Abstract
Background
Whole-body low-dose CT is the recommended initial imaging modality to evaluate bone destruction as a result of multiple myeloma. Accurate interpretation of these scans to detect small lytic bone lesions is time intensive. A functional deep learning) algorithm to detect lytic lesions on CTs could improve the value of these CTs for myeloma imaging. Our objectives were to develop a DL algorithm and determine its performance at detecting lytic lesions of multiple myeloma.
Methods
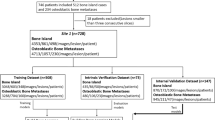
Axial slices (2-mm section thickness) from whole-body low-dose CT scans of subjects with biochemically confirmed plasma cell dyscrasias were included in the study. Data were split into train and test sets at the patient level targeting a 90%/10% split. Two musculoskeletal radiologists annotated lytic lesions on the images with bounding boxes. Subsequently, we developed a two-step deep learning model comprising bone segmentation followed by lesion detection. Unet and “You Look Only Once” (YOLO) models were used as bone segmentation and lesion detection algorithms, respectively. Diagnostic performance was determined using the area under the receiver operating characteristic curve (AUROC).
Results
Forty whole-body low-dose CTs from 40 subjects yielded 2193 image slices. A total of 5640 lytic lesions were annotated. The two-step model achieved a sensitivity of 91.6% and a specificity of 84.6%. Lesion detection AUROC was 90.4%.
Conclusion
We developed a deep learning model that detects lytic bone lesions of multiple myeloma on whole-body low-dose CTs with high performance. External validation is required prior to widespread adoption in clinical practice.








Similar content being viewed by others
Abbreviations
- IMWG:
-
International Myeloma Working Group
- CNN:
-
Convolutional neural network
- DL:
-
Deep learning
- DICOM:
-
Digital Imaging and Communications in Medicine
- PNG:
-
Portable Network Graphics
- YOLO:
-
You Only Look Once
- AUROC:
-
Area under the receiver operating characteristic curve
- DSC:
-
Dice similarity coefficient
- mAP:
-
Mean average precision
- SD:
-
Standard deviation
References
Durie BG, Salmon SE. A clinical staging system for multiple myeloma. Correlation of measured myeloma cell mass with presenting clinical features, response to treatment, and survival. Cancer. 1975;36:842–54.
Hillengass J, Usmani S, Rajkumar SV, Durie BGM, Mateos M-V, Lonial S, et al. International myeloma working group consensus recommendations on imaging in monoclonal plasma cell disorders. Lancet Oncol. 2019;20:e302–12.
Moulopoulos LA, Koutoulidis V, Hillengass J, Zamagni E, Aquerreta JD, Roche CL, et al. Recommendations for acquisition, interpretation and reporting of whole body low dose CT in patients with multiple myeloma and other plasma cell disorders: a report of the IMWG Bone Working Group. Blood Cancer J. 2018;8:95.
Horger M, Claussen CD, Bross-Bach U, Vonthein R, Trabold T, Heuschmid M, et al. Whole-body low-dose multidetector row-CT in the diagnosis of multiple myeloma: an alternative to conventional radiography. Eur J Radiol. 2005;54:289–97.
Gavriatopoulou M, Βoultadaki A, Koutoulidis V, Ntanasis-Stathopoulos I, Bourgioti C, Malandrakis P, et al. The role of low dose whole body CT in the detection of progression of patients with smoldering multiple myeloma. Blood Cancer J. 2020;10:93.
Akkus Z, Galimzianova A, Hoogi A, Rubin DL, Erickson BJ. Deep learning for brain MRI segmentation: state of the art and future directions. J Digit Imaging. 2017;30:449–59.
Weston AD, Korfiatis P, Kline TL, Philbrick KA, Kostandy P, Sakinis T, et al. Automated abdominal segmentation of CT scans for body composition analysis using deep learning. Radiology. 2019;290:669–79.
Kline TL, Korfiatis P, Edwards ME, Blais JD, Czerwiec FS, Harris PC, et al. Performance of an artificial multi-observer deep neural network for fully automated segmentation of polycystic kidneys. J Digit Imaging. 2017;30:442–8.
Korfiatis P, Kline TL, Erickson BJ. Automated segmentation of hyperintense regions in FLAIR MRI using deep learning. Tomography. 2016;2:334–40.
Qu R, Yang Y, Wang Y. COVID-19 detection using CT image based on YOLOv5 network [Internet]. arXiv [eess.IV]. 2022. Available from: http://arxiv.org/abs/2201.09972
Hossain A, Islam MT, Almutairi AF. A deep learning model to classify and detect brain abnormalities in portable microwave based imaging system. Sci Rep. 2022;12:6319.
Vyshnav MT, Sowmya V, Gopalakrishnan EA, Variyar V.V. S, Menon VK, Soman P K. Deep learning based approach for multiple myeloma detection. 2020 11th International Conference on Computing, Communication and Networking Technologies (ICCCNT). 2020. p. 1–7.
He J, Zhang K. Medical image analysis of multiple myeloma based on convolutional neural network. Expert Syst [Internet]. Wiley; 2022;39. Available from: https://onlinelibrary.wiley.com/doi/https://doi.org/10.1111/exsy.12810
Xu L, Tetteh G, Lipkova J, Zhao Y, Li H, Christ P, et al. Automated whole-body bone lesion detection for multiple myeloma on 68Ga-pentixafor PET/CT imaging using deep learning methods. Contrast Media Mol Imaging. 2018;2018:2391925.
Allegra A, Tonacci A, Sciaccotta R, Genovese S, Musolino C, Pioggia G, et al. Machine learning and deep learning applications in multiple myeloma diagnosis, prognosis, and treatment selection. Cancers [Internet]. 2022;14. Available from: http://dx.doi.org/https://doi.org/10.3390/cancers14030606
Faghani S, Codipilly DC, Vogelsang D, Moassefi M, Rouzrokh P, Khosravi B, et al. Development of a deep learning model for the histological diagnosis of dysplasia in Barrett’s esophagus. Gastrointest Endosc [Internet]. 2022; Available from: https://www.sciencedirect.com/science/article/pii/S0016510722017643
Ganaie MA, Hu M, Malik AK, Tanveer M, Suganthan PN. Ensemble deep learning: a review [Internet]. arXiv [cs.LG]. 2021. Available from: http://arxiv.org/abs/2104.02395
Ren M, Yi PH. Deep learning detection of subtle fractures using staged algorithms to mimic radiologist search pattern. Skeletal Radiol. 2022;51:345–53.
Franklin J. The elements of statistical learning: data mining, inference and prediction. Math Intelligencer. 2005;27:83–5.
Philbrick KA, Weston AD, Akkus Z, Kline TL, Korfiatis P, Sakinis T, et al. RIL-Contour: a medical imaging dataset annotation tool for and with deep learning. J Digit Imaging. 2019;32:571–81.
Russell BC, Torralba A, Murphy KP, Freeman WT. LabelMe: a database and web-based tool for image annotation. Int J Comput Vis. 2008;77:157–73.
MONAI Consortium. MONAI: medical open network for AI [Internet]. 2022. Available from: https://zenodo.org/record/6639453
Girshick R, Donahue J, Darrell T, Malik J. (2014) Rich feature hierarchies for accurate object detection and semantic segmentation. 2014 IEEE Conference on Computer Vision and Pattern Recognition. p. 580–7.
Redmon J, Divvala S, Girshick R, Farhadi A. You only look once: unified, real-time object detection [Internet]. arXiv [cs.CV]. 2015. Available from: http://arxiv.org/abs/1506.02640
Hansen LK, Salamon P. Neural network ensembles. IEEE Trans Pattern Anal Mach Intell. 1990;12:993–1001.
Pedregosa, Varoquaux, Gramfort, Michel, Thirion, Grisel, et al. Scikit-learn: machine learning in Python. J Mach Learn Res [Internet]. Available from: https://jmlr.csail.mit.edu/papers/v12/pedregosa11a.html
Yang S, Yin B, Cao W, Feng C, Fan G, He S. Diagnostic accuracy of deep learning in orthopaedic fractures: a systematic review and meta-analysis. Clin Radiol. 2020;75:713.e17-713.e28.
Chang CY, Buckless C, Yeh KJ, Torriani M. Automated detection and segmentation of sclerotic spinal lesions on body CTs using a deep convolutional neural network. Skeletal Radiol. 2022;51:391–9.
Hayashi D, Kompel AJ, Ventre J, Ducarouge A, Nguyen T, Regnard N-E, et al. Automated detection of acute appendicular skeletal fractures in pediatric patients using deep learning. Skeletal Radiol [Internet]. 2022; Available from: http://dx.doi.org/https://doi.org/10.1007/s00256-022-04070-0
Yan W, Shi H, He T, Chen J, Wang C, Liao A, et al. Employment of artificial intelligence based on routine laboratory results for the early diagnosis of multiple myeloma. Front Oncol. 2021;11:608191.
Xiong X, Wang J, Hu S, Dai Y, Zhang Y, Hu C. Differentiating between multiple myeloma and metastasis subtypes of lumbar vertebra lesions using machine learning-based radiomics. Front Oncol. 2021;11:601699.
Lin T-Y, Maire M, Belongie S, Bourdev L, Girshick R, Hays J, et al. Microsoft COCO: common objects in context [Internet]. arXiv [cs.CV]. 2014. Available from: http://arxiv.org/abs/1405.0312
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Conflicts of interest
Bradley Erickson: officer: FlowSIGMA, Inc. and Yunu, Inc. The other others declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Bradley J. Erickson is the senior author.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Faghani, S., Baffour, F.I., Ringler, M.D. et al. A deep learning algorithm for detecting lytic bone lesions of multiple myeloma on CT. Skeletal Radiol 52, 91–98 (2023). https://doi.org/10.1007/s00256-022-04160-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00256-022-04160-z