in S2 (35.7%) followed by S1 (23.4%).
Conclusion
We identified distinct subtypes of AD with different clinicopathologic features. The deep learning-based approach to distinguish AD subtypes on FDG PET could have implications for predicting individual outcomes and provide a clue to understanding the heterogeneous pathophysiology of AD.





Similar content being viewed by others
Data availability
All raw data including FDG PET, CDR-SB score, demographic, cognitive, and biomarker variables are available through the ADNI data archive (http://adni.loni.usc.edu/). The custom codes for the deep learning model may be available for research purposes from the corresponding authors on reasonable request.
References
Lam B, Masellis M, Freedman M, Stuss DT, Black SE. Clinical, imaging, and pathological heterogeneity of the Alzheimer’s disease syndrome. Alzheimers Res Ther. 2013;5:1. https://doi.org/10.1186/alzrt155.
Friedland RP, Koss E, Haxby JV, Grady CL, Luxenberg J, Schapiro MB, et al. NIH conference. Alzheimer disease: clinical and biological heterogeneity. Ann Intern Med. 1988;109:298–311. https://doi.org/10.7326/0003-4819-109-4-298.
Noh Y, Jeon S, Lee JM, Seo SW, Kim GH, Cho H, et al. Anatomical heterogeneity of Alzheimer disease: based on cortical thickness on MRIs. Neurology. 2014;83:1936–44. https://doi.org/10.1212/WNL.0000000000001003.
Dujardin S, Commins C, Lathuiliere A, Beerepoot P, Fernandes AR, Kamath TV, et al. Tau molecular diversity contributes to clinical heterogeneity in Alzheimer’s disease. Nat Med. 2020;26:1256–63. https://doi.org/10.1038/s41591-020-0938-9.
Ferreira D, Nordberg A, Westman E. Biological subtypes of Alzheimer disease: a systematic review and meta-analysis. Neurology. 2020;94:436–48. https://doi.org/10.1212/wnl.0000000000009058.
Vogel JW, Young AL, Oxtoby NP, Smith R, Ossenkoppele R, Strandberg OT, et al. Four distinct trajectories of tau deposition identified in Alzheimer’s disease. Nat Med. 2021;27:871–81. https://doi.org/10.1038/s41591-021-01309-6.
Ferreira D, Verhagen C, Hernández-Cabrera JA, Cavallin L, Guo C-J, Ekman U, et al. Distinct subtypes of Alzheimer’s disease based on patterns of brain atrophy: longitudinal trajectories and clinical applications. Sci Rep. 2017;7:1–13. https://doi.org/10.1038/srep46263.
Mosconi L, Tsui WH, Herholz K, Pupi A, Drzezga A, Lucignani G, et al. Multicenter standardized 18F-FDG PET diagnosis of mild cognitive impairment, Alzheimer’s disease, and other dementias. J Nucl Med. 2008;49:390–8. https://doi.org/10.2967/jnumed.107.045385.
Meyer PT, Frings L, Rücker G, Hellwig S. 18F-FDG PET in parkinsonism: differential diagnosis and evaluation of cognitive impairment. J Nucl Med. 2017;58:1888–98. https://doi.org/10.2967/jnumed.116.186403.
Herholz K, Westwood S, Haense C, Dunn G. Evaluation of a calibrated 18F-FDG PET score as a biomarker for progression in Alzheimer disease and mild cognitive impairment. J Nucl Med. 2011;52:1218–26. https://doi.org/10.2967/jnumed.111.090902.
Landau S, Harvey D, Madison C, Reiman E, Foster N, Aisen P, et al. Comparing predictors of conversion and decline in mild cognitive impairment. Neurology. 2010;75:230–8. https://doi.org/10.1212/WNL.0b013e3181e8e8b8.
Besson FL, La Joie R, Doeuvre L, Gaubert M, Mézenge F, Egret S, et al. Cognitive and brain profiles associated with current neuroimaging biomarkers of preclinical Alzheimer’s disease. J Neurosci. 2015;35:10402–11. https://doi.org/10.1523/JNEUROSCI.0150-15.2015.
Laforce R Jr, Soucy J-P, Sellami L, Dallaire-Théroux C, Brunet F, Bergeron D, et al. Molecular imaging in dementia: past, present, and future. Alzheimers Dement. 2018;14:1522–52. https://doi.org/10.1016/j.jalz.2018.06.2855.
Vanhoutte M, Semah F, Sillaire AR, Jaillard A, Petyt G, Kuchcinski G, et al. 18F-FDG PET hypometabolism patterns reflect clinical heterogeneity in sporadic forms of early-onset Alzheimer’s disease. Neurobiol Aging. 2017;59:184–96. https://doi.org/10.1016/j.neurobiolaging.2017.08.009.
Levin F, Ferreira D, Lange C, Dyrba M, Westman E, Buchert R, et al. Data-driven FDG-PET subtypes of Alzheimer’s disease-related neurodegeneration. Alzheimers Res Ther. 2021;13:49. https://doi.org/10.1186/s13195-021-00785-9.
Groot C, Risacher SL, Chen JQA, Dicks E, Saykin AJ, Mac Donald CL, et al. Differential trajectories of hypometabolism across cognitively-defined Alzheimer’s disease subgroups. Neuroimage Clin. 2021;31:102725. https://doi.org/10.1016/j.nicl.2021.102725.
Jo T, Nho K, Saykin AJ. Deep learning in Alzheimer’s disease: diagnostic classification and prognostic prediction using neuroimaging data. Front Aging Neurosci. 2019;11:220. https://doi.org/10.3389/fnagi.2019.00220.
Zhao Z, Chuah JH, Lai KW, Chow CO, Gochoo M, Dhanalakshmi S, et al. Conventional machine learning and deep learning in Alzheimer’s disease diagnosis using neuroimaging: a review. Front Comput Neurosci. 2023;17:1038636. https://doi.org/10.3389/fncom.2023.1038636.
Young AL, Marinescu RV, Oxtoby NP, Bocchetta M, Yong K, Firth NC, et al. Uncovering the heterogeneity and temporal complexity of neurodegenerative diseases with subtype and stage inference. Nat Commun. 2018;9:4273. https://doi.org/10.1038/s41467-018-05892-0.
Kingma DP, Welling M. Auto-encoding variational bayes. arXiv preprint arXiv:1312.6114, 2013.
Kingma DP, Mohamed S, Rezende DJ, Welling M. Semi-supervised learning with deep generative models. arXiv preprint arXiv:1406.5298, 2014.
Jagust WJ, Landau SM, Koeppe RA, Reiman EM, Chen K, Mathis CA, et al. The Alzheimer’s disease neuroimaging initiative 2 PET core: 2015. Alzheimers Dement. 2015;11:757–71. https://doi.org/10.1016/j.jalz.2015.05.001.
Hughes CP, Berg L, Danziger W, Coben LA, Martin RL. A new clinical scale for the staging of dementia. Br J Psychiatry. 1982;140:566–72. https://doi.org/10.1192/bjp.140.6.566.
Choi H, Kang H, Lee DS. Predicting aging of brain metabolic topography using variational autoencoder. Front Aging Neurosci. 2018;10:212. https://doi.org/10.3389/fnagi.2018.00212.
Lloyd S. Least squares quantization in PCM. IEEE Trans Inf Theory. 1982;28:129–37. https://doi.org/10.1109/TIT.1982.1056489.
MacQueen J. Some methods for classification and analysis of multivariate observations. Proceedings of the 5th Berkeley symposium on mathematical statistics and probability. 1967:281-97.
Kodinariya TM, Makwana PR. Review on determining number of Cluster in K-Means Clustering. Int J Adv Res Comput Sci Manage Stud. 2013;1:90–5.
Marutho D, Handaka SH, Wijaya E. The determination of cluster number at k-mean using elbow method and purity evaluation on headline news. Proceedings of the 2018 International Seminar on Application for Technology of Information and Communication. 2018:533-8.
Mukherjee S, Choi SE, Lee ML, Scollard P, Trittschuh EH, Mez J, et al. Cognitive domain harmonization and cocalibration in studies of older adults. Neuropsychology. 2023;37:409–23. https://doi.org/10.1037/neu0000835.
Sohn K, Lee H, Yan X. Learning structured output representation using deep conditional generative models. Proceedings of the 28th International Conference on Neural Information Processing Systems. 2015:3483-91.
Misra C, Fan Y, Davatzikos C. Baseline and longitudinal patterns of brain atrophy in MCI patients, and their use in prediction of short-term conversion to AD: results from ADNI. Neuroimage. 2009;44:1415–22. https://doi.org/10.1016/j.neuroimage.2008.10.031.
Davatzikos C, Bhatt P, Shaw LM, Batmanghelich KN, Trojanowski JQ. Prediction of MCI to AD conversion, via MRI, CSF biomarkers, and pattern classification. Neurobiol Aging. 2011;32:2322.e19-e27. https://doi.org/10.1016/j.neurobiolaging.2010.05.023.
Cheng B, Liu M, Zhang D, Munsell BC, Shen D. Domain transfer learning for MCI conversion prediction. IEEE Trans Biomed Eng. 2015;62:1805–17. https://doi.org/10.1109/TBME.2015.2404809.
Kwak K, Giovanello KS, Bozoki A, Styner M, Dayan E. Subtyping of mild cognitive impairment using a deep learning model based on brain atrophy patterns. Cell Rep Med. 2021;2:100467. https://doi.org/10.1016/j.xcrm.2021.100467.
Chen P, Yao H, Tijms BM, Wang P, Wang D, Song C, et al. Four distinct subtypes of Alzheimer’s disease based on resting-state connectivity biomarkers. Biol Psychiatry. 2023;93:759–69. https://doi.org/10.1016/j.biopsych.2022.06.019.
Iaccarino L, Sala A, Perani D. Predicting long-term clinical stability in amyloid-positive subjects by FDG-PET. Ann Clin Transl Neurol. 2019;6:1113–20. https://doi.org/10.1002/acn3.782.
van der Flier WM, Pijnenburg YA, Fox NC, Scheltens P. Early-onset versus late-onset Alzheimer’s disease: the case of the missing APOE ɛ4 allele. Lancet Neurol. 2011;10:280–8. https://doi.org/10.1016/s1474-4422(10)70306-9.
Rogalski E, Sridhar J, Rader B, Martersteck A, Chen K, Cobia D, et al. Aphasic variant of Alzheimer disease: clinical, anatomic, and genetic features. Neurology. 2016;87:1337–43. https://doi.org/10.1212/wnl.0000000000003165.
Schott JM, Crutch SJ, Carrasquillo MM, Uphill J, Shakespeare TJ, Ryan NS, et al. Genetic risk factors for the posterior cortical atrophy variant of Alzheimer’s disease. Alzheimers Dement. 2016;12:862–71. https://doi.org/10.1016/j.jalz.2016.01.010.
Petersen RC. Mild cognitive impairment as a diagnostic entity. J Intern Med. 2004;256:183–94. https://doi.org/10.1111/j.1365-2796.2004.01388.x.
Jack CR Jr, Knopman DS, Weigand SD, Wiste HJ, Vemuri P, Lowe V, et al. An operational approach to National Institute on Aging-Alzheimer’s Association criteria for preclinical Alzheimer disease. Ann Neurol. 2012;71:765–75. https://doi.org/10.1002/ana.22628.
Wisse LEM, Butala N, Das SR, Davatzikos C, Dickerson BC, Vaishnavi SN, et al. Suspected non-AD pathology in mild cognitive impairment. Neurobiol Aging. 2015;36:3152–62. https://doi.org/10.1016/j.neurobiolaging.2015.08.029.
Jack CR Jr, Knopman DS, Chételat G, Dickson D, Fagan AM, Frisoni GB, et al. Suspected non-Alzheimer disease pathophysiology–concept and controversy. Nat Rev Neurol. 2016;12:117–24. https://doi.org/10.1038/nrneurol.2015.251.
Acknowledgements
This paper is based on the first author’s Ph.D. thesis in the Graduate School of Convergence Science and Technology, and College of Medicine or College of Pharmacy, Seoul National University, Seoul, Republic of Korea.
Funding
This research was supported by the National Research Foundation of Korea grant funded by the Korea Government (NRF-2019K1A3A1A14065446), and Korea Medical Device Development Fund grant funded by the Korea government (the Ministry of Science and ICT, the Ministry of Trade, Industry and Energy, the Ministry of Health & Welfare, the Ministry of Food and Drug Safety) (Project Number: 1711137868, RS-2020-KD000006).
Data collection and sharing for this project was funded by the Alzheimer's Disease Neuroimaging Initiative (ADNI) (National Institutes of Health Grant U01 AG024904) and DOD ADNI (Department of Defense award number W81XWH-12-2-0012). ADNI is funded by the National Institute on Aging, the National Institute of Biomedical Imaging and Bioengineering, and through generous contributions from the following: AbbVie, Alzheimer’s Association; Alzheimer’s Drug Discovery Foundation; Araclon Biotech; BioClinica, Inc.; Biogen; Bristol-Myers Squibb Company; CereSpir, Inc.; Cogstate; Eisai Inc.; Elan Pharmaceuticals, Inc.; Eli Lilly and Company; EuroImmun; F. Hoffmann-La Roche Ltd and its affiliated company Genentech, Inc.; Fujirebio; GE Healthcare; IXICO Ltd.; Janssen Alzheimer Immunotherapy Research & Development, LLC.; Johnson & Johnson Pharmaceutical Research & Development LLC.; Lumosity; Lundbeck; Merck & Co., Inc.; Meso Scale Diagnostics, LLC.; NeuroRx Research; Neurotrack Technologies; Novartis Pharmaceuticals Corporation; Pfizer Inc.; Piramal Imaging; Servier; Takeda Pharmaceutical Company; and Transition Therapeutics. The Canadian Institutes of Health Research is providing funds to support ADNI clinical sites in Canada. Private sector contributions are facilitated by the Foundation for the National Institutes of Health (www.fnih.org). The grantee organization is the Northern California Institute for Research and Education, and the study is coordinated by the Alzheimer’s Therapeutic Research Institute at the University of Southern California. ADNI data are disseminated by the Laboratory for Neuro Imaging at the University of Southern California.
Author information
Authors and Affiliations
Consortia
Contributions
H.C. designed the study. H.G.R. and H.C. developed the model. H.G.R. performed experiments. A.R. and D.S.L. contributed to analyze PET images. D.Y.L. and D.S.L. contributed to analyze clinical data. K.S. supported developing the model. K.S., A.R., D.Y.L., and D.S.L. contributed to data interpretation and analysis. H.G.R. wrote the manuscript mainly and all authors critically reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Ethics approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Informed consent
Written informed consent for cognitive testing and neuroimaging prior to participation of the ADNI cohort was obtained from all subjects, and the study protocols were approved by the institutional review boards of all participating institutions.
Competing interests
Dr. Choi is a co-founder of Portrai, Inc. All other authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Data used in preparation of this article were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (adni.loni.usc.edu). As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in analysis or writing of this report. A complete listing of ADNI investigators can be found at http://adni.loni.usc.edu/wp-content/uploads/how_to_apply/ADNI_Acknowledgement_List.pdf.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ryoo, H.G., Choi, H., Shi, K. et al. Distinct subtypes of spatial brain metabolism patterns in Alzheimer’s disease identified by deep learning-based FDG PET clusters. Eur J Nucl Med Mol Imaging 51, 443–454 (2024). https://doi.org/10.1007/s00259-023-06440-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-023-06440-9