Abstract
Cell proliferation is achieved through numerous enzyme reactions. Temperature governs the activity of each enzyme, ultimately determining the optimal growth temperature. The synthesis of useful chemicals and fuels utilizes a fraction of available metabolic pathways, primarily central metabolic pathways including glycolysis and the tricarboxylic acid cycle. However, it remains unclear whether the optimal temperature for these pathways is correlated with that for cell proliferation. Here, we found that wild-type Corynebacterium glutamicum displayed increased glycolytic activity under non-growing anaerobic conditions at 42.5 °C, at which cells do not proliferate under aerobic conditions. At this temperature, glucose consumption was not inhibited and increased by 28% compared with that at the optimal growth temperature of 30 °C. Transcriptional analysis revealed that a gene encoding glucose transporter (iolT2) was upregulated by 12.3-fold compared with that at 30 °C, with concomitant upregulation of NCgl2954 encoding the iolT2-regulating transcription factor. Deletion of iolT2 decreased glucose consumption rate at 42.5 °C by 28%. Complementation of iolT2 restored glucose consumption rate, highlighting the involvement of iolT2 in the accelerating glucose consumption at an elevated temperature. This study shows that the optimal temperature for glucose metabolism in C. glutamicum under anaerobic conditions differs greatly from that for cell growth under aerobic conditions, being beyond the upper limit of the growth temperature. This is beneficial for fuel and chemical production not only in terms of increasing productivity but also for saving cooling costs.
Key points
• C. glutamicum accelerated anaerobic glucose consumption at elevated temperature.
• The optimal temperature for glucose consumption was above the upper limit for growth.
• Gene expression involved in glucose transport was upregulated at elevated temperature.

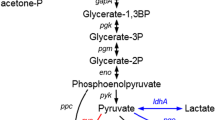
Graphical abstract





Similar content being viewed by others
References
Auchter M, Cramer A, Huser A, Ruckert C, Emer D, Schwarz P, Arndt A, Lange C, Kalinowski J, Wendisch VF, Eikmanns BJ (2011) RamA and RamB are global transcriptional regulators in Corynebacterium glutamicum and control genes for enzymes of the central metabolism. J Biotechnol 154(2–3):126–139. https://doi.org/10.1016/j.jbiotec.2010.07.001
Babul J (1978) Phosphofructokinases from Escherichia coli. Purification and characterization of the nonallosteric isozyme. J Biol Chem 253(12):4350–4355
Baldwin SA, Perham RN, Stribling D (1978) Purification and characterization of the class-II D-fructose 1,6-bisphosphate aldolase from Escherichia coli (Crookes’ strain). Biochem J 169(3):633–641
Baumgart M, Unthan S, Ruckert C, Sivalingam J, Grunberger A, Kalinowski J, Bott M, Noack S, Frunzke J (2013) Construction of a prophage-free variant of Corynebacterium glutamicum ATCC 13032 for use as a platform strain for basic research and industrial biotechnology. Appl Environ Microbiol 79(19):6006–6015. https://doi.org/10.1128/AEM.01634-13
Blombach B, Riester T, Wieschalka S, Ziert C, Youn JW, Wendisch VF, Eikmanns BJ (2011) Corynebacterium glutamicum tailored for efficient isobutanol production. Appl Environ Microbiol 77(10):3300–3310. https://doi.org/10.1128/AEM.02972-10
Brinkrolf K, Brune I, Tauch A (2006) Transcriptional regulation of catabolic pathways for aromatic compounds in Corynebacterium glutamicum. Genet Mol Res 5(4):773–789
Bussmann M, Emer D, Hasenbein S, Degraf S, Eikmanns BJ, Bott M (2009) Transcriptional control of the succinate dehydrogenase operon sdhCAB of Corynebacterium glutamicum by the cAMP-dependent regulator GlxR and the LuxR-type regulator RamA. J Biotechnol 143(3):173–182. https://doi.org/10.1016/j.jbiotec.2009.06.025
D’Alessio G, Josse J (1971) Glyceraldehyde phosphate dehydrogenase, phosphoglycerate kinase, and phosphoglyceromutase of Escherichia coli. Simultaneous purification and physical properties. J Biol Chem 246(13):4319–4325
Delaunay S, Gourdon P, Lapujade P, Mailly E, Oriol E, Engasser JM, Lindley ND, Goergen JL (1999) An improved temperature-triggered process for glutamate production with Corynebacterium glutamicum. Enzym Microb Technol 25:762–768
Delaunay S, Lapujade P, Engasser JM, Goergen JL (2002) Flexibility of the metabolism of Corynebacterium glutamicum 2262, a glutamic acid-producing bacterium, in response to temperature upshocks. J Mol Microbiol Biotechnol 28(6):333–337. https://doi.org/10.1038/sj/jim/7000251
Durfee T, Hansen AM, Zhi H, Blattner FR, Jin DJ (2008) Transcription profiling of the stringent response in Escherichia coli. J Bacteriol 190(3):1084–1096. https://doi.org/10.1128/JB.01092-07
Guisbert E, Yura T, Rhodius VA, Gross CA (2008) Convergence of molecular, modeling, and systems approaches for an understanding of the Escherichia coli heat shock response. Microbiol Mol Biol Rev 72(3):545–554. https://doi.org/10.1128/MMBR.00007-08
Hasegawa S, Suda M, Uematsu K, Natsuma Y, Hiraga K, Jojima T, Inui M, Yukawa H (2013) Engineering of Corynebacterium glutamicum for high-yield L-valine production under oxygen deprivation conditions. Appl Environ Microbiol 79(4):1250–1257. https://doi.org/10.1128/AEM.02806-12
Holtmann G, Brigulla M, Steil L, Schutz A, Barnekow K, Volker U, Bremer E (2004) RsbV-independent induction of the SigB-dependent general stress regulon of Bacillus subtilis during growth at high temperature. J Bacteriol 186(18):6150–6158. https://doi.org/10.1128/JB.186.18.6150-6158.2004
Inui M, Kawaguchi H, Murakami S, Vertès AA, Yukawa H (2004a) Metabolic engineering of Corynebacterium glutamicum for fuel ethanol production under oxygen-deprivation conditions. J Mol Microbiol Biotechnol 8(4):243–254. https://doi.org/10.1159/000086705
Inui M, Murakami S, Okino S, Kawaguchi H, Vertès AA, Yukawa H (2004b) Metabolic analysis of Corynebacterium glutamicum during lactate and succinate productions under oxygen deprivation conditions. J Mol Microbiol Biotechnol 7(4):182–196. https://doi.org/10.1159/000079827
Jojima T, Fujii M, Mori E, Inui M, Yukawa H (2010) Engineering of sugar metabolism of Corynebacterium glutamicum for production of amino acid L-alanine under oxygen deprivation. Appl Microbiol Biotechnol 87(1):159–165. https://doi.org/10.1007/s00253-010-2493-7
Jojima T, Noburyu R, Sasaki M, Tajima T, Suda M, Yukawa H, Inui M (2015) Metabolic engineering for improved production of ethanol by Corynebacterium glutamicum. Appl Microbiol Biotechnol 99(3):1165–1172. https://doi.org/10.1007/s00253-014-6223-4
Kimura E (2005) L-Glutamate production. In: Eggeling L, Bott M (eds) Handbook of Corynebacterium glutamicum. CRC Press, Boca Raton, pp 439–463
Kohl TA, Baumbach J, Jungwirth B, Puhler A, Tauch A (2008) The GlxR regulon of the amino acid producer Corynebacterium glutamicum: in silico and in vitro detection of DNA binding sites of a global transcription regulator. J Biotechnol 135(4):340–350. https://doi.org/10.1016/j.jbiotec.2008.05.011
Litsanov B, Brocker M, Bott M (2012) Toward homosuccinate fermentation: metabolic engineering of Corynebacterium glutamicum for anaerobic production of succinate from glucose and formate. Appl Environ Microbiol 78(9):3325–3337. https://doi.org/10.1128/AEM.07790-11
Matsumoto N, Hattori H, Matsutani M, Matayoshi C, Toyama H, Kataoka N, Yakushi T, Matsushita K (2018) A singlenucleotide insertion in a drug transporter geneinduces a thermotolerance phenotype in Gluconobacter frateurii by increasing the NADPH/NADP+ ratio via metabolic change. Appl Environ Microbiol 84(10):e00354–e00318. https://doi.org/10.1128/AEM.00354-18
Miller BG, Raines RT (2004) Identifying latent enzyme activities: substrate ambiguity within modern bacterial sugar kinases. Biochemistry 43(21):6387–6392. https://doi.org/10.1021/bi049424m
Muslin EH, Li D, Stevens FJ, Donnelly M, Schiffer M, Anderson LE (1995) Engineering a domain-locking disulfide into a bacterial malate dehydrogenase produces a redox-sensitive enzyme. Biophys J 68(6):2218–2223. https://doi.org/10.1016/S0006-3495(95)80430-3
Nazina TN, Tourova TP, Poltaraus AB, Novikova EV, Grigoryan AA, Ivanova AE, Lysenko AM, Petrunyaka VV, Osipov GA, Belyaev SS, Ivanov MV (2001) Taxonomic study of aerobic thermophilic bacilli: descriptions of Geobacillus subterraneus gen. nov., sp. nov. and Geobacillus uzenensis sp. nov. from petroleum reservoirs and transfer of Bacillus stearothermophilus, Bacillus thermocatenulatus, Bacillus thermoleovorans, Bacillus kaustophilus, Bacillus thermodenitrificans to Geobacillus as the new combinations G. stearothermophilus, G. th. Int J Syst Evol Microbiol 51(Pt 2):433–446. https://doi.org/10.1099/00207713-51-2-433
Ohnishi J, Hayashi M, Mitsuhashi S, Ikeda M (2003) Efficient 40 degrees C fermentation of L-lysine by a new Corynebacterium glutamicum mutant developed by genome breeding. Appl Microbiol Biotechnol 62(1):69–75. https://doi.org/10.1007/s00253-003-1254-2
Oide S, Gunji W, Moteki Y, Yamamoto S, Suda M, Jojima T, Yukawa H, Inui M (2015) Thermal and solvent stress cross-tolerance conferred to Corynebacterium glutamicum by adaptive laboratory evolution. Appl Environ Microbiol 81(7):2284–2298. https://doi.org/10.1128/AEM.03973-14
Okino S, Inui M, Yukawa H (2005) Production of organic acids by Corynebacterium glutamicum under oxygen deprivation. Appl Microbiol Biotechnol 68(4):475–480. https://doi.org/10.1007/s00253-005-1900-y
Okino S, Noburyu R, Suda M, Jojima T, Inui M, Yukawa H (2008) An efficient succinic acid production process in a metabolically engineered Corynebacterium glutamicum strain. Appl Microbiol Biotechnol 81(3):459–464. https://doi.org/10.1007/s00253-008-1668-y
Ozcan N, Ejsing CS, Shevchenko A, Lipski A, Morbach S, Kramer R (2007) Osmolality, temperature, and membrane lipid composition modulate the activity of betaine transporter BetP in Corynebacterium glutamicum. J Bacteriol 189(20):7485–7496. https://doi.org/10.1128/JB.00986-07
Patek M, Nesvera J (2011) Sigma factors and promoters in Corynebacterium glutamicum. J Biotechnol 154(2–3):101–113. https://doi.org/10.1016/j.jbiotec.2011.01.017
Sambrook J, Fritsh E, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press edn. Cold Spring Harbor
Schreyer R, Bock A (1980) Phosphoglucose isomerase from Escherischia coli K 10: purification, properties and formation under aerobic and anaerobic condition. Arch Microbiol 127(3):289–298
Toyoda K, Teramoto H, Inui M, Yukawa H (2009) Involvement of the LuxR-type transcriptional regulator RamA in regulation of expression of the gapA gene, encoding glyceraldehyde-3-phosphate dehydrogenase of Corynebacterium glutamicum. J Bacteriol 191(3):968–977. https://doi.org/10.1128/JB.01425-08
Tsuge Y, Yamamoto S, Suda M, Inui M, Yukawa H (2013) Reactions upstream of glycerate-1,3-bisphosphate drive Corynebacterium glutamicum (D)-lactate productivity under oxygen deprivation. Appl Microbiol Biotechnol 97(15):6693–6703. https://doi.org/10.1007/s00253-013-4986-7
Tsuge Y, Yamamoto S, Kato N, Suda M, Vertès AA, Yukawa H, Inui M (2015) Overexpression of the phosphofructokinase encoding gene is crucial for achieving high production of D-lactate in Corynebacterium glutamicum under oxygen deprivation. Appl Microbiol Biotechnol 99(11):4679–4689. https://doi.org/10.1007/s00253-015-6546-9
Vogt M, Brusseler C, Ooyen JV, Bott M, Marienhagen J (2016) Production of 2-methyl-1-butanol and 3-methyl-1-butanol in engineered Corynebacterium glutamicum. Metab Eng 38:436–445. https://doi.org/10.1016/j.ymben.2016.10.007
Wendisch VF (2003) Genome-wide expression analysis in Corynebacterium glutamicum using DNA microarrays. J Biotechnol 104:273–285. https://doi.org/10.1016/s0168-1656(03)00147-0
Wennerhold J, Krug A, Bott M (2005) The AraC-type regulator RipA represses aconitase and other iron proteins from Corynebacterium under iron limitation and is itself repressed by DtxR. J Biol Chem 280(49):40500–40508. https://doi.org/10.1074/jbc.M508693200
Wohl RC, Markus G (1972) Phosphoenolpyruvate carboxylase of Escherichia coli. Purification and some properties. J Biol Chem 247(18):5785–5792
Yamamoto S, Gunji W, Suzuki H, Toda H, Suda M, Jojima T, Inui M, Yukawa H (2012) Overexpression of genes encoding glycolytic enzymes in Corynebacterium glutamicum enhances glucose metabolism and alanine production under oxygen deprivation conditions. Appl Environ Microbiol 78(12):4447–4457. https://doi.org/10.1128/AEM.07998-11
Yamamoto S, Suda M, Niimi S, Inui M, Yukawa H (2013) Strain optimization for efficient isobutanol production using Corynebacterium glutamicum under oxygen deprivation. Biotechnol Bioeng 110(11):2938–2948. https://doi.org/10.1002/bit.24961
Zhang R, Zhang CT (2005) Genomic islands in the Corynebacterium efficiens genome. Appl Environ Microbiol 71(6):3126–3130. https://doi.org/10.1128/AEM.71.6.3126-3130.2005
Zhou J, Liu L, Shi Z, Du G, Chen J (2009) ATP in current biotechnology: regulation, applications and perspectives. Biotechnol Adv 27(1):94–101. https://doi.org/10.1016/j.biotechadv.2008.10.005
Funding
This work was supported by JSPS KAKENHI Grant Number JP17K14866 and Noda Institute for Scientific Research. This work was also partly supported by JST ALCA Grant Number JPMJAL1104, Japan.
Author information
Authors and Affiliations
Contributions
HU conducted anaerobic reaction and HPLC analysis. KT conducted DNA microarray analysis, qRT-PCR, and revised the manuscript. HMa constructed plasmids and strains. HMi conducted anaerobic reaction and HPLC analysis. KN, KT, and MI collaborated on the research. YT conceived and designed the research, conducted anaerobic reaction, and wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 165 kb)
Rights and permissions
About this article
Cite this article
Uchikura, H., Toyoda, K., Matsuzawa, H. et al. Anaerobic glucose consumption is accelerated at non-proliferating elevated temperatures through upregulation of a glucose transporter gene in Corynebacterium glutamicum. Appl Microbiol Biotechnol 104, 6719–6729 (2020). https://doi.org/10.1007/s00253-020-10739-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10739-0