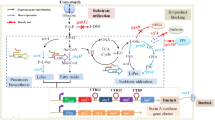
Abstract
The interspecies communication roles of γ-butyrolactones (GBLs) have been described for a long time but are still poorly understood. Herein, we analyzed more than 1000 Streptomyces strains and noticed a big quantitative gap between the strains with GBL biosynthetic genes and the strains with GBL receptor genes, which implies the wide-spread of GBLs as interspecies signals in Streptomyces and their great potential in the activation of silent natural product gene clusters. Streptomyces albidoflavus J1074, which has one GBL receptor gene but no GBL biosynthetic gene, was chosen as a target to study the possible interspecies communication roles of GBLs. At first, the GBL biosynthetic genes from Streptomyces coelicolor M145 were expressed in S. albidoflavus J1074, which enabled the S. albidoflavus strains to synthesize Streptomyces coelicolor butanolides (SCBs) and activated the production of paulomycins. Further studies showed that this activation process requires the participation of the GBL receptor gene XNR_4681. The results suggest that the expression of exogenous GBL biosynthetic genes can modulate the metabolisms of GBL non-producing strains, and this regulation role might be meaningful for silent gene cluster activation in Streptomyces. At final, we synthesized racemic-SCB2 and tried to simplify the activation process by adding SCB2 directly to S. albidoflavus J1074, which unfortunately failed to induce paulomycin production.




Similar content being viewed by others
References
Ahmed Y, Rebets Y, Tokovenko B, Brotz E, Luzhetskyy A (2017) Identification of butenolide regulatory system controlling secondary metabolism in Streptomyces albus J1074. Sci Rep 7(1):9784. https://doi.org/10.1038/s41598-017-10316-y
Baltz RH (2010) Streptomyces and Saccharopolyspora hosts for heterologous expression of secondary metabolite gene clusters. J Ind Microbiol Biotechnol 37(8):759–772. https://doi.org/10.1007/s10295-010-0730-9
Bentley SD, Chater KF, A-M COT, Challis GL, Thomson NR, James KD, Harris DE, Quail MA, Kieser H, Harper D (2002) Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417(6885):141–147. https://doi.org/10.1038/417141a
Berdy J (2012) Thoughts and facts about antibiotics: where we are now and where we are heading. J Antibiot (Tokyo) 65(8):385–395. https://doi.org/10.1038/ja.2012.27
Biarnes-Carrera M, Lee CK, Nihira T, Breitling R, Takano E (2018) Orthogonal regulatory circuits for Escherichia coli based on the gamma-butyrolactone system of Streptomyces coelicolor. ACS Synth Biol 7(4):1043–1055. https://doi.org/10.1021/acssynbio.7b00425
Bierman M, Logan R, O’brien K, Seno E, Rao RN, Schoner B (1992) Plasmid cloning vectors for the conjugal transfer of DNA from Escherichia coli to Streptomyces spp. Gene 116(1):43–49. https://doi.org/10.1016/0378-1119(92)90627-2
Cuthbertson L, Nodwell JR (2013) The TetR family of regulators. Microbiol Mol Biol Rev 77(3):440–475. https://doi.org/10.1128/MMBR.00018-13
Daniel-Ivad M, Pimentel-Elardo S, Nodwell JR (2018) Control of specialized metabolism by signaling and transcriptional regulation: opportunities for new platforms for drug discovery? Annu Rev Microbiol 72:25–48. https://doi.org/10.1146/annurev-micro-022618-042458
Genilloud O (2017) Actinomycetes: still a source of novel antibiotics. Nat Prod Rep 34(10):1203–1232. https://doi.org/10.1039/c7np00026j
Gonzalez A, Rodriguez M, Brana AF, Mendez C, Salas JA, Olano C (2016) New insights into paulomycin biosynthesis pathway in Streptomyces albus J1074 and generation of novel derivatives by combinatorial biosynthesis. Microb Cell Factories 15:56. https://doi.org/10.1186/s12934-016-0452-4
Hsiao NH, Soding J, Linke D, Lange C, Hertweck C, Wohlleben W, Takano E (2007) ScbA from Streptomyces coelicolor A3(2) has homology to fatty acid synthases and is able to synthesize gamma-butyrolactones. Microbiology 153(Pt 5):1394–1404. https://doi.org/10.1099/mic.0.2006/004432-0
Kato J-Y, Funa N, Watanabe H, Ohnishi Y, Horinouchi S (2007) Biosynthesis of γ-butyrolactone autoregulators that switch on secondary metabolism and morphological development in Streptomyces. Proc Natl Acad Sci U S A 104(7):2378–2383. https://doi.org/10.1073/pnas.0607472104
Khokhlov A, Tovarova I, Borisova L, Pliner S, Shevchenko L, Kornitskaia E, Ivkina N, Rapoport I (1967) The A-factor, responsible for streptomycin biosynthesis by mutant strains of Actinomyces streptomycini. Dokl Akad Nauk SSSR 177(1):232–235
Kieser T, Bibb MJ, Buttner MJ, Chater KF, Hopwood DA (2000) Practical streptomyces genetics. John Innes Foundation, Norwich
Kitani S, Doi M, Shimizu T, Maeda A, Nihira T (2010) Control of secondary metabolism by farX, which is involved in the γ-butyrolactone biosynthesis of Streptomyces lavendulae FRI-5. Arch Microbiol 192(3):211–220. https://doi.org/10.1007/s00203-010-0550-3
Li J, Guo Z, Huang W, Meng X, Ai G, Tang G, Chen Y (2013) Mining of a streptothricin gene cluster from Streptomyces sp. TP-A0356 genome via heterologous expression. Sci China Life Sci 56(7):619–627. https://doi.org/10.1007/s11427-013-4504-2
Li J, Xie Z, Wang M, Ai G, Chen Y (2015a) Identification and analysis of the paulomycin biosynthetic gene cluster and titer improvement of the paulomycins in Streptomyces paulus NRRL 8115. PLoS One 10(3):e0120542. https://doi.org/10.1371/journal.pone.0120542
Li P, Li J, Guo Z, Tang W, Han J, Meng X, Hao T, Zhu Y, Zhang L, Chen Y (2015b) An efficient blue-white screening based gene inactivation system for Streptomyces. Appl Microbiol Biotechnol 99(4):1923–1933. https://doi.org/10.1007/s00253-014-6369-0
Li P, Guo Z, Tang W, Chen Y (2018) Activation of three natural product biosynthetic gene clusters from Streptomyces lavendulae CGMCC 4.1386 by a reporter-guided strategy. Synth Syst Biotechnol 3(4):254–260. https://doi.org/10.1016/j.synbio.2018.10.010
Nguyen TB, Kitani S, Shimma S, Nihira T (2018) Butenolides from Streptomyces albus J1074 act as external signals to stimulate avermectin production in Streptomyces avermitilis. Appl Environ Microbiol 84(9). https://doi.org/10.1128/AEM.02791-17
Niu G, Chater KF, Tian Y, Zhang J, Tan H (2016) Specialised metabolites regulating antibiotic biosynthesis in Streptomyces spp. FEMS Microbiol Rev 40(4):554–573. https://doi.org/10.1093/femsre/fuw012
Olano C, Garcia I, Gonzalez A, Rodriguez M, Rozas D, Rubio J, Sanchez-Hidalgo M, Brana AF, Mendez C, Salas JA (2014) Activation and identification of five clusters for secondary metabolites in Streptomyces albus J1074. Microb Biotechnol 7(3):242–256. https://doi.org/10.1111/1751-7915.12116
Recio E, Colinas Á, Rumbero Á, Aparicio JF, Martín JF (2004) PI factor, a novel type quorum-sensing inducer elicits pimaricin production in Streptomyces natalensis. J Biol Chem 279(40):41586–41593. https://doi.org/10.1074/jbc.M402340200
Rutledge PJ, Challis GL (2015) Discovery of microbial natural products by activation of silent biosynthetic gene clusters. Nat Rev Microbiol 13(8):509–523. https://doi.org/10.1038/nrmicro3496
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Sarkale AM, Kumar A, Appayee C (2018) Organocatalytic approach for short asymmetric synthesis of (R)-paraconyl alcohol: application to the total syntheses of IM-2, SCB2, and A-Factor γ-Butyrolactone autoregulators. J Organomet Chem 83(7):4167–4172. https://doi.org/10.1021/acs.joc.8b00122
Short JM, Sorge JA, Fernandez JM, Huse WD (1988) λ ZAP: a bacteriophage λ expression vector with in vivo excision properties. Nucleic Acids Res 16(15):7583–7600. https://doi.org/10.1093/nar/16.15.7583
Takano E (2006) Gamma-butyrolactones: Streptomyces signalling molecules regulating antibiotic production and differentiation. Curr Opin Microbiol 9(3):287–294. https://doi.org/10.1016/j.mib.2006.04.003
Takano E, Nihira T, Hara Y, Jones JJ, Gershater CJ, Yamada Y, Bibb M (2000) Purification and structural determination of SCB1, a γ-butyrolactone that elicits antibiotic production in Streptomyces coelicolor A3(2). J Biol Chem 275(15):11010–11016. https://doi.org/10.1074/jbc.275.15.11010
Takano E, Chakraburtty R, Nihira T, Yamada Y, Bibb MJ (2001) A complex role for the γ-butyrolactone SCB1 in regulating antibiotic production in Streptomyces coelicolor A3(2). Mol Microbiol 41(5):1015–1028. https://doi.org/10.1046/j.1365-2958.2001.02562.x
Takano E, Kinoshita H, Mersinias V, Bucca G, Hotchkiss G, Nihira T, Smith CP, Bibb M, Wohlleben W, Chater K (2005) A bacterial hormone (the SCB1) directly controls the expression of a pathway-specific regulatory gene in the cryptic type I polyketide biosynthetic gene cluster of Streptomyces coelicolor. Mol Microbiol 56(2):465–479. https://doi.org/10.1111/j.1365-2958.2005.04543.x
Wei J, He L, Niu G (2018) Regulation of antibiotic biosynthesis in actinomycetes: perspectives and challenges. Synth Syst Biotechnol 3(4):229–235. https://doi.org/10.1016/j.synbio.2018.10.005
Yang YH, Joo HS, Lee K, Liou KK, Lee HC, Sohng JK, Kim BG (2005) Novel method for detection of butanolides in Streptomyces coelicolor culture broth, using a His-tagged receptor (ScbR) and mass spectrometry. Appl Environ Microbiol 71(9):5050–5055. https://doi.org/10.1128/AEM.71.9.5050-5055.2005
Zaburannyi N, Rabyk M, Ostash B, Fedorenko V, Luzhetskyy A (2014) Insights into naturally minimised Streptomyces albus J1074 genome. BMC Genomics 15(1):97. https://doi.org/10.1186/1471-2164-15-97
Zou Z, Du D, Zhang Y, Zhang J, Niu G, Tan H (2014) A γ-butyrolactone-sensing activator/repressor, JadR3, controls a regulatory mini-network for jadomycin biosynthesis. Mol Microbiol 94(3):490–505. https://doi.org/10.1111/mmi.12752
Acknowledgments
We thank Prof. Huarong Tan and A/Prof. Jihui Zhang, Institute of Microbiology, Chinese Academy of Sciences, for generously provided us with SCB standards and bacterial strains.
Funding
This study was funded in part by the Ministry of Science and Technology of China (2018YFA0901901 and 2015CB150600) and the National Natural Science Foundation of China (31670032).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 428 kb)
Rights and permissions
About this article
Cite this article
Zhang, Y., Wang, M., Tian, J. et al. Activation of paulomycin production by exogenous γ-butyrolactone signaling molecules in Streptomyces albidoflavus J1074. Appl Microbiol Biotechnol 104, 1695–1705 (2020). https://doi.org/10.1007/s00253-019-10329-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-10329-9