Abstract
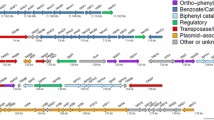
Cupriavidus basilensis WS degrades diphenyl ether (DE) and its lower brominated derivatives using enzymes encoded by the bph operon. However, it is not yet known under what circumstances bph genes are expressed and how they are regulated in C. basilensis WS. To answer these questions, we used transposon mutagenesis and identified a new two-component regulatory system, BphS/BphT, in C. basilensis WS, which is indispensable for the expression of the bph operon. When BphS or BphT is inactivated, C. basilensis WS no longer exhibits the ability to decompose DE. Using a β-galactosidase reporter system and RT-qPCR, we showed that bph genes are constitutively transcribed in C. basilensis WS and that deletion of bphS or bphT strongly inhibited the transcription of bph genes. We also showed that the gene ORF0, which is upstream of bphA1 and is similar to the GntR-family regulators of the bph operon, is not involved in the constitutive transcription of the bph operon in C. basilensis WS. The cis-acting elements required for the expression and regulation of bph genes in the DE degradation pathway are included in the intergenic region between ORF0 and bphA1. Our results suggest that BphS/BphT represents a new two-component regulatory system for the bph operon that is necessary for the constitutive expression of bph genes.







Similar content being viewed by others
References
Anthamatten D, Hennecke H (1991) The regulatory status of the fixL- and fixJ-like genes in Bradyrhizobium japonicum may be different from that in Rhizobium meliloti. Mol Gen Genet 225(1):38–48
Beltrametti F, Reniero D, Backhaus S, Hofer B (2001) Analysis of transcription of the bph locus of Burkholderia sp. strain LB400 and evidence that the ORF0 gene product acts as a regulator of the bphA1 promoter. Microbiology 147:2169–2182
Breland EJ, Eberly AR, Hadjifrangiskou M (2017) An Overview of Two-Component Signal Transduction Systems Implicated in Extra-Intestinal Pathogenic E-coli Infections. Front Cell Infect Microbiol 7:–162
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55:611–622
Capra EJ, Laub MT (2012) Evolution of two-component signal transduction systems. Annu Rev Microbiol 66:325–347
El-Shahawi MS, Hamza A, Bashammakh AS, Al-Saggaf WT (2010) An overview on the accumulation, distribution, transformations, toxicity and analytical methods for the monitoring of persistent organic pollutants. Talanta 80(5):1587–1597
Erickson BD, Mondello FJ (1992) Nucleotide sequencing and transcriptional mapping of the genes encoding biphenyl dioxygenase, a multicomponent polychlorinated-biphenyl-degrading enzyme in Pseudomonas strain LB400. J Bacteriol 174(9):2903–2912
Fortnagel P, Harms H, Wittich R-M, Krohn S, Meyer H, Sinnwell V, Wilkes H, Francke W (1990) Metabolism of dibenzofuran by Pseudomonas sp. strain HH69 and the mixed culture HH27. Appl Environ Microbiol 56(4):1148–1156
Furukawa K, Fujihara H (2008) Microbial degradation of polychlorinated biphenyls: biochemical and molecular features. J Biosci Bioeng 105(5):433–449
Goncalves ER, Hara H, Miyazawa D, Davies JE, Eltis LD, Mohn WW (2006) Transcriptomic assessment of isozymes in the biphenyl pathway of Rhodococcus sp. strain RHA1. Appl Environ Microbiol 72(9):6183–6193
Hamblin MJ, Shaw JG, Kelly DJ (1993) Sequence analysis and interposon mutagenesis of a sensor-kinase (DctS) and response-regulator (DctR) controlling synthesis of the high-affinity C4-dicarboxylate transport system in Rhodobacter capsulatus. Mol Gen Genet 237(1–2):215–224
Herrero M, de Lorenzo V, Timmis KN (1990) Transposon vectors containing non-antibiotic resistance selection markers for cloning and stable chromosomal insertion of foreign genes in gram-negative bacteria. J Bacteriol 172(11):6557–6567
Higuchi R, Krummel B, Saiki RK (1988) A general method of in vitro preparation and specific mutagenesis of DNA fragments: study of protein and DNA interactions. Nucleic Acids Res 16(15):7351–7367
Liu Q, Yeo WS, Bae T (2016) The SaeRS two-component system of Staphylococcus aureus. Genes 7(10)
Liu YG, Chen Y (2007) High-efficiency thermal asymmetric interlaced PCR for amplification of unknown flanking sequences. Biotechniques 43(5):649–650 652, 654 passim
Masai E, Yamada A, Healy JM, Hatta T, Kimbara K, Fukuda M, Yano K (1995) Characterization of biphenyl catabolic genes of gram-positive polychlorinated biphenyl degrader Rhodococcus sp. strain RHA1. Appl Environ Microbiol 61(6):2079–2085
Mattos-Graner RO, Duncan MJ (2017) Two-component signal transduction systems in oral bacteria. J Oral Microbiol 9(1):1400858
Mikkelsen H, Sivaneson M, Filloux A (2011) Key two-component regulatory systems that control biofilm formation in Pseudomonas aeruginosa. Environ Microbiol 13(7):1666–1681
Miller JH (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor
Nguyen PAT, Trinh THT, Fukumitsu Y, Shimodaira J, Miyauchi K, Tokuda M, Kasai D, Masai E, Fukuda M (2013) Gene cluster and regulation system for 1,1-dichloro-2,2-bis(4-chlorophenyl) ethylene (DDE) degradation in Janibacter sp. TYM3221. J Biosci Bioeng 116(1):91–100
Ohtsubo Y, Delawary M, Kimbara K, Takagi M, Ohta A, Nagata Y (2001) BphS, a key transcriptional regulator of bph genes involved in polychlorinated biphenyl/biphenyl degradation in Pseudomonas sp. KKS102. J Biol Chem 276(39):36146–36154
Ohtsubo Y, Goto H, Nagata Y, Kudo T, Tsuda M (2006) Identification of a response regulator gene for catabolite control from a PCB-degrading beta-proteobacteria, Acidovorax sp. KKS102. Mol Microbiol 60(6):1563–1575
Pieper DH, Seeger M (2008) Bacterial metabolism of polychlorinated biphenyls. J Mol Microbiol Biotechnol 15(2–3):121–138
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Schafer A, Tauch A, Jager W, Kalinowski J, Thierbach G, Puhler A (1994) Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: selection of defined deletions in the chromosome of Corynebacterium glutamicum. Gene 145(1):69–73
Solovyev V, Salamov A (2011) Automatic annotation of microbial genomes and metagenomic sequences. Metagenomics and its application in agriculture, biomedicine and environmental studies:61–78
Takeda H, Shimodaira J, Yukawa K, Hara N, Kasai D, Miyauchi K, Masai E, Fukuda M (2010) Dual two-component regulatory systems are involved in aromatic compound degradation in a polychlorinated-biphenyl degrader, Rhodococcus jostii RHA1. J Bacteriol 192(18):4741–4751
Wang S, Bai NL, Wang B, Feng Z, Hutchins WC, Yang CH, Zhao YH (2015) Characterization of the molecular degradation mechanism of diphenyl ethers by Cupriavidus sp. WS. Environ Sci Pollut Res 22(21):16914–16926
Wang S, Zhang MN, Bai NL, Ding HT, Zhu XF, Zhao YH (2017) Construction, properties, and application of the pCB5 plasmid, a novel conjugative shuttle vector with a Cupriavidus basilensis origin of replication. Appl Microbiol Biotechnol 101(3):1217–1226
Watanabe T, Fujihara H, Furukawa K (2003) Characterization of the second LysR-type regulator in the biphenyl-catabolic gene cluster of Pseudomonas pseudoalcaligenes KF707. J Bacteriol 185(12):3575–3582
Watanabe T, Inoue R, Kimura N, Furukawa K (2000) Versatile transcription of biphenyl catabolic bph operon in Pseudomonas pseudoalcaligenes KF707. J Biol Chem 275(40):31016–31023
Wong SM, Mekalanos JJ (2000) Genetic footprinting with mariner-based transposition in Pseudomonas aeruginosa. Proc Natl Acad Sci U S A 97(18):10191–10196
Funding
This study was supported by the National Key Basic Research Program of China (2015CB150502), the National Natural Science Foundation of China (31470191, 41671314, 41877114), the Zhejiang Provincial Natural Science Foundation of China (LQ17D010001), and the Key Research and Development Program of Zhejiang Province (2015C03011).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no conflict of interest.
Human and animal rights and informed consent
This article does not contain any studies with human participants performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 959 kb)
Rights and permissions
About this article
Cite this article
Wang, S., Li, Y., Wang, B. et al. New two-component regulatory system required for the constitutive expression of bph operon in Cupriavidus basilensis WS. Appl Microbiol Biotechnol 103, 3099–3109 (2019). https://doi.org/10.1007/s00253-019-09686-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-09686-2