Abstract
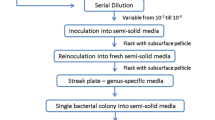
We developed a sensitive quantitative assay for detecting Ralstonia solanacearum in soil by most probable number (MPN) analysis based on bio-PCR results. For development of the detection method, we optimized an elution buffer containing 5 g/L skim milk for extracting bacteria from soil and reducing contamination of polymerase inhibitors in soil extracts. Because R. solanacearum can grow in water without any added nutrients, we used a cultivation buffer in the culture step of the bio-PCR that contained only the buffer and antibiotics to suppress the growth of other soil microorganisms. To quantify the bacterial population in soil, the elution buffer was added to 10 g soil on a dry weight basis so that the combined weight of buffer, soil, and soil-water was 50 g; 5 mL of soil extract was assumed to originate from 1 g of soil. The soil extract was divided into triplicate aliquots each of 5 mL and 500, 50, and 5 μL. Each aliquot was diluted with the cultivation buffer and incubated at 35 °C for about 24 h. After incubation, 5 μL of culture was directly used for nested PCR. The number of aliquots showing positive results was collectively checked against the MPN table. The method could quantify bacterial populations in soil down to 3 cfu/10 g dried soil and was successfully applied to several types of soil. We applied the method for the quantitative detection of R. solanacearum in horticultural soils, which could quantitatively detect small populations (9.3 cfu/g), but the semiselective media were not able to detect the bacteria.




Similar content being viewed by others
References
Boudazin G, Le Roux AC, Josi K, Labarre P, Jouan B (1999) Design of division specific primers of Ralstonia solanacearum and application to the identification of European isolates. Eur J Plant Pathol 105:373–380
Caruso P, Bertolini E, Cambra M, Lopez MM (2003) A new and sensitive co-operational polymerase chain reaction for rapid detection of Ralstonia solanacearum in water. J Microbiol Meth 55:257–272
Caruso P, Palomo JL, Bertolini E, Alvarez B, Lopez MM, Biosca EG (2005) Seasonal variation of Ralstonia solanacearum biovar 2 populations in a Spanish river, recovery of stressed cells at low temperatures. Appl Environ Microbiol 71:140–148
Chen W-Y, Echandi E (1982) Bacteriocin production and semiselective medium for detection, isolation and quantification of Pseudomonas solanacearum in soil. Phytopathology 72:310–313
Chen Y, Zhang W-Z, Liu X, Ma Z-H, Li B, Allen C, Guo J-H (2010) A real-time PCR assay for the quantitative detection of Ralstonia solanacearum in horticultural soil and plant tissues. J Microbiol Biotechnol 20:193–201
Denny TP (2006) Plant pathogenic Ralstonia species. In: Gnanamanickam SS (ed) Plant-associated bacteria. Springer, Dordrecht, The Netherlands, pp 573–644
Dittapongpitch V, Surat S (2003) Detection of Ralstonia solanacearum in soil and weeds from commercial tomato fields using immunocapture and the polymerase chain reaction. J Phytopathol 151:239–246
Elphinstone JG, Henessy J, Wilson JK, Stead D (1996) Sensitivity of different methods for the detection of Ralstonia solanacearum in potato tuber extracts. Bull OEPP/EPPO Bull 26:663–678
Fegan M, Prior P (2005) How complex is the Ralstonia solanacearum species complex. In: Allen C, Prior P, Hayward AC (eds) Bacterial wilt disease and the Ralstonia solanacearum species complex. APS Press, St. Paul, MN, pp 449–461
Fredslund L, Ekelund F, Jacobsen CS, Kaare Johnsen K (2001) Development and application of a most-probable-number-PCR assay to quantify flagellate populations in soil samples. Appl Environ Microbiol 67:1613–1618
Goto M (1992) Lethal dilution effect. Pages 61–63 in. Fundamentals of bacterial plant pathology. Academic Press, New York
Granada GA, Sequeira L (1983) A new selective medium for Pseudomonas solanacearum. Plant Dis 67:1084–1088
Grover A, Azmi W, Khurana SMP, Chakrabarti SK (2009) Multiple displacement amplification as a pre-polymerase chain reaction (pre-PCR) to detect ultra low population of Ralstonia solanacearum (Smith 1896) Yabuchi et al. (1996). Lett Appl Microbiol 49:539–543
Ha Y, Kim J-S, Denny TP, Schell MA (2012) A rapid, sensitive assay for Ralstonia solanacearum race 3 biovar 2 in plant and soil samples using magnetic beads and real-time PCR. Plant Dis 96:258–264
Hara H, Ono K (1983) Ecological studies on the bacterial wilt of tobacco, caused by Pseudomonas solanacearum E. F. Smith. I. A selective medium for isolation and detection of P. solanacearum. Bull Okayama Tob Exp Stn 42:127–138
Hayward AC (1991) Biology and epidemiology of bacterial wilt caused by Pseudomonas solanacearum. Annu Rev Phytopathol 29:65–87
Hoshino YT, Matsumoto N (2004) An improved DNA extraction method using skim milk from soils that strongly adsorb DNA. Microbes Environ 19:13–19
Huang J, Wu J, Li C, Xiao C, Wang G (2009) Specific and sensitive detection of Ralstonia solanacearum in soil with quantitative, real-time PCR assays. J Appl Microbiol 107:1729–1739
Ito S, Ushijima Y, Fujii T, Tanaka S, Kameya-Iwaki M, Yoshiwara S, Kishi F (1998) Detection of viable cells of Ralstonia solanacearum in soil using a semiselective medium and a PCR technique. J Phytopathol 146:379–384
Janse JD (1988) A detection method for Pseudomonas solanacearum in symptomless potato tubers and some data on its sensitivity and specificity. Bull OEPP/EPPO Bull 18:343–351
Ji P, Allen C, Sanchez-Perez A, Yao J, Elphinstone JG, Jones JB, Momol MT (2007) New diversity of Ralstonia solanacearum strains associated with vegetable and ornamental crops in Florida. Plant Dis 91:195–203
Kang MJ, Lee MH, Shim JK, Seo ST, Shrestha R, Cho MS, Hahn JH, Park DS (2007) PCR-based specific detection of Ralstonia solanacearum by amplification of cytochrome c1 signal peptide sequences. J Microbiol Biotechnol 17:1765–1771
Karganilla AD, Buddenhagen IW (1972) Development of a selective medium for Pseudomonas solanacearum. Phytopathology 62:1373–1376
Kelman A (1954) The relationship of pathogenicity in Pseudomonas solanacearum to colony appearance on a tetrazolium medium. Phytopathology 44:693–695
Kikuchi T, Iwasaki K, Nishihara H, Takamura Y, Yagi O (2001) Quantitative and specific detection of a trichloroethylene-degrading methanotroph, Methylocystis sp. strain M, by a most probable number-polymerase chain reaction method. Biosci Biotechnol Biochem 65:2673–2681
Kutin RK, Alvarez A, Jenkins DM (2009) Detection of Ralstonia solanacearum in natural substrates using phage amplification integrated with real-time PCR assay. J Microbiol Meth 76:241–246
Lee YA, Wang CC (2000) The design of specific primers for the detection of Ralstonia solanacearum in soil samples by polymerase chain reaction. Bot Bull of Acad Sin 41:121–128
Luan X, Chen J, Liu Y, Li Y, Jia J, Liu R, Zhang X-H (2008) Rapid quantitative detection of Vibrio parahaemolyticus in seafood by MPN-PCR. Curr Microbiol 57:218–221
Miwa N, Nishio T, Arita Y, Kawamori F, Masuda T, Akiyama M (2003) Evaluation of MPN method combined with PCR procedure for detection and enumeration of Vibrio parahaemolyticus in seafood. J Food Hyg Soc Japan 44:289–293
Miwa N, Kashiwagi M, Kawamori F, Masuda T, Sano Y, Hiroi M, Kurashige H (2006) Levels of Vibrio parahaemolyticus and thermostable direct hemolysin gene-positive organisms in retail seafood determined by the most probable number-polymerase chain reaction (MPN-PCR) method. J Food Hyg Soc Japan 47:41–45
Nesmith WC, Jenkins SF Jr (1979) A selective medium for the isolation and quantification of Pseudomonas solanacearum from soil. Phytopathology 69:182–185
Okabe N (1969) Population changes of Pseudomonas solanacearum and soil microorganisms in artificially infested natural field soil. Bull Fac Agr Shizuoka Univ 19:1–29
Opina N, Tavner F, Hollway G, Wang J-F, Li T-H, Maghirang R, Fegan M, Hayward AC, Krishnapillai V, Hong WF, Holloway BW, Timmis JN (1997) A novel method for development of species and strain specific DNA probes and PCR primers for identifying Burkholderia solanacearum (formerly Pseudomonas solanacearum). Asia Pac J Mol Biol Biotechnol 5:19–30
Ozakman M, Schaad NW (2003) A real-time BIO-PCR assay for detection of Ralstonia solanacearum race 3, biovar 2, in asymptomatic potato tubers. Can J Plant Pathol 25:232–239
Pastrik KH, Maiss E (2000) Detection of Ralstonia solanacearum in potato tubers by polymerase chain reaction. J Phytopathol 148:619–626
Pastrik KH, Elphinstone JG, Pukall R (2002) Sequence analysis and detection of Ralstonia solanacearum by multiplex PCR amplification of 16S-23S ribosomal intergenic spacer region with internal positive control. Eur J Plant Pathol 108:831–842
Poussier S, Cheron JJ, Couteau A, Luisetti J (2003) Evaluation of procedures for reliable PCR detection of Ralstonia solanacearum in common natural substrates. J Microbiol Meth 51:349–359
Pradhanang PM, Elphinstone JG, Fox RTV (2000) Sensitive detection of Ralstonia solanacearum in soil: a comparison of different detection techniques. Plant Pathol 49:414–422
Robinson-Smith A, Jones P, Elphinstone JG, Forde SMD (1995) Production of antibodies to Pseudomonas solanacearum, the causative agent of bacterial wilt. Food Agric Immunol 7:67–79
Schönfeld J, Heuer H, van Elsas JD, Smalla K (2003) Specific and sensitive detection of Ralstonia solanacearum in soil on the basis of PCR amplification of fliC fragments. Appl Environ Microbiol 69:7248–7256
Seal SE, Jackson LA, Young JPW, Daniels MJ (1993) Differentiation of Pseudomonas solanacearum, Pseudomonas syzygii, Pseudomonas pickettii and the blood disease bacterium by partial 16S rRNA sequencing: construction of oligonucleotide primers for sensitive detection by polymerase chain reaction. J Gen Microbiol 139:1587–1594
Thaithongnum S, Ratanama P, Weeradechapol K, Sukhoom A, Vuddhakul V (2006) Detection of V. harveyi in shrimp postlarvae and hatchery tank water by the most probable number technique with PCR. Aquaculture 261:1–9
Thammakijjawat P, Thaveechai N, Kositratana W, Chunwongse J, Frederick RD, Schaad NW (2006) Detection of Ralstonia solanacearum in ginger rhizomes by real-time PCR. Can J Plant Pathol 28:391–400
Volossiouk T, Robb E, Nazar R (1995) Direct DNA extraction for PCR-mediated assays of soil organisms. Appl Environ Microbiol 61:3972–3976
Wakimoto S (1960) Classification of strains of Xanthomonas oryzae on the basis of their susceptibility against bacteriophages. Ann Phytopathol Soc Jpn 25:193–198
Weller SA, Elphinstone JG, Smith NC, Boonham N, Stead DE (2000) Detection of Ralstonia solanacearum strains with a quantitative, multiplex, real-time, fluorogenic PCR (TaqMan) assay. Appl Environ Microbiol 66:2853–2858
Acknowledgments
We thank Ms. A Notsu (Ornamental Plants and Vegetables Research Center, Hokkaido Research Organization), Dr. M Maeda (Niigata Agricultural Research Institute), and Dr. H Kajihara (Yamaguchi Prefectural Technology Center for Agricultural and Forestry) for kindly providing the horticultural soil samples. This work was supported by a Grant-in-Aid for Science and Technology Research Promotion Program for Agriculture, Forestry, Fisheries and Food Industry from the Ministry of Agriculture, Forestry and Fisheries, Japan (25062C).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 159 kb)
Rights and permissions
About this article
Cite this article
Inoue, Y., Nakaho, K. Sensitive quantitative detection of Ralstonia solanacearum in soil by the most probable number-polymerase chain reaction (MPN-PCR) method. Appl Microbiol Biotechnol 98, 4169–4177 (2014). https://doi.org/10.1007/s00253-014-5604-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-5604-z