Abstract
Purpose
The role that gut microbiota plays in determining the efficacy of the anti-tumor effect of immune checkpoint inhibitors is gaining increasing attention, and fecal bacterial transplantation has been recognized as a promising strategy for improving or rescuing the effect of immune checkpoint inhibition. However, techniques for the precise monitoring of in vivo bacterial behaviors after transplantation are limited. In this study, we aimed to use metabolic labeling and subsequent positron emission tomography (PET) imaging to track the in vivo behaviors of gut bacteria that are responsible for the efficacy of anti-PD-1 therapy in living mice.
Methods
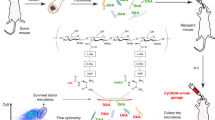
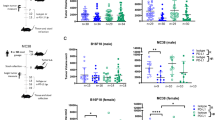
The antitumor effect of anti-PD-1 blockade was tested in a low-response 4T1 syngeneic mouse model with or without fecal transplantation and with or without broad-spectrum antibiotic imipenem treatment. High-throughput sequencing analyses of 16S rRNA gene amplicons in feces of 4T1 tumor-bearing mice pre- and post-anti-PD-1 treatment were performed. The identified bacteria, Bacteroides fragilis (B. fragilis), were labeled with 64Cu and fluorescence dye by the metabolic labeling of N3 followed by click chemistry. In vivo PET and optical imaging of B. fragilis were performed in mice after oral gavage.
Results
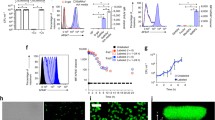
The disturbance of gut microbiota reduced the efficacy of anti-PD-1 treatment, and the combination of B. fragilis gavage and PD-1 blockade was beneficial in rescuing the antitumor effect of anti-PD-1 therapy. Metabolic oligosaccharide engineering and biorthogonal click chemistry resulted in successful B. fragilis labeling with 64Cu and fluorescence dye with high in vitro and in vivo stability and no effect on viability. PET imaging successfully detected the in vivo behaviors of B. fragilis after transplantation.
Conclusion
PET tracking by metabolic labeling is a powerful, noninvasive tool for the real-time tracking and quantitative imaging of gut microbiota. This strategy is clinically translatable and may also be extended to the PET tracking of other functional cells to guide cell-based adoptive therapies.






Similar content being viewed by others
References
Syn NL, Teng MWL, Mok TSK, Soo RA. De-novo and acquired resistance to immune checkpoint targeting. Lancet Oncol. 2017;18:e731–e41.
Chen DS, Mellman I. Oncology meets immunology: the cancer-immunity cycle. Immunity. 2013;39:1–10.
Rosenberg SA. Raising the bar: the curative potential of human cancer immunotherapy. Sci Transl Med. 2012;4:127ps8.
Nishino M, Ramaiya NH, Hatabu H, Hodi FS. Monitoring immune-checkpoint blockade: response evaluation and biomarker development. Nat Rev Clin Oncol. 2017;14:655–68.
Maleki Vareki S, Garrigos C, Duran I. Biomarkers of response to PD-1/PD-L1 inhibition. Crit Rev Oncol Hematol. 2017;116:116–24.
Heskamp S, Hobo W, Molkenboer-Kuenen JD, Olive D, Oyen WJ, Dolstra H, et al. Noninvasive imaging of tumor PD-L1 expression using radiolabeled anti-PD-L1 antibodies. Cancer Res. 2015;75:2928–36.
Josefsson A, Nedrow JR, Park S, Banerjee SR, Rittenbach A, Jammes F, et al. Imaging, biodistribution, and dosimetry of radionuclide-labeled PD-L1 antibody in an immunocompetent mouse model of breast cancer. Cancer Res. 2016;76:472–9.
Levi J, Lam T, Goth SR, Yaghoubi S, Bates J, Ren G, et al. Imaging of activated T cells as an early predictor of immune response to anti-PD-1 therapy. Cancer Res. 2019;79:3455–65.
Gibney GT, Weiner LM, Atkins MB. Predictive biomarkers for checkpoint inhibitor-based immunotherapy. Lancet Oncol. 2016;17:e542–e51.
Vetizou M, Pitt JM, Daillere R, Lepage P, Waldschmitt N, Flament C, et al. Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science. 2015;350:1079–84.
Sivan A, Corrales L, Hubert N, Williams JB, Aquino-Michaels K, Earley ZM, et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science. 2015;350:1084–9.
Routy B, Le Chatelier E, Derosa L, Duong CPM, Alou MT, Daillere R, et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science. 2018;359:91–7.
Gopalakrishnan V, Spencer CN, Nezi L, Reuben A, Andrews MC, Karpinets TV, et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science. 2018;359:97–103.
Aso Y, Akazan H. Prophylactic effect of a Lactobacillus casei preparation on the recurrence of superficial bladder cancer. BLP Study Group Urol Int. 1992;49:125–9.
Aso Y, Akaza H, Kotake T, Tsukamoto T, Imai K, Naito S. Preventive effect of a Lactobacillus casei preparation on the recurrence of superficial bladder cancer in a double-blind trial. The BLP Study Group Eur Urol. 1995;27:104–9.
Lenoir M, Del Carmen S, Cortes-Perez N, Lozano-Ojalvo D, Muñoz-Provencio D, Chain F, et al. Lactobacillus casei BL23 regulates Treg and Th17 T-cell populations and reduces DMH-associated colorectal cancer. J Gastroenterol. 2016;51:862–73.
Li J, Sung C, Lee N, Ni Y, Pihlajamäki J, Panagiotou G, et al. Probiotics modulated gut microbiota suppresses hepatocellular carcinoma growth in mice. Proc Natl Acad Sci U S A. 2016;113:E1306–15.
Campbell CT, Sampathkumar SG, Yarema KJ. Metabolic oligosaccharide engineering: perspectives, applications, and future directions. Mol BioSyst. 2007;3:187–94.
Prescher JA, Bertozzi CR. Chemistry in living systems. Nat Chem Biol. 2005;1:13–21.
Sletten EM, Bertozzi CR. Bioorthogonal chemistry: fishing for selectivity in a sea of functionality. Angew Chem Int Ed Engl. 2009;48:6974–98.
Geva-Zatorsky N, Alvarez D, Hudak JE, Reading NC, Erturk-Hasdemir D, Dasgupta S, et al. In vivo imaging and tracking of host-microbiota interactions via metabolic labeling of gut anaerobic bacteria. Nat Med. 2015;21:1091–100.
Weissleder R. Molecular imaging: exploring the next frontier. Radiology. 1999;212:609–14.
Margolis DJ, Hoffman JM, Herfkens RJ, Jeffrey RB, Quon A, Gambhir SS. Molecular imaging techniques in body imaging. Radiology. 2007;245:333–56.
Gao D, Gao L, Zhang C, Liu H, Jia B, Zhu Z, et al. A near-infrared phthalocyanine dye-labeled agent for integrin alphavbeta6-targeted theranostics of pancreatic cancer. Biomaterials. 2015;53:229–38.
Wexler HM. Bacteroides: the good, the bad, and the nitty-gritty. Clin Microbiol Rev. 2007;20:593–621.
Iida N, Dzutsev A, Stewart CA, Smith L, Bouladoux N, Weingarten RA, et al. Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science. 2013;342:967–70.
Weinstein EA, Ordonez AA, DeMarco VP, Murawski AM, Pokkali S, MacDonald EM, et al. Imaging Enterobacteriaceae infection in vivo with 18F-fluorodeoxysorbitol positron emission tomography. Sci Transl Med. 2014;6:259ra146.
Wang W, Zhu Y, Chen X. Selective imaging of gram-negative and gram-positive microbiotas in the mouse gut. Biochemistry. 2017;56:3889–93.
Weissleder R. Scaling down imaging: molecular mapping of cancer in mice. Nat Rev Cancer. 2002;2:11–8.
Negrin RS, Contag CH. In vivo imaging using bioluminescence: a tool for probing graft-versus-host disease. Nat Rev Immunol. 2006;6:484–90.
Berger C, Flowers ME, Warren EH, Riddell SR. Analysis of transgene-specific immune responses that limit the in vivo persistence of adoptively transferred HSV-TK-modified donor T cells after allogeneic hematopoietic cell transplantation. Blood. 2006;107:2294–302.
Acknowledgments
We thank Prof. Changtao Jiang at Peking University Health Science Center for kindly providing the Bacteroides fragilis and also thank Miss Xuemei Wang for her technical assistance with the culture of Bacteroides fragilis. In addition, we thank the cyclotron teams of the Department of Nuclear Medicine, Peking University Cancer Hospital and Institute for 64Cu production.
Funding
This work was supported, in part, by the National Key R&D Program of China (2018YFC1313300), National Natural Science Foundation of China (81873907, 81671747, and 81920108020), Beijing Nova Program Interdisciplinary Cooperation Project (Z181100006218136), Beijing Natural Science Foundation (JQ19026 and L172007), and the Clinical Medicine Plus X-Young Scholars Project of Peking University (PKU2019LCXQ023).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed. This article does not contain any studies with human participants performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Preclinical Imaging.
Electronic supplementary material
ESM 1
(DOCX 1088 kb).
Rights and permissions
About this article
Cite this article
Wang, Y., Zhang, C., Lai, J. et al. Noninvasive PET tracking of post-transplant gut microbiota in living mice. Eur J Nucl Med Mol Imaging 47, 991–1002 (2020). https://doi.org/10.1007/s00259-019-04639-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-019-04639-3