Abstract
Purpose
Although functional brain imaging has been used for the early and objective assessment of cognitive dysfunction, there is a lack of generalized image-based biomarker which can evaluate individual’s cognitive dysfunction in various disorders. To this end, we developed a deep learning-based cognitive signature of FDG brain PET adaptable for Parkinson’s disease (PD) as well as Alzheimer’s disease (AD).
Methods
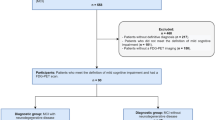
A deep learning model for discriminating AD from normal controls (NCs) was built by a training set consisting of 636 FDG PET obtained from Alzheimer’s Disease Neuroimaging Initiative database. The model was directly transferred to images of mild cognitive impairment (MCI) patients (n = 666) for identifying who would rapidly convert to AD and another independent cohort consisting of 62 PD patients to differentiate PD patients with dementia. The model accuracy was measured by area under curve (AUC) of receiver operating characteristic (ROC) analysis. The relationship between all images was visualized by two-dimensional projection of the deep learning-based features. The model was also designed to predict cognitive score of the subjects and validated in PD patients. Cognitive dysfunction-related regions were visualized by feature maps of the deep CNN model.
Results
AUC of ROC for differentiating AD from NC was 0.94 (95% CI 0.89–0.98). The transfer of the model could differentiate MCI patients who would convert to AD (AUC = 0.82) and PD with dementia (AUC = 0.81). The two-dimensional projection mapping visualized the degree of cognitive dysfunction compared with normal brains regardless of different disease cohorts. Predicted cognitive score, an output of the model, was highly correlated with the mini-mental status exam scores. Individual cognitive dysfunction-related regions included cingulate and high frontoparietal cortices, while they showed individual variability.
Conclusion
The deep learning-based cognitive function evaluation model could be successfully transferred to multiple disease domains. We suggest that this approach might be extended to an objective cognitive signature that provides quantitative biomarker for cognitive dysfunction across various neurodegenerative disorders.





Similar content being viewed by others
Data availability
The imaging data can be found in ADNI database (http://adni.loni.usc.edu/). The application of our method is developed for a web-based resource (https://fdgbrainpet.appspot.com/).
References
Chaudhuri KR, Healy DG, Schapira AH. Non-motor symptoms of Parkinson’s disease: diagnosis and management. Lancet Neurol. 2006;5(3):235–45.
Emre M, et al. Clinical diagnostic criteria for dementia associated with Parkinson’s disease. Mov Disord. 2007;22(12):1689–707.
McKinlay A, et al. A profile of neuropsychiatric problems and their relationship to quality of life for Parkinson’s disease patients without dementia. Parkinsonism Relat Disord. 2008;14(1):37–42.
Schrag A, Jahanshahi M, Quinn N. What contributes to quality of life in patients with Parkinson’s disease? J Neurol Neurosurg Psychiatry. 2000;69(3):308–12.
Ravina B, et al. The role of radiotracer imaging in Parkinson disease. Neurology. 2005;64(2):208–15.
Brooks DJ. Imaging approaches to Parkinson disease. J Nucl Med. 2010;51(4):596–609.
Eidelberg D. Metabolic brain networks in neurodegenerative disorders: a functional imaging approach. Trends Neurosci. 2009;32(10):548–57.
Eckert T, et al. FDG PET in the differential diagnosis of parkinsonian disorders. Neuroimage. 2005;26(3):912–21.
Huang C, et al. Metabolic abnormalities associated with mild cognitive impairment in Parkinson disease. Neurology. 2008;70(16 Part 2):1470–7.
Huang C, et al. Metabolic brain networks associated with cognitive function in Parkinson’s disease. Neuroimage. 2007;34(2):714–23.
Svenningsson P, Westman E, Ballard C, Aarsland D. Cognitive impairment in patients with Parkinson’s disease: diagnosis, biomarkers, and treatment. Lancet Neurol. 2012;11(8):697–707.
Choi H. Deep learning in nuclear medicine and molecular imaging: current perspectives and future directions. Nucl Med Mol Imaging. 2017:1–10.
Oquab M, Bottou L, Laptev I, & Sivic J. Learning and transferring mid-level image representations using convolutional neural networks. Proceedings of the IEEE conference on computer vision and pattern recognition. 2014; pp 1717–1724.
Yosinski J, Clune J, Bengio Y, & Lipson H. How transferable are features in deep neural networks? Advances in neural information processing systems. 2014 pp 3320–3328.
Gibb W, Lees A. The relevance of the Lewy body to the pathogenesis of idiopathic Parkinson’s disease. J Neurol Neurosurg Psychiatry. 1988;51(6):745–52.
Jagust WJ, et al. The Alzheimer’s disease neuroimaging initiative 2 PET Core: 2015. Alzheimer’s & dementia: the journal of the Alzheimer’s Association. 2015;11(7):757–71.
Kingma D & Ba J. Adam: a method for stochastic optimization. arXiv preprint arXiv. 2014;1412.6980.
van der Maaten L. Learning a parametric embedding by preserving local structure. RBM. 2009;500(500):26.
Zhou B, Khosla A, Lapedriza A, Oliva A, & Torralba A. Learning deep features for discriminative localization. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. 2016; pp 2921–2929.
Zintgraf LM, Cohen TS, Adel T, & Welling M. Visualizing deep neural network decisions: prediction difference analysis. arXiv preprint arXiv. 2017;1702.04595.
Rajpurkar P, et al. Chexnet: radiologist-level pneumonia detection on chest x-rays with deep learning. arXiv preprint arXiv. 2017;1711.05225.
Aarsland D, Andersen K, Larsen JP, Lolk A. Prevalence and characteristics of dementia in Parkinson disease: an 8-year prospective study. Arch Neurol. 2003;60(3):387–92.
Schroeter ML, Stein T, Maslowski N, Neumann J. Neural correlates of Alzheimer’s disease and mild cognitive impairment: a systematic and quantitative meta-analysis involving 1351 patients. Neuroimage. 2009;47(4):1196–206.
Bohnen NI, et al. Cerebral glucose metabolic features of Parkinson disease and incident dementia: longitudinal study. J Nucl Med. 2011;52(6):848–55.
Greenspan H, van Ginneken B, Summers RM. Guest editorial deep learning in medical imaging: overview and future promise of an exciting new technique. IEEE Trans Med Imaging. 2016;35(5):1153–9.
Tajbakhsh N, et al. Convolutional neural networks for medical image analysis: full training or fine tuning? IEEE Trans Med Imaging. 2016;35(5):1299–312.
Gulshan V, et al. Development and validation of a deep learning algorithm for detection of diabetic retinopathy in retinal fundus photographs. Jama. 2016;316(22):2402–10.
Esteva A, et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature. 2017;542(7639):115.
Coudray N, et al. Classification and mutation prediction from non–small cell lung cancer histopathology images using deep learning. Nat Med. 2018:1.
Cheng B, Zhang D, & Shen D. Domain transfer learning for MCI conversion prediction. Medical Image Computing and Computer-Assisted Intervention–MICCAI. 2012;2012:82–90
Choi H, Jin KH, Initiative AsDN. Predicting cognitive decline with deep learning of brain metabolism and amyloid imaging. Behav Brain Res. 2018;344:103–9.
Choi H, et al. Deep learning only by normal brain PET identify unheralded brain anomalies. EBioMedicine. 2019. 2019;43:447-53.
Funding
This work was supported by a clinical research grant-in-aid from the Seoul Metropolitan Government Seoul National University Boramae Medical Center (02-2017-5) and National Research Foundation grant funded by the Ministry of Education, Science, and Technology in Korea (NRF-2016R1D1A1B03936159, NRF-2018R1C1B3008971, and NRF-2018R1A5A2025964).
Data collection and sharing for this project was funded by the Alzheimer’s Disease Neuroimaging Initiative (ADNI) (National Institutes of Health Grant U01 AG024904) and DOD ADNI (Department of Defense award number W81XWH-12-2-0012). ADNI is funded by the National Institute on Aging, the National Institute of Biomedical Imaging and Bioengineering, and through generous contributions from the following: AbbVie, Alzheimer’s Association; Alzheimer’s Drug Discovery Foundation; Araclon Biotech; BioClinica, Inc.; Biogen; Bristol-Myers Squibb Company; CereSpir, Inc.;Eisai Inc.; Elan Pharmaceuticals, Inc.; Eli Lilly and Company; EuroImmun; F. Hoffmann-La Roche Ltd. and its affiliated company Genentech, Inc.; Fujirebio; GE Healthcare; IXICO Ltd.; Janssen Alzheimer Immunotherapy Research & Development, LLC.; Johnson & Johnson Pharmaceutical Research & Development LLC.; Lumosity; Lundbeck; Merck & Co., Inc.; Meso Scale Diagnostics, LLC.; NeuroRx Research; Neurotrack Technologies; Novartis Pharmaceuticals Corporation; Pfizer Inc.; Piramal Imaging; Servier; Takeda Pharmaceutical Company; and Transition Therapeutics. The Canadian Institutes of Health Research is providing funds to support ADNI clinical sites in Canada. Private sector contributions are facilitated by the Foundation for the National Institutes of Health (www.fnih.org). The grantee organization is the Northern California Institute for Research and Education, and the study is coordinated by the Alzheimer’s Disease Cooperative Study at the University of California, San Diego. ADNI data are disseminated by the Laboratory for Neuro Imaging at the University of Southern California.
Author information
Authors and Affiliations
Consortia
Contributions
Y.K.K., C.H. and J.Y.L. designed the study. C.H. developed the deep learning model and analyzed the data. Y.K.K. and E.J.Y. collected the data. J.Y.K. performed clinical assessment and supported the analysis. D.S.L supervised the study. All authors interpreted data results, drafted, and edited manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent to clinical testing and neuroimaging prior to participation of the ADNI cohort was obtained, approved by the institutional review boards (IRB) of all participating institutions. The Institutional Review Board of Seoul National University Boramae Hospital approved this study and informed consents were waived for a retrospective cohort of Parkinson’s disease patients.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the topical collection on Advanced Image Analyses (Radiomics and Artificial Intelligence)
Data used in preparation of this article were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (adni.loni.usc.edu). As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in analysis or writing of this report. A complete listing of ADNI investigators can be found at: http://adni.loni.usc.edu/wp-content/uploads/how_to_apply/ADNI_Acknowledgement_List.pdf
Electronic supplementary material
ESM 1
(DOCX 432 kb)
Rights and permissions
About this article
Cite this article
Choi, H., Kim, Y.K., Yoon, E.J. et al. Cognitive signature of brain FDG PET based on deep learning: domain transfer from Alzheimer’s disease to Parkinson’s disease. Eur J Nucl Med Mol Imaging 47, 403–412 (2020). https://doi.org/10.1007/s00259-019-04538-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-019-04538-7