Abstract
Objective
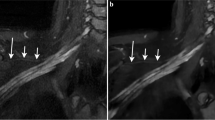
To evaluate whether ‘fast,’ unilateral, brachial plexus, 3D magnetic resonance neurography (MRN) acquisitions with deep learning reconstruction (DLR) provide similar image quality to longer, ‘standard’ scans without DLR.
Materials and methods
An IRB-approved prospective cohort of 30 subjects (13F; mean age = 50.3 ± 17.8y) underwent clinical brachial plexus 3.0 T MRN with 3D oblique-coronal STIR-T2-weighted-FSE. ‘Standard’ and ‘fast’ scans (time reduction = 23–48%, mean = 33%) were reconstructed without and with DLR. Evaluation of signal-to-noise ratio (SNR) and edge sharpness was performed for 4 image stacks: ‘standard non-DLR,’ ‘standard DLR,’ ‘fast non-DLR,’ and ‘fast DLR.’ Three raters qualitatively evaluated ‘standard non-DLR’ and ‘fast DLR’ for i) bulk motion (4-point scale), ii) nerve conspicuity of proximal and distal suprascapular and axillary nerves (5-point scale), and iii) nerve signal intensity, size, architecture, and presence of a mass (binary). ANOVA or Wilcoxon signed rank test compared differences. Gwet’s agreement coefficient (AC2) assessed inter-rater agreement.
Results
Quantitative SNR and edge sharpness were superior for DLR versus non-DLR (SNR by + 4.57 to + 6.56 [p < 0.001] for ‘standard’ and + 4.26 to + 4.37 [p < 0.001] for ‘fast;’ sharpness by + 0.23 to + 0.52/pixel for ‘standard’ [p < 0.018] and + 0.21 to + 0.25/pixel for ‘fast’ [p < 0.003]) and similar between ‘standard non-DLR’ and ‘fast DLR’ (SNR: p = 0.436–1, sharpness: p = 0.067–1). Qualitatively, ‘standard non-DLR’ and ‘fast DLR’ had similar motion artifact, as well as nerve conspicuity, signal intensity, size and morphology, with high inter-rater agreement (AC2: ‘standard’ = 0.70–0.98, ‘fast DLR’ = 0.69–0.97).
Conclusion
DLR applied to faster, 3D MRN acquisitions provides similar image quality to standard scans. A faster, DL-enabled protocol may replace currently optimized non-DL protocols.




Similar content being viewed by others
Data Availability
The raw data used to support the findings of this study are available upon reasonable request from the corresponding author.
References
Sun S, Tan ET, Mintz DN, et al. Evaluation of deep learning reconstructed high-resolution 3D lumbar spine MRI. Eur Radiol. 2022;32(9):6167–77. https://doi.org/10.1007/s00330-022-08708-4.
Sahr M, Tan ET, Sneag DB. 3D MRI of the Spine. Semin Musculoskelet Radiol. 2021;25(3):433–40. https://doi.org/10.1055/s-0041-1731060.
Davidson EJ, Tan ET, Pedrick EG, Sneag DB. Brachial Plexus Magnetic Resonance Neurography: Technical Challenges and Solutions. Invest Radiol. 2023;58(1):14–27. https://doi.org/10.1097/RLI.0000000000000906.
Sneag DB, Mendapara P, Zhu JC, et al. Prospective respiratory triggering improves high-resolution brachial plexus MRI quality. J Magn Reson Imaging. 2019;49(6):1723–9. https://doi.org/10.1002/jmri.26559.
Chaudhari AS, Fang Z, Kogan F, et al. Super-resolution musculoskeletal MRI using deep learning. Magn Reson Med. 2018;80(5):2139–54. https://doi.org/10.1002/mrm.27178.
Balsiger F, Steindel C, Arn M, et al. Segmentation of Peripheral Nerves From Magnetic Resonance Neurography: A Fully-Automatic, Deep Learning-Based Approach. Front Neurol. 2018;19(9):777. https://doi.org/10.3389/fneur.2018.00777.
Mori R, Kassai Y, Masuda A, et al. Ultrashort echo time time-spatial labeling inversion pulse magnetic resonance angiography with denoising deep learning reconstruction for the assessment of abdominal visceral arteries. J Magn Reson Imaging. 2021;53(6):1926–37. https://doi.org/10.1002/jmri.27481.
Kang H, Witanto JN, Pratama K, et al. Fully Automated MRI Segmentation and Volumetric Measurement of Intracranial Meningioma Using Deep Learning. J Magn Reson Imaging. 2023;57(3):871–81. https://doi.org/10.1002/jmri.28332.
van der Velde N, Hassing HC, Bakker BJ, et al. Improvement of late gadolinium enhancement image quality using a deep learning-based reconstruction algorithm and its influence on myocardial scar quantification. Eur Radiol. 2021;31(6):3846–55. https://doi.org/10.1007/s00330-020-07461-w.
Kidoh M, Shinoda K, Kitajima M, et al. Deep Learning Based Noise Reduction for Brain MR Imaging: Tests on Phantoms and Healthy Volunteers. Magn Reson Med Sci. 2020;19(3):195–206. https://doi.org/10.2463/mrms.mp.2019-0018.
Hammernik K, Klatzer T, Kobler E, et al. Learning a variational network for reconstruction of accelerated MRI data. Magn Reson Med. 2018;79(6):3055–71. https://doi.org/10.1002/mrm.26977.
Lebel RM. Performance characterization of a novel deep learning-based MR image reconstruction pipeline. ArXiv 2020;abs/2008.06559. https://api.semanticscholar.org/CorpusID:221139899
Zochowski KC, Tan ET, Argentieri EC, et al. Improvement of peripheral nerve visualization using a deep learning-based MR reconstruction algorithm. Magn Reson Imaging. 2022;85:186–92. https://doi.org/10.1016/j.mri.2021.10.038.
Chazen JL, Tan ET, Fiore J, Nguyen JT, Sun S, Sneag DB. Rapid lumbar MRI protocol using 3D imaging and deep learning reconstruction. Skeletal Radiol. 2023;52(7):1331–8. https://doi.org/10.1007/s00256-022-04268-2.
Jardon M, Tan ET, Chazen JL, Sahr M, Wen Y, Schneider B, Sneag DB. Deep-learning-reconstructed high-resolution 3D cervical spine MRI for foraminal stenosis evaluation. Skeletal Radiol. 2023;52(4):725–32. https://doi.org/10.1007/s00256-022-04211-5.
Sneag DB, Daniels SP, Geannette C, et al. Post-Contrast 3D Inversion Recovery Magnetic Resonance Neurography for Evaluation of Branch Nerves of the Brachial Plexus. Eur J Radiol. 2020;132:109304. https://doi.org/10.1016/j.ejrad.2020.109304.
R Development Core Team. R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2021.
Ensle F, Kaniewska M, Tiessen A, Lohezic M, Getzmann JM, Guggenberger R. Diagnostic performance of deep learning-based reconstruction algorithm in 3D MR neurography. Skeletal Radiol. 2023. https://doi.org/10.1007/s00256-023-04362-z.
Yoon D, Antil N, Biswal S, Lutz AM. A robust 3D fast spin-echo technique for fast examination of the brachial plexus. Skeletal Radiol. 2022;51(9):1865–72. https://doi.org/10.1007/s00256-022-04021-9.
Oudeman J, Coolen BF, Mazzoli V, et al. Diffusion-prepared neurography of the brachial plexus with a large field-of-view at 3T. J Magn Reson Imaging. 2016;43(3):644–54. https://doi.org/10.1002/jmri.25025.
Acknowledgements
The authors thank Shiv Kaushik and Maggie Fung for providing technical advice.
Funding
HSS receives institutional research support from GE Healthcare.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
D. B. Sneag, S. C. Queler joint first author.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sneag, D.B., Queler, S.C., Campbell, G. et al. Optimized 3D brachial plexus MR neurography using deep learning reconstruction. Skeletal Radiol 53, 779–789 (2024). https://doi.org/10.1007/s00256-023-04484-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00256-023-04484-4