Abstract
Physcion is one of natural anthraquinones, registered as a novel plant-derived fungicide due to its excellent prevention of plant disease. However, the current production of physcion via plant extraction limits its yield promotion and application. Here, a pair of polyketide synthases (PKS) in emodin biosynthesis were used as probes to mining the potential O-methyltransferase (OMT) responsible for physcion biosynthesis. Further refinement using the phylogenetic analysis of the mined OMTs revealed a distinct OMT (AcOMT) with the ability of transferring a methyl group to C-6 hydroxyl of emodin to form physcion. Through introducing AcOMT, we successfully obtained the de novo production of physcion in Aspergillus nidulans. The physcion biosynthetic pathway was further rationally engineered by expressing the decarboxylase genes from different fungi. Finally, the titer of physcion reached to 64.6 mg/L in shake-flask fermentation through enhancing S-adenosylmethionine supply. Our work provides a native O-methyltransferase for physcion biosynthesis and lays the foundation for further improving the production of physcion via a sustainable route.
Key points
• Genome mining of the native O-methyltransferase responsible for physcion biosynthesis
• De novo biosynthesis of physcion in the engineered Aspergillus nidulans
• Providing an alternative way to produce plant-derived fungicide physcion
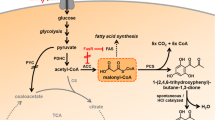
Graphical abstract







Similar content being viewed by others
References
Adnan M, Rasul A, Hussain G, Shah AM, Sarfraz I, Nageen B, Riaz A, Khalid R, Asrar M, Selamoglu Z, Adem Ş, Sarker DS (2021) Physcion and physcion 8-O-D-glucopyranoside: natural anthraquinones with potential anticancer activities. Curr Drug Targets 22(5):488–504. https://doi.org/10.2174/1389450121999201013154542
Awakawa T, Yokota K, Funa N, Doi F, Mori N, Watanabe H, Horinouchi S (2009) Physically discrete β-lactamase-type thioesterase catalyzes product release in atrochrysone synthesis by iterative type I polyketide synthase. Chem Biol 16(6):613–623. https://doi.org/10.1016/j.chembiol.2009.04.004
Bachmann M, Lüthy J, Schlatter C (1979) Toxicity and mutagenicity of molds of the Aspergillus glaucus group. identification of physcion and three related anthraquinones as main toxic constituents from Aspergillus chevalieri. J Agric Food Chem 27(6):1342–1347. https://doi.org/10.1021/jf60226a021
Cardenas J, Da Silva NA (2014) Metabolic engineering of Saccharomyces cerevisiae for the production of triacetic acid lactone. Metab Eng 25:194–203. https://doi.org/10.1016/j.ymben.2014.07.008
Chiang YM, Szewczyk E, Davidson AD, Entwistle R, Keller NP, Wang CC, Oakley BR (2010) Characterization of the Aspergillus nidulans monodictyphenone gene cluster. Appl Environ Microbiol 76(7):2067–2074. https://doi.org/10.1128/aem.02187-09
Chiang YM, Oakley CE, Ahuja M, Entwistle R, Schultz A, Chang SL, Sung CT, Wang CC, Oakley BR (2013) An efficient system for heterologous expression of secondary metabolite genes in Aspergillus nidulans. J Am Chem Soc 135(20):7720–7731. https://doi.org/10.1021/ja401945a
Chiang YM, Ahuja M, Oakley CE, Entwistle R, Asokan A, Zutz C, Wang CC, Oakley BR (2016) Development of genetic dereplication strains in Aspergillus nidulans results in the discovery of aspercryptin. Angew Chem Int Ed Eng 55(5):1662–1665. https://doi.org/10.1002/anie.201507097
Chiang Y-M, Lin T-S, Chang S-L, Ahn G, Wang CCC (2021) An Aspergillus nidulans platform for the complete cluster refactoring and total biosynthesis of fungal natural products. ACS Synth Biol 10(1):173–182. https://doi.org/10.1021/acssynbio.0c00536
Cravens A, Payne J, Smolke CD (2019) Synthetic biology strategies for microbial biosynthesis of plant natural products. Nat Commun 10(1):2142. https://doi.org/10.1038/s41467-019-09848-w
Du L, Zhu T, Liu H, Fang Y, Zhu W, Gu Q (2008) Cytotoxic polyketides from a marine-derived fungus Aspergillus glaucus. J Nat Prod 71(11):1837–1842. https://doi.org/10.1021/np800303t
Gibson DG, Young L, Chuang RY, Venter JC, Hutchison CA 3rd, Smith HO (2009) Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods 6(5):343–345. https://doi.org/10.1038/nmeth.1318
Hoefgen S, Lin J, Fricke J, Stroe MC, Mattern DJ, Kufs JE, Hortschansky P, Brakhage AA, Hoffmeister D, Valiante V (2018) Facile assembly and fluorescence-based screening method for heterologous expression of biosynthetic pathways in fungi. Metab Eng 48:44–51. https://doi.org/10.1016/j.ymben.2018.05.014
Jones EW (1991) Tackling the protease problem in Saccharomyces cerevisiae. Methods Enzymol 194:428–453. https://doi.org/10.1016/0076-6879(91)94034-a
Kotopka BJ, Li Y, Smolke CD (2018) Synthetic biology strategies toward heterologous phytochemical production. Nat Prod Rep 35(9):902–920. https://doi.org/10.1039/c8np00028j
Kouprina N, Larionov V (2016) Transformation-associated recombination (TAR) cloning for genomics studies and synthetic biology. Chromosoma 125(4):621–632. https://doi.org/10.1007/s00412-016-0588-3
Kozbial PZ, Mushegian AR (2005) Natural history of S-adenosylmethionine-binding proteins. BMC Struct Biol 5(1):19. https://doi.org/10.1186/1472-6807-5-19
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547–1549. https://doi.org/10.1093/molbev/msy096
Li S, Li Y, Smolke CD (2018) Strategies for microbial synthesis of high-value phytochemicals. Nat Chem 10(4):395–404. https://doi.org/10.1038/s41557-018-0013-z
Liscombe DK, Louie GV, Noel JP (2012) Architectures, mechanisms and molecular evolution of natural product methyltransferases. Nat Prod Rep 29(10):1238–1250. https://doi.org/10.1039/C2NP20029E
Ma X, Yang X, Zeng F, Yang L, Yu D, Ni H (2010) Physcion, a natural anthraquinone derivative, enhances the gene expression of leaf-specific thionin of barley against Blumeria graminis. Pest Manag Sci 66(7):718–724. https://doi.org/10.1002/ps.1933
Mirdita M, Schütze K, Moriwaki Y, Heo L, Ovchinnikov S, Steinegger M (2022) ColabFold: making protein folding accessible to all. Nat Methods 19(6):679–682. https://doi.org/10.1038/s41592-022-01488-1
Nayak T, Szewczyk E, Oakley CE, Osmani A, Ukil L, Murray SL, Hynes MJ, Osmani SA, Oakley BR (2006) A versatile and efficient gene-targeting system for Aspergillus nidulans. Genetics 172(3):1557–1566. https://doi.org/10.1534/genetics.105.052563
Nicholas KB, Nicholas HB, Deerfield DW (1997) GeneDoc: analysis and visualization of genetic variation. Embnew News 4:14
Okorafor IC, Chen M, Tang Y (2021) High-titer production of olivetolic acid and analogs in engineered fungal host using a nonplant biosynthetic pathway. ACS Synth Biol 10(9):2159–2166. https://doi.org/10.1021/acssynbio.1c00309
Qi F, Zhang W, Xue Y, Geng C, Jin Z, Li J, Guo Q, Huang X, Lu X (2022) Microbial production of the plant-derived fungicide physcion. Metab Eng 74:130–138. https://doi.org/10.1016/j.ymben.2022.10.007
Semwal RB, Semwal DK, Combrinck S, Viljoen A (2021) Emodin - a natural anthraquinone derivative with diverse pharmacological activities. Phytochemistry 190:112854. https://doi.org/10.1016/j.phytochem.2021.112854
Sun L, Liu G, Li Y, Jiang D, Guo W, Xu H, Zhan R (2019) Metabolic engineering of Saccharomyces cerevisiae for efficient production of endocrocin and emodin. Metab Eng 54:212–221. https://doi.org/10.1016/j.ymben.2019.04.008
Sun Q, Huang M, Wei Y (2021) Diversity of the reaction mechanisms of SAM-dependent enzymes. Acta Pharm Sin B 11(3):632–650. https://doi.org/10.1016/j.apsb.2020.08.011
Suntar I, Cetinkaya S, Haydaroglu US, Habtemariam S (2021) Bioproduction process of natural products and biopharmaceuticals: biotechnological aspects. Biotechnol Adv 50:107768. https://doi.org/10.1016/j.biotechadv.2021.107768
Ter-Hovhannisyan V, Lomsadze A, Chernoff YO, Borodovsky M (2008) Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training. Genome Res 18(12):1979–1990. https://doi.org/10.1101/gr.081612.108
Throckmorton K, Lim FY, Kontoyiannis DP, Zheng W, Keller NP (2016) Redundant synthesis of a conidial polyketide by two distinct secondary metabolite clusters in Aspergillus fumigatus. Environ Microbiol 18(1):246–259. https://doi.org/10.1111/1462-2920.13007
Upshall A (1986) Genetic and molecular characterization of argB+ transformants of Aspergillus nidulans. Curr Genet 10(8):593–599. https://doi.org/10.1007/BF00418126
Xu X, Liu L, Zhang F, Wang W, Li J, Guo L, Che Y, Liu G (2014) Identification of the first diphenyl ether gene cluster for pestheic acid biosynthesis in plant endophyte Pestalotiopsis fici. ChemBioChem 15(2):284–292. https://doi.org/10.1002/cbic.201300626
Xue Y, Liang Y, Zhang W, Geng C, Feng D, Huang X, Dong S, Zhang Y, Sun J, Qi F (2022) Characterization and structural analysis of emodin-O-methyltransferase from. Aspergillus terreus 70(18):5728–5737. https://doi.org/10.1021/acs.jafc.2c01281
XunLi LY, Chu S, Yang S, Peng Y, Ren S, Wen B, Chen N (2019) Physcion and physcion 8-O-β-glucopyranoside: a review of their pharmacology, toxicities and pharmacokinetics. Chem Biol Interact 310:108722. https://doi.org/10.1016/j.cbi.2019.06.035
Yang XJ, Yang LJ, Zeng FS, Xiang LB, Wang SN, Yu DZ, Ni H (2008) Distribution of baseline sensitivities to natural product physcion among isolates of Sphaerotheca fuliginea and Pseudoperonospora cubensis. Plant Dis 92(10):1451–1455. https://doi.org/10.1094/pdis-92-10-1451
Yao Y, Wang W, Shi W, Yan R, Zhang J, Wei G, Liu L, Che Y, An C, Gao S-S (2022) Overproduction of medicinal ergot alkaloids based on a fungal platform. Metab Eng 69:198–208. https://doi.org/10.1016/j.ymben.2021.12.002
Acknowledgements
This study was supported by the National Key Research and Development Program (2021YFC2100600) and the National Natural Science Foundation of China (grant no. 32001036).
Availability of data and materials
Data and materials described in this work are available from the authors upon reasonable request and availability.
Code availability
Not applicable
Author information
Authors and Affiliations
Contributions
YY and EY designed and performed main experiments and data analyses. YY performed data curation and writing of the original draft. YP helped to revise the manuscript and gave valuable advices on experimental design and data analyses. XS conducted experiments. GL did conceptualization and writing review and editing. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval
This study does not contain any studies with human participants or animals performed by any of the authors.
Consent to participate
Not applicable
Consent for publication
Not applicable
Conflict of interest
YY, YP, and GL are inventors on a provisional patent application related to this work (no. CN202211098075.9, filed 8 September 2022). All other authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1:
Tables S1-S7 and Figures S1-S7 (PDF 937 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yao, Y., Yang, E., Pan, Y. et al. Mining an O-methyltransferase for de novo biosynthesis of physcion in Aspergillus nidulans. Appl Microbiol Biotechnol 107, 1177–1188 (2023). https://doi.org/10.1007/s00253-023-12373-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-023-12373-y