Abstract
Objectives
To report the diagnostic performance of machine learning-based CT texture analysis for differentiating multiple myeloma from osteolytic metastatic bone lesions in the peripheral skeleton.
Methods
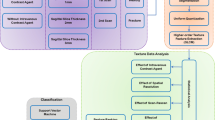
We retrospectively evaluated 172 patients with multiple myeloma (n = 70) and osteolytic metastatic bone lesions (n = 102) in the peripheral skeleton. Two radiologists individually used two-dimensional manual segmentation to extract texture features from non-contrast CT. In total, 762 radiomic features were extracted. Dimension reduction was performed in three stages: inter-observer agreement analysis, collinearity analysis, and feature selection. Data were randomly divided into training (n = 120) and test (n = 52) groups. Eight machine learning algorithms were used for model development. The primary performance metrics were the area under the receiver operating characteristic curve and accuracy.
Results
In total, 476 of the 762 texture features demonstrated excellent interobserver agreement. The number of features was reduced to 22 after excluding those with strong collinearity. Of these features, six were included in the machine learning algorithms using the wrapper-based classifier-specific technique. When all eight machine learning algorithms were considered for differentiating multiple myeloma from osteolytic metastatic bone lesions in the peripheral skeleton, the area under the receiver operating characteristic curve and accuracy were 0.776–0.932 and 78.8–92.3%, respectively. The k-nearest neighbors model performed the best, with the area under the receiver operating characteristic curve and accuracy values of 0.902 and 92.3%, respectively.
Conclusion
Machine learning-based CT texture analysis is a promising method for discriminating multiple myeloma from osteolytic metastatic bone lesions.



Similar content being viewed by others
References
Roodman GD. Pathogenesis of myeloma bone disease. J Cell Biochem. 2010;109:283–91.
Panaroni C, Yee AJ, Raje NS. Myeloma and bone disease. Curr Osteoporos Rep. 2017;15:483–98.
Hillengass J. Evolving concepts in the diagnosis and staging of multiple myeloma. J Natl Compr Cancer Netw. 2020;18:1770–2.
Roodman GD. Pathogenesis of myeloma bone disease. Leukemia. 2009;23:435–41.
Terpos E, Dimopoulos MA, Moulopoulos LA. The role of imaging in the treatment of patients with multiple myeloma in 2016. Am Soc Clin Oncol Educ B. 2016;36:407–17.
Macedo F, Ladeira K, Pinho F, et al. Bone metastases: an overview. Oncol Rev. 2017;11:321.
Roodman GD. Mechanisms of bone metastasis. N Engl J Med. 2004;15:1655–64.
Giuliani N, Rizzoli V, Roodman GD. Multiple myeloma bone disease: patho-physiology of osteoblast inhibition. Blood. 2006;108:3992–6.
Terpos E, Ntanasis-Stathopoulos I, Gavriatopoulou M, et al. Pathogenesis of bone disease in multiple myeloma: from bench to bedside. Blood Cancer J. 2018;8:7.
Bernstein ZS, Kim EB, Raje N. Bone disease in multiple myeloma: biologic and clinical implications. Cells. 2022;11(15):2308.
Clézardin P, Coleman R, Puppo M, et al. Bone metastasis: mechanisms, therapies, and biomarkers. Physiol Rev. 2021;101(3):797–855.
Chandra H, Chandra S, Verma SK. Multiple myeloma or metastatic carcinoma breast: diagnostic dilemma in a case presenting with lytic bony lesion. Breast Dis. 2015;35:199–201.
Tomono H, Fujioka S, Kato K, Yoshida K, Nimura Y. Multiple myeloma mimicking bone metastasis from breast cancer: report of a case. Surg Today. 1998;28:1304–6.
Hough B, Brufsky A, Lentzsch S. Metastatic breast cancer or multiple myeloma? Camouflage by lytic lesions. J Oncol. 2010;2010:7–9.
Kang YM, Lee HJ, Kim S-J. Metastatic breast cancer with osteolytic skull lesions suspected to be multiple myeloma. Korean J Clin Oncol. 2017;13:152–5.
Hsu Y-T, Chang K-C. Breast cancer with extensive bone metastasis mimicking myeloma. Clin Case Reports. 2017;5:203–4.
Zhang H, Lv J, Lv C, Zhang H. Presentation of multiple myeloma mimicking bone metastasis from colon adenocarcinoma: a case report and literature review. Mol Clin Oncol. 2016;4:31–4.
Merrild EH, Baerentzen S, Bouchelouche K, Buus S. Vertebral myeloma mimicking prostatic carcinoma metastasis in 68Ga-PSMA PET/CT. Clin Nucl Med. 2017;42:790–2.
Shah E, Azhar W, Saleem S. A trail to diagnosis-finding the primary lesions of bone metastasis. Cureus. e2022;44:1–6
Nagao T, Bando K, Iura A, et al. Malignant paraganglioma mimicking multiple myeloma. Rinsho Ketsueki Japan. 2022;63:1373–8.
Mutlu U, Balci A, Özsan GH, Özkal S, Şeyhanli A, Özgül HA. Computed tomography characteristics of multiple myeloma and other osteolytic metastatic bone lesions. Acta Radiol. 2021;62:1639–47.
Varghese BA, Cen SY, Hwang DH, et al. Texture analysis of imaging: what radiologists need to know. AJR Am J Roentgenol. 2019;212:520–8.
Gillies RJ, Kinahan PE, Hricak H. Radiomics: images are more than pictures, they are data. Radiology. 2016;278:563–77.
Wang S, Summers RM. Machine learning and radiology. Med Image Anal. 2012;16:933–51.
Kocak B, Kus EA, Yardimci AH, Bektas CT, Kilickesmez O. Machine learning in radiomic renal mass characterization: fundamentals, applications, challenges, and future directions. AJR Am J Roentgenol. 2020;215:920–8.
Gitto S, Cuocolo R, Albano D, et al. MRI radiomics-based machine-learning classification of bone chondrosarcoma. Eur J Radiol. 2020;128:109043.
Basara Akin I, Ozgul H, Simsek K, Altay C, Secil M, Balci P. Texture analysis of ultrasound images to differentiate simple fibroadenomas from complex fibroadenomas and benign phyllodes tumors. J Ultrasound Med. 2020;39:1993–2003.
Jin Z, Wang Y, Wang Y, Mao Y, Zhang F. Application of 18F-FDG PET-CT images based radiomics in identifying vertebral multiple myeloma and bone metastases. Front Med. 2022;9:1–12.
Baykara M, Yıldırım M. Differentiation of multiple myeloma and metastases with apparent diffusion coefficient map histogram analysis. North Clin Istanb. 2022;9:256–60.
Tagliafico AS, Cea M, Rossi F, et al. Differentiating diffuse from focal pattern on computed tomography in multiple myeloma: added value of a radiomics approach. Eur J Radiol. 2019;121:108739.
Park H, Lee S, Lee J, et al. Detecting multiple myeloma infiltration of the bone marrow on CT scans in patients with osteopenia : feasibility of radiomics analysis. Diagnostics. 2022;12:923.
Rajkumar SV, Dimopoulos MA, Palumbo A, et al. International myeloma working group updated criteria for the diagnosis of multiple myeloma. Lancet Oncol. 2014;15:e538–48.
Koçak B, Durmaz EŞ, Ateş E, Kılıçkesmez Ö. Radiomics with artificial intelligence: a practical guide for beginners. Diagn Interv Radiol. 2019;25:485–95.
Dormann CF, Elith J, Bacher S, et al. Collinearity: a review of methods to deal with it and a simulation study evaluating their performance. Ecography. 2013;36:27–46.
Bermejo P, Gamez JA, Puerta JM. Improving incremental wrapper-based subset selection via replacement and early stopping. Int J Pattern Recognit Artif Intell. 2011;25:605–25.
Frank E, Hall MA, Witten IH. The WEKA workbench. Online appendix for “data mining: practical machine learning tools and techniques”. Data Min Pract Mach Learn Tools Tech. 2016. https://www.cs.waikato.ac.nz/ml/weka/Witten_et_al_2016_appendix.pdf. Accessed 12 Dec 2022.
Mwangi B, Tian TS, Soares JC. A review of feature reduction techniques in neuroimaging. Neuroinformatics. 2014;12:229–44.
Varma S, Simon R. Bias in error estimation when using cross-validation for model selection. BMC Bioinformatics. 2006;7:91.
Rajkumar SV. Multiple myeloma: 2020 update on diagnosis, risk-stratification and management. Am J Hematol. 2020;95:548–67.
Deng C, Han D, Zhang X, Lv Z, Li D. Differential performances in lesions and radio-tracer of 18F-FDG PET/CT between multiple myeloma and unknown osteolytic metastasis. Curr Med Imaging. 2022. https://doi.org/10.2174/1573405618666220516120230.
Vicentini JRT, Bredella MA. Whole body imaging in musculoskeletal oncology: when, why, and how. Skeletal Radiol. 2022;52(3):281–95. https://doi.org/10.1007/s00256-022-04112-7.
Li X, Wu N, Zhang W, Liu Y, Ming Y. Differential diagnostic value of 18F-FDG PET/CT in osteolytic lesions. J Bone Oncol. 2020;24:100302.
Lecouvet FE, Van Nieuwenhove S, Jamar F, Lhommel R, Guermazi A, Pasoglou VP. Whole-body MR imaging: the novel, “intrinsically hybrid,” approach to metastases, myeloma, lymphoma, in bones and beyond. PET Clin. 2018;13:505–22.
Matteucci F, Paganelli G, Martinelli G, Cerchione C. PET/CT in multiple myeloma: beyond FDG. Front Oncol. 2021;10:622501.
Ganeshan B, Miles KA. Quantifying tumour heterogeneity with CT. Cancer Imaging. 2013;13:140–9.
Mutlu İN, Koçak B, Ateş Kuş E, Baykara Ulusan M, Kılıçkesmez Ö. Machine learning-based computed tomography texture analysis of lytic bone lesions needing biopsy: a preliminary study. Istanbul Med J. 2021;22:223–31.
Ng F, Kozarski R, Ganeshan B, Goh V. Assessment of tumor heterogeneity by CT texture analysis: can the largest cross-sectional area be used as an alternative to whole tumor analysis? Eur J Radiol. 2013;82:342–8.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The English in this document has been checked by at least two professional editors, both native speakers of English. For a certificate, please see: http://www.textcheck.com/certificate/DmpUp8.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Özgül, H.A., Akin, I.B., Mutlu, U. et al. Diagnostic value of machine learning-based computed tomography texture analysis for differentiating multiple myeloma from osteolytic metastatic bone lesions in the peripheral skeleton. Skeletal Radiol 52, 1703–1711 (2023). https://doi.org/10.1007/s00256-023-04333-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00256-023-04333-4