Abstract
Rare Actinomycetes from underexplored marine environments are targeted in drug discovery studies due to the Actinomycetes’ potentially huge resource of structurally diverse natural products with unusual biological activity. Of all marine bacteria, 10 % are Actinomycetes, which have proven an outstanding and fascinating resource for new and potent bioactive molecules. Past and present efforts in the isolation of rare Actinomycetes from underexplored diverse natural habitats have resulted in the isolation of about 220 rare Actinomycete genera of which more than 50 taxa have been reported to be the producers of 2,500 bioactive compounds. That amount represents greater than 25 % of the total Actinomycetes metabolites, demonstrating that selective isolation methods are being developed and extensively applied. Due to the high rediscovery rate of known compounds from Actinomycetes, a renewed interest in the development of new antimicrobial agents from rare and novel Actinomycetes is urgently required to combat the increasing number of multidrug-resistant human pathogens. To facilitate that discovery, this review updates all selective isolation media including pretreatment and enrichment methods for the isolation of marine rare Actinomycetes. In addition, this review demonstrates that discovering new compounds with novel scaffolds can be increased by intensive efforts in isolating and screening rare marine genera of Actinomycetes. Between 2007 and mid-2013, 80 new rare Actinomycete species were reported from marine habitats. They belong to 23 rare families, of which three are novel, and 20 novel genera. Of them, the family Micromonosporaceae is dominant as a producer of promising chemical diversity.
Similar content being viewed by others
Introduction
Natural products have continued to play a highly significant role in the drug discovery and development process; about 28 % of the new chemical entities and 42 % of the anticancer drugs introduced into the worldwide market between 1981 and 2006 were natural products and their derivatives (Newman and Cragg 2007).
Microbial natural products represent an important route to the discovery of novel chemicals for the development of new therapeutic agents—more than 22,000 biologically active compounds have been obtained from microbes. Among them, 45 % were produced by Actinobacteria, especially the excellent producers in the genus Streptomyces (Berdy 2005). Actinobacteria have made a significant contribution to the health and well-being of people throughout the world (Demain and Sanchez 2009). Even so, the emergence of antibiotic resistance developed in various bacterial pathogens and the increase in numbers of new diseases and pathogens (such as acquired immunodeficiency syndrome, severe acute respiratory syndrome and H1N1 flu virus) has caused a resurgence of interest in finding new biologically active compounds for drug discovery. However, the ‘law of diminishing returns’ (Fischbach and Walsh 2009) has resulted in fewer new discoveries from the traditional sources (such as plants and soil Actinomycetes) of natural products. Thus, it is critical that new groups of microbes from unexplored habitats be pursued as sources of novel antibiotics and other therapeutic agents (Bull et al. 2005).
The oceans are home to high microbial diversity (Stach and Bull 2005; Sogin et al. 2006). These are also being screened intensively throughout the world for their biodiversity potential (Jensen et al. 2005a, b). Moreover, until now, representatives of a relatively few taxa have been isolated from marine as opposed to terrestrial habitats (Goodfellow 2010). Thus, considering the vastness of the marine environment, the potential rewards of this treasure house represented by the oceans are large (Tiwari and Gupta 2012b).
Novel (new genera in Actinobacteria), new (new species of previously reported rare genera) or rare microbes need to be examined in the search for bioactive compounds with diverse biological activity. Rare Actinomycetes are usually considered as non-streptomycete Actinomycete strains. The isolation frequency of rare Actinomycetes is much lower than that of the streptomycete strains isolated by conventional methods (Baltz 2006)—only 11 genera had been isolated by 1970, increasing to 100 genera by 2005 and 220 genera by 2010 (Berdy 2005; Tiwari and Gupta 2012a). This number is quickly increasing due to recently developed taxonomically selective isolation and genetic techniques. Table 1 shows the approximate number of antibiotics produced by Streptomyces and rare Actinomycetes between 1974 and 2005. By 1974, 125 antibiotics had been isolated from rare Actinomycetes, increasing to 2,250 by 2005, and a recent update by Kurtböke (2012) indicates that there were about 2,500 by 2010. Thus, it is clear that isolation of antibiotics and biologically active metabolites has steadily been increasing from rare Actinomycetes (Fenical and Jensen 2006; Lam 2006; Subramani and Aalbersberg 2012). Furthermore, contemporary bioprospecting of soil Actinobacteria (particularly streptomycetes), the most significant source of new antibiotics in the twentieth century has largely resulted in the rediscovery of already-known compounds (Walsh 2003; Fischbach and Walsh 2009); rare Actinomycetes should be targeted for novel drug discovery programmes. Many excellent reviews describing Actinomycetes diversity, secondary metabolism, natural product discovery and genetics have appeared over the last 20 years. However, fewer reviews have described rare Actinomycetes diversity and their increasing contribution to the production of novel compounds (Lazzarini et al. 2000; Kurtböke 2012; Tiwari and Gupta 2012a, b).
The goal of this review is to summarize isolation and cultivation methods, and discuss the new and rare Actinobacteria findings in studies since 2007 particularly from marine habitats, also to discuss their enormous biotechnological potential in the area of natural products discovery and related applications.
High rate of rediscovery of known compounds
It is important to speculate on the reasons for the high rate of rediscovery of antimicrobial compounds in previous screening programmes. According to Stach (2010), the reasons are likely to include bias in the screening programmes and limitations in analytical technology, but more importantly in the organisms being screened themselves. Many new antibiotics were isolated from Actinomycetes (particularly from a single genus Streptomyces) between the late 1940s and 1960s—a period which came to be known as the Golden Age of antibiotic discovery—but the rate of new discoveries plummeted thereafter due in large part to the frequent rediscovery of highly abundant existing compounds. Stach (2010) suggested that the distribution of microbial species is probably similar to that of other organisms, i.e. there are small numbers of very rich species and those species may also be those that are readily cultured (as is the case for Streptomyces); thus, they represent a small fraction of the available diversity. In addition, many streptomycetes, although isolated from different environments, evidently produce the same known compounds, probably due to the frequent genetic exchange between them (Bredholdt et al. 2007).
However, recent genome sequence information suggests that this Streptomyces source of novel compounds is still not yet exhausted. Whole-genome sequencing of several streptomycetes (Bentley et al. 2002; Ikeda et al. 2003; Ohnishi et al. 2008; Song et al. 2010; Medema et al. 2010) revealed that each member can produce on average 20–30 bioactive small molecules, but only a small fraction of these molecules have ever been detected under various culture conditions. Consequently, over the past decade, researchers have been attempting several methods such as cloning (Peiru et al. 2005) and heterologous expression (Mutka et al. 2006) of biosynthetic gene clusters, interfering with regulatory pathways (Laureti et al. 2011), varying culture conditions (Sánchez et al. 2010), co-culturing two or more organisms together (Kurosawa et al. 2008), the adaptive evolution (Charusanti et al. 2012) and other strategies (Baltz 2011) to stimulate the production of new compounds. Furthermore, the biosynthetic gene pathways used to make the antimicrobial compounds are distributed among the Actinomycetes at varying frequencies, such that a single compound may be found in one in ten strains screened. In other words, thousands of compounds should be found in 1 in 107 are screened (Baltz 2007; Stach 2010). Previous screening activities appeared to be focused on limited species diversity, and those few species produced a number of common compounds (rediscovered antimicrobials) that would obscure the detection of novel antimicrobials in the lower frequency ranges (Stach 2010). Baltz (2007) defined the challenge as finding the resources necessary to discover new antibiotics at frequencies of <1 in 107 within a background noise of 2,000 known antibiotics (Baltz 2007; Stach 2010). Understanding the reasons for rediscovery, coupled with disappointing returns from small molecule libraries, has led to a revival of interest in microbes as sources of new antimicrobial compounds. Proponents of this renaissance have suggested focusing on rare Actinomycetes, the assumption being that species novelty will lead to chemical novelty. In this instance, rare Actinomycetes are not necessarily those that are scarce in nature, but those that are rarely brought into culture (Stach 2010). Thus, it is reasonable to predict that focusing on environments that have been underexplored, and use of selective isolation methods, will lead to the isolation of novel genera and species of Actinomycetes and hence new antimicrobial compounds.
Rare Actinomycetes: selective isolation methods
In a report released by the American Academy of Microbiology entitled “The Microbial World: Foundation of the Biosphere”, Young (1997), estimated that less than 1 % of bacterial species are known, and recent evidence indicates that millions of microbial species are undiscovered (Cragg and Newman 2005). Surprisingly, the approach to the search for potentially valuable bacteria has been largely empirical and restricted to sampling a tiny fraction of the microbial community found in natural habitats. Therefore, techniques that enhance the growth of desirable microorganisms in natural samples (enrichment) or eliminate the undesirable streptomycete propagules and other contaminants from the primary isolation plate (pretreatment) must be developed and employed for selectively isolating particularly rare genera of Actinomycetes (Tiwari and Gupta 2012a).
Different pretreatment methods and media combinations are effective in the isolation of rare Actinomycetes (Tiwari and Gupta 2012a), and many researchers have been attempting to develop methods for isolating desirable rare Actinomycete genera from natural habitats (Nonomura 1988; Nonomura and Hayakawa 1988; Hayakawa et al. 1991a, b, c, d; Hayakawa 1990, 1994, 2003; Hayakawa and Nonomura 1993; Seong et al. 2001; Hamaki et al. 2005; Tan et al. 2006; Qiu et al. 2008; Qin et al. 2009; Nakaew et al. 2009; Baskaran et al. 2011; Istianto et al. 2012; Wang et al. 2013a). Their methods include a variety of pretreatments in combination with different enrichment techniques that selectively supplement isolation media with chemicals and selective antimicrobial agents to successfully increase the selectivity of the isolation media for desirable rare Actinomycetes.
Humic acid vitamin agar (HVA), first developed by Hayakawa and Nonomura (1987a), is one of the milestones in rare Actinomycetes isolation: this medium contains soil humic acid as the sole carbon and nitrogen sources which are suitable for recovery of rare Actinomycetes from natural samples. Although humic acid is an extremely heterogeneous cross-linked polymer resistant to biological decomposition and restricts the growth of non-filamentous bacteria colonies (Seong et al. 2001), Actinomycetes can utilize it as a nutrient source and also use it to support sporulation. A number of rare genera isolated by researchers described in this review have been discovered through the use of HVA together with different pretreatment and enrichment techniques, to successfully isolate rare Actinomycetes. Rare Actinomycetes as well as Streptomyces grow well on HVA. Although the growth rate of Actinomycetes is low, discrimination of typical morphology of colonies is easy on HVA because the black colour of HVA also makes it suitable for determining the morphology of white Actinomycetes colonies. The activation of spore germination by humic acid is believed to be one of the causes that increases the number of diverse Actinomycetes colonies on HVA (Hayakawa and Nonomura 1987b).
Moist and dry heat treatment
Samples secured from natural habitats cultured without pretreatment surrendered (in order of frequency) bacteria other than Actinomycetes, Streptomyces, fungi and non-streptomycete Actinomycetes (Seong et al. 2001). Consequently, different pretreatment procedures and selective isolation media have been recommended for the selective isolation of novel and rare Actinomycetes. The aerial spores of most Actinomycete genera resist desiccation and show a slightly higher resistance to wet or dry heat than do the corresponding vegetative hyphae (Seong et al. 2001). Pretreatments of natural habitat samples by drying and heating stimulated the isolation of rare Actinomycetes (Nolan and Cross 1988; Kim et al. 1995). In comparison to the other genera of rare Actinomycetes, the rare genera Streptosporangium are difficult to isolate by traditional methods as their sporangiospores are able to withstand and resist physical or chemical pretreatments: Hayakawa et al. (1991a) found that dry heat treatment (120 °C for 1 h) of natural samples greatly induces the growth of Streptosporangium spp. After surface sterilization, Qin et al. (2009) subjected different medicinal plant samples to continuous drying at 100 °C for 15 min: directly plating on different selective media enabled the isolation of 280 strains belonging to the genera Pseudonocardia, Nocardiopsis, Micromonospora and Streptosporangium. Additionally, along with dry heating of samples treated with chemicals such as 0.01 % benzethonium chloride, 0.03 % chlorhexidine gluconate, 0.05 % sodium dodecylsulfate (SDS), 6 % yeast extract and 1.5 % phenol and supplemented with different selective antibiotics such as leucomycin, nalidixic acid on HVA drastically eliminated the unicellular bacteria and other unwanted Actinomycete propagules (including Streptomyces spp.) from the isolation plates and increased the selectivity for Streptosporangium spp., Microbispora spp., Acitinomadura spp., Micromonospora spp., Nocardia spp. and Nonomurea spp. (Hayakawa et al. 1988, 1991a, b, c, d; Hayakawa 2008; Khamna et al. 2009). Recently, Niyomvong et al. (2012) showed that pretreatment of samples with moist (50 °C for 6 min) and dry (120 °C for 1 h) heating and 1.5 % phenol reduced the number of undesirable bacteria and enhanced the selective isolation of Actinoplanes, Gordonia, Microbispora, Micromonospora, Nocardia and Nonomuraea. The successful isolation of members of the genera Actinomadura and Saccharopolyspora from caves was reported for the first time using these pretreatments with selective isolation media (Niyomvong et al. 2012).
Phenol treatment
An alternative approach is to make the isolation procedure more selective by adding chemicals such as phenol to the natural samples (Nonomura 1988; Hayakawa et al. 1991c). Phenol is a biocide and toxic to bacteria, fungi and streptomycetes, so treatment with 1.5 % phenol reduces the number of those organisms by removing sensitive species (Hayakawa et al. 1991b, 2004). Khamna et al. (2009) selectively isolated 11 % of non-streptomycetes including the rare genera Actinomadura, Microbispora, Micromonospora, Nocardia and Nonomurea by pretreating the samples with 1.5 % phenol and then plating on HVA. Although phenol treatment of soil suspension lowered the number of fungi and other bacteria, the Actinomycetes were less affected: 65 % of the colonies were rare Actinomycetes. The phenol pretreatment of the soil killed bacteria and streptomycetes in the samples, while keeping Micromonosporae and Microbisporae alive (Hayakawa et al. 1991b; Qiu et al. 2008). In another study, the rare genera Micromonospora (49.2 %), Actinomadura (13.1 %), Microbispora (9.8 %) and Polymorphospora (3.3 %) were successfully obtained from soil samples using 1.5 % phenol pretreatment (Istianto et al. 2012).
Selective antimicrobial agents
Several rare Actinomycetes are resistant to a wide spectrum of antibiotics. Thus, several antibiotic molecules have been used in selective media to inhibit the competing bacteria including fast-growing Actinomycetes (Okami and Hotta 1988). Selective isolation plates containing novobiocin significantly increased the numbers of Micromonospora-like colonies (Qiu et al. 2008). Gentamicin is also one of the selective agents used to access Micromonospora spp. (Williams and Wellington 1982). Specialized growth media have also been developed to isolate specific Actinomycete genera (Seong et al. 2001). Hayakawa and Nonomura (1987a, b) and Cho et al. (1994) chose macromolecules such as casein, chitin, hair hydrolysate and humic acid as carbon and nitrogen sources of rare Actinomycetes.
Calcium carbonate treatment
Treatment of natural habitat samples with calcium carbonate increased the populations of rare genera of Actinomycetes (Alferova and Terekhova 1988). The mechanism of the calcium carbonate effect is not clear; however, Tsao et al. (1960) described natural samples mixed with powdered calcium carbonate where the pH is altered in favour of the growth of Actinomycete propagules and the calcium ions have the ability to stimulate the formation of aerial mycelia by several Actinomycete cultures (Natsume et al. 1989). Furthermore, Tsao et al. (1960) demonstrated significant increases in the relative plate counts of the Actinomycete populations in soil samples treated with calcium carbonate. In addition, using a combined calcium carbonate rehydration and centrifugation (RC) procedure, Otoguro et al. (2001) successfully isolated diverse Actinokineospora spp. and other Actinomycetes from soils and plant litter. Recently, Qin et al. (2009) demonstrated that the enrichment stage with calcium carbonate and the RC procedure was also suitable for the isolation of zoosporic and other rare Actinobacteria; they were the first to the isolation of Saccharopolyspora, Dietzia, Blastococcus, Dactylosporangium, Promicromonospora, Oerskovia, Actinocorallia and Jiangella species from endophytic environments. Therefore, the calcium carbonate procedure, in combination with other selective isolation methods, is recommended for the isolation of rare genera of Actinomycetes from soil samples (Tiwari and Gupta 2012a).
Microwave irradiation
Many studies have examined the use of microwave energy for sterilization of soil (Wang et al. 2013a), yet there are few reports about the effect of microwave irradiation on the culturability of microorganisms, and especially the culturability of Actinomycetes (Bulina et al. 1997; Yang et al. 2008; Xue et al. 2010). Ferriss (1984) reported that microwave irradiation of soil reduced total fungal and total prokaryote counts in soil extracts. Bulina et al. (1997) reported that microwave irradiation significantly increased the number of culturable rare Actinomycetes taxa in soil, including Micromonospora, Micropolyspora, Norcardia and Actinomadura. Yang et al. (2008) reported that short periods of microwave irradiation increased culturable Actinomycete counts and the number of culturable Actinomycete isolates in a sandy aeolian soil; they also found that irradiation increased the number of antagonistic Actinomycete isolates as a percentage of the total number of cultural Actinomycete isolates. Recently, Xue et al. (2010) reported that microwave irradiation of a calcareous soil increased the total counts of culturable Actinomycetes such as Streptomyces spp. and Micromonospora spp. Furthermore, Wang et al. (2013a) isolated biologically active Streptomyces spp., Nocardia spp., Streptosporangium spp. and Lentzea spp. using microwave irradiation of soil samples. In addition, some researchers used other physical agents such as electromagnetic radiation (Miguélez et al. 1993; Niyomvong et al. 2012), electric pulses and super high frequency radiation (Bulina et al. 1997), ultrasonic waves (Jiang et al. 2010) and extremely high-frequency radiation (Li et al. 2003) for the selective isolation of Actinomycetes in natural samples. All of these methods have significantly increased the total number of rare Actinomycetes isolated.
Centrifugation process
Another physical method, centrifugation, eliminates streptomycetes and other non-motile Actinomycetes from the liquid phase, thereby facilitating the selective growth of rare—especially motile Actinomycetes—on isolation plates subsequent to inoculation (Hayakawa et al. 2000; Qin et al. 2009). The combined enzymatic hydrolysis and differential centrifugation method was particularly useful for isolating endophytic rare Actinobacteria Pseudonocardia, Nocardiopsis and Micromonospora species and species of other genera, including Amycolatopsis, Nocardia, Nonomuraea, Actinomadura, Gordonia, Promicromonospora and Mycobacterium (Qin et al. 2009).
Chemoattractants and chlorination methods
Selective isolation of sporulating Actinomycetes known to produce motile spores is done by the use of xylose, chloride, γ-collidine, bromide and vanillin (Hayakawa 2008) which act as chemoattractants for accumulating spores of Actinoplanes, Dactylosporangium and Catenuloplanes (Hayakawa 2008). Further, selective isolation of rare genera Herbidospora, Microbispora, Microtetraspora and Streptosporangium can be achieved by chloramine treatment (Hong et al. 2009), as chlorination is known to suppress growth of contaminant bacteria and promote the growth of rare Actinomycetes when plated on humic acid–vitamin-enriched media (Hong et al. 2009).
Finally, Tiwari and Gupta (2012a) found that selective isolation of rare Actinomycetes from natural habitats using combined physical and chemical treatments of natural samples can increase the chance of isolation of rare genera of Actinomycetes.
Other methods
Several terms have been used in the literature, including ‘uncultured’, ‘unculturable’ and ‘uncultivable’ to describe bacteria that are not readily cultured in the laboratory. Sampling of diverse environments, such as soil, marine sediment or hot springs shows that only 0.01–1 % of cells visible under the microscope will form colonies on a Petri dish, leaving the remaining majority ‘uncultured’ (D’Onofrio et al. 2010). In recent years, researchers have been attempting various methods such as co-culture (D’Onofrio et al. 2010; Stewart 2012), simulation of the natural environment in vitro (Stewart 2012), colony hybridization, flow cytometry and cell sorting, micromanipulation of single bacterial cells (Vartoukian et al. 2010), design and application of the diffusion chamber, ichip and the microbial trap (Gavrish et al. 2008; Lewis et al. 2010) for isolating unculturable microorganisms. Of all of these methods, co-culture has proven successful and so is widely used method for the cultivation of unculturable, novel or rare microorganisms.
Co-culture method
Recently, D’Onofrio et al. (2010) experimentally described the success of the co-culture methods on the culture of unculturable bacteria. Briefly, pairs of colonies growing within a 2-cm distance of each other were selected from high-density isolation plates (50–200 colonies per plate) and re-streaked in close proximity to each other. Each of the two isolates was streaked on one half of an R2Asea plate and cross-streaked through the centre of the plate; the result was regions of proximal, distal and overlapping inoculation (D’Onofrio et al. 2010). Using this method, they were able to isolate an uncultured marine bacterium Maribacter polysiphoniae in the presence of helper strain Micrococcus luteus that was isolated from the same environment. Similarly, an uncultured bacterium Bacillus marisflavi was obtained from fresh water sediment in the presence of the helper strain Bacillus megaterium from the same environment (Stewart 2012). Some unculturable colony-forming microorganisms can grow on a Petri dish only in the presence of other species from the same environment (Kaeberlein et al. 2002; Nichols et al. 2008; D’Onofrio et al. 2010). Interspecies symbiosis based on nutrient exchange (syntrophy) is well known in the bacterial world (McInerney et al. 2008). Bacteria are also known to communicate using an interspecies quorum-sensing factor [autoinducer 2 (AI-2)] that induces synthesis of proteins such as toxins or polymer hydrolases that are useful for a community rather than a single cell (Williams et al. 2007a). Uncultured bacteria, however, do not grow on rich synthetic media (such media should largely obviate the need for nutrient supply by other species), and AI-2 has not been found to act as a growth-promoting factor, raising questions about the nature of unknown growth-promoting factors in microbial communities.
Diverse habitats and genera of rare Actinomycetes
Soil and plants
Soil is well-studied for Actinomycetes populations and most of the rare Actinomycetes reported so far have come from different types of soil (Tiwari and Gupta 2012a, b). The isolation of several new and rare genera discussed in this review under the section ‘Selective isolation methods’ was mostly derived from different soil types. Many rare Actinomycetes are now being isolated from plants (Matsumoto et al. 1998; Taechowisan et al. 2003; Janso and Carter 2010), often for the purpose of finding novel microbial resources for use in screening for new bioactive compounds (Inahashi et al. 2011). For example, Qin et al. (2009) reported for the first time the isolation of Saccharopolyspora, Dietzia, Blastococcus, Dactylosporangium, Promicromonospora, Oerskovia, Actinocorallia and Jiangella species from endophytic environments. A typical endophytic Actinomycete, Frankia, has nitrogen-fixing activity, a function which plays an important role in ecological systems (Xu et al. 2007).
Extreme environments
Extreme environments have unusual growth conditions such as high and low temperature, salt, alkaline and acidic pH, radioactivity and high pressure. Microorganisms from extreme environments have received great attention owing to their special mechanisms of adapting to the conditions in their extreme environments and also because they can produce unusual compounds (Meklat et al. 2011). Despite the interest however, only a few investigations have been performed with Actinomycetes growing under extreme environments: Actinopolyspora halophila is an accidentally discovered pioneer (Gochnauer et al. 1975). In recent years, researchers from Yunnan Institute of Microbiology at Yunnan University discovered many novel Actinomycetes from salt and alkaline soils in Xinjiang and Qinghai, P. R. China (Jiang and Xu 1996; Jiang et al. 2006). These researchers described a new family Yaniaceae, several novel genera including Streptomonospora, Jiangella, Myceligenerans, Naxibacter, and a great number of new species of the genera Actinopolyspora, Amycolatopsis, Citricoccus, Halomonas, Isoptericola, Jonesia, Kocuria, Kribbella, Liuella, Marinococcus, Massilia, Microbacterium, Nesterenkonia, Nocardia, Nocardiopsis, Prauserella, Rhodococcus, Saccharomonospora, Saccharopolyspora, Sphingomonas, Thermobifida and Virgibacillus. Recently, Meklat et al. (2011) reported a wide spectrum of biologically active halophilic Actinomycetes evaluated using a polyphasic approach which showed the presence of a new genus and many new species of the Actinopolyspora, Nocardiopsis, Saccharomonospora, Streptomonospora and Saccharopolyspora genera. Furthermore, their discovery that from among the rare genera isolated from saline conditions, Nocardiopsis strains having high frequency of NRPS genes could be evidence of the high potential of halophilic Actinomycetes for producing a large number of biologically active compounds.
Caves
Generally, caves are low in nutrients, temperature and light intensity but they have high humidity (Schabereiter-Gurtner et al. 2002). These factors might encourage competition which could enhance the production of substances such as antibiotics and hydrolytic enzymes that inhibit the growth of other microorganisms (Nakaew et al. 2009). Recently, several new species of Actinomycetes have been isolated from caves, including from a gold mine in Korea (Lee et al. 2000a, b, 2001; Lee 2006a, b, c), the Reed Flute Cave in China (Groth et al. 1999), the Grotta Dei Cervi Cave in Italy (Jurado et al. 2005a) and a cave occupied by bats in Spain (Jurado et al. 2005b). Nakaew et al. (2009) reported for the first time the isolation of Spirillospora and Nonomuraea from a cave soil along with very rare genera such as Spirillospora, Catellatospora, Nonomuraea and Micromonospora, and Niyomvong et al. (2012) isolated members of the genera Actinomadura and Saccharopolyspora from caves along with other rare genera Actinoplanes, Gordonia, Microbispora, Micromonospora, Nocardia, Nonomuraea and the predominant genus Streptomyces. These studies confirm that caves may be excellent sources of rare Actinomycetes that produce novel compounds.
Insects
The insect world is another unexplored environment for exploring new and novel microorganisms. Fungi culture in the insect world is practised by ants, termites, beetles and gall midges (Kaltenpoth 2009) and there is evidence that the fungal cultivar produces antibiotics in order to defend itself (Wang et al. 1999; Currie et al. 1999; Little et al. 2006). Ant workers also defend their fungal gardens through a combination of grooming and weeding (Little et al. 2006), producing their own antimicrobials through metapleural gland secretions (Bot et al. 2002) and the application of weedkillers. These weedkillers are natural product antimicrobials produced by symbiotic Actinomycete bacteria (Currie et al. 1999; Sen et al. 2009; Haeder et al. 2009; Oh et al. 2009). A long-standing theory suggests that bacteria from the genus Pseudonocardia co-evolved with the ants and are transmitted vertically by the gynes (reproductive females) along with the fungal cultivar. However, more recent evidence has emerged that suggests attine ants are also associated with bacteria from the Actinomycete genera Streptomyces and Amycolatopsis and that antibiotic-producing Actinomycetes can be horizontally acquired through male dispersal and sampling of Actinomycetes from soils (Currie et al. 1999; Mueller et al. 2008). The identities of the antifungal compounds produced by attine ant-associated Actinomycetes remain largely unknown. Only two compounds have been identified so far: a previously unknown antifungal named ‘Dentigerumycin’ that is produced by Pseudonocardia species isolated from the lower attines Apterostigma dentigerum and ‘Candicidin’, a well-known antifungal that is produced by Streptomyces species isolated from the higher attine ants belonging to the genus Acromyrmex (Haeder et al. 2009; Oh et al. 2009). Pseudonocardia isolated from Acromyrmex octospinosus also inhibit the growth of Escovopsis in bioassays, but the antifungal compounds have not been isolated nor identified (Haeder et al. 2009). Recently, Barke et al. (2010) identified a Pseudonocardia species in the ant Acromyrmex octospinosus that produces an unusual polyene antifungal metabolite. Exploring new bioactive molecules could be increased by switching the search away from explored environments to unexplored ones (Clardy et al. 2009). In this line, the insect world is emerging rapidly as a source to discover Actinomycetes for unusual and novel bioactive molecules.
Aquatic environments
Actinomycetes are predominant in river, lake and marine environments, despite some of them being introduced from terrestrial habitats (Cross 1981). High numbers of Micromonospora, an indigenous inhabitant of the water and mud from freshwater lakes (Cross 1981), can be isolated from lake sediments as much as 10–50 % of the total microbial population in lake water. Nebish Lake had 3,300 bacteria mL−1 of which 15 % was Micromonospora, and Crystal Lake 3,600 bacteria mL−1 with 16 % Micromonospora.
Actinoplanes with sporangium and zoospores will grow at moist conditions and survive as spores in the dry environment (Cross 1981): it colonizes vegetable and animal remains, ranging from pollen and hair to leaves and twigs. Rehydration stimulates the release of zoospores, which swim in the water film of soil or in stream and lake waters until they are able to recolonize a suitable substrate (Cross 1981). Representatives of Thermoactinomyces, Streptomyces and Rhodococcus live in aquatic environments (Cross 1981). Xu and Jiang (1996) studied Actinomycete populations of 12 lakes in the middle plateau of Yunnan (China) and found that Micromonospora was the dominant genus (39–89 %) in the Actinomycetes population in sediments of those lakes. Furthermore, Streptomyces was the second most abundant genus. Members of rare genera Actinoplanes, Actinomadura, Microbispora, Micropolyspora, Microtetraspora, Mycobacterium, Nocardiopsis, Nocardia, Promicromonospora, Rhodococcus, Saccharomonospora, Saccharopolyspora, Streptosporangium, Thermoactinomyces, Thermomonospora and Thermopolyspora have also been reported from lake sediments (Xu and Jiang 1996).
Other habitats
Rare genera of Actinomycetes such as Microbispora, Nocardia, Microtetraspora, Actinomadura, Amycolatopsis and Saccharothrix have been successfully isolated from desert soil (Takahashi et al. 1996), and the novel rare Actinomycete genera Beutenbergia (Groth et al. 1999) and Terrabacter (Lee et al. 2008c) have been reported from small stones collected from caves and agricultural fields, respectively. Recently, rare genera of Actinomycetes such as Streptosporangium, Actinomadura, Saccharopolyspora, Thermoactinomyces and Nocardia were isolated from soils in the nests of solitary wasps and swallow birds (Kumar et al. 2012).
Marine environment: a source of rare Actinomycetes
Many natural environments are still either unexplored or underexplored and thus can be considered as potential resources for the isolation of lesser studied microorganisms, including rare Actinomycetes (Tiwari and Gupta 2012a). Unexplored marine environments, for example, are now a popular research area due to the potentially huge resources present within them. A recent study (Stach and Bull 2005) of the microbial diversity of deep-sea sediments has shown that this environment might contain more than 1,300 different actinobacterial operational taxonomic units, a great proportion of which are predicted to represent novel species and genera. Furthermore, it is recognized that marine microbes can sense, adapt and respond quickly to diverse environments, and can compete for defense and survival by producing unique secondary metabolites (Knight et al. 2003; Zhang et al. 2005). The hidden wealth of this source needs to be explored further.
The historical paradigm of the deep ocean as a biological ‘desert’ has shifted to one of a ‘rainforest’ owing to the isolation of many novel microorganisms and their associated unusual bioactive compounds (Zhang 2005). The marine environment has emerged as an important source of bioactive natural products. There are, for example, several exciting marine-derived molecules on the pharmaceutical market and dozens more progressing through the development pipeline (Mayer et al. 2010). Thus, unexplored and new microbial habitats need to be examined for microbial resources that produce useful bioactive compounds. As with terrestrial soils, marine sediments contain limited amounts of readily available organic matter, with most sources of carbon (such as cellulose and chitin) being present in complex forms. However, culture-independent studies have shown that marine sediment environments contain a wide diversity of Actinomycetes and many unique taxa are very different from their terrestrial counterparts (Stach et al. 2003; Gontang et al. 2007). In addition, culture-dependent studies have shown that marine Actinomycetes are ubiquitous in marine sediment environments (Maldonado et al. 2005; Jensen et al. 2005a). Many novel bioactive secondary metabolites isolated from marine Actinomycetes have been reported (Subramani and Aalbersberg 2012), and they may be a source of novel compounds with pharmaceutical potential (Mayer et al. 2010).
The isolation of a seawater-obligate marine Actinomycete species of the genus Salinispora was reported in 2005 (Maldonado et al. 2005) and that discovery was followed by the discovery of other genera such as Demequina, Marinispora, Solwaraspora, Lamerjespora, Serinicoccus, Salinibacterium, Aeromicrobium, Williamsia, Marinactinospora and Sciscionella that so far appear to be exclusively marine (Subramani and Aalbersberg 2012). Further, these indigenous Actinomycetes are robust sources of natural products, such as the genera Salinispora [salinosporamide A (NPI-0052), sporolides, saliniquinone A-F, salinosporamide K], Verrucosispora (abyssomicins), Micromonospora [diazepinomicin (ECO-4601)] (Lam 2006) and Marinispora (marinomycins, marinisporolides) (Kwon et al. 2009). The discovery of novel marine actinomycetal taxa is very important for potential new sources of pharmaceuticals.
Rare Actinomycetes are widely present in marine habitats (Goodfellow and Williams 1986; Subramani and Aalbersberg 2012). Rare or unusual Actinomycetes produce diverse, unique, unprecedented and occasionally complicated compounds with excellent antibacterial potency and usually low toxicity (Berdy 2005). The oceans represent a rich microbial diversity and population (Stach and Bull 2005; Sogin et al. 2006), and intensive research is ongoing for the microbial biodiversity potential in the marine environment (Heidelberg et al. 2010). Moreover, until now, very few marine obligate taxa have been isolated (Goodfellow 2010). Therefore, oceans are expected to harbour prolific sources of new/novel microbial taxa, and Tiwari and Gupta (2012a) argued that to obtain a novel metabolite, a diverse and less exploited reserve of microbes is required. Isolation of rare Actinomycetes thus becomes the first and most crucial step towards Actinomycetes resource development for drug discovery (Cai et al. 2009).
Marine sediments, seawater, symbiotic and mangroves
Deep-sea sediments cover 63.5 % of the Earth’s surface (Emery 1969) and represent the most undersampled marine habitat (Butman and Carlton 1995). As early as 1884, marine bacterial strains were isolated from deep-sea sediments, to depths of 5,100 m (Zobell 1946). Recently, the concept of ‘marine microorganism’ has been accepted worldwide (Tian et al. 2012), yet the common recognition for ‘marine Actinomycetes’ has undergone a long period of dispute concerning their actual source (Goodfellow and Haynes 1984). Originally, Actinomycetes generally were considered to be indigenous to terrestrial habitats because no convincing evidence was available to demonstrate that Actinomycetes could adapt to marine habitats (Tian et al. 2012). Nevertheless, the novel genus Salinispora (Maldonado et al. 2005) was described and subsequently accepted as the first obligate marine Actinomycetes due to its stringent requirement of seawater for growth. Tian et al. (2009b) described another marine actinobacterial genus, Sciscionella, which can tolerate high salt concentrations (up to 13 %) for growth. To date, more than 14 novel actinobacterial genera have been discovered from the marine environment (Goodfellow and Fiedler 2010; Kurahashi et al. 2010; Chang et al. 2011; Xiao et al. 2011a). It is becoming increasingly obvious that Actinomycetes are an important part of the indigenous microflora in marine ecosystems.
Generally, the pretreatments and enrichment of the samples used for isolation of rare Actinomycetes from soil (see earlier) are the same methods followed for treatment of marine samples. Tables 2, 3, 4, 5 and 6 re-emphasize the pretreatment of samples and enrichment culture methods used, particularly for isolation of marine-derived rare Actinomycetes. This review summarizes the source, treatment of samples and isolation media for all new rare Actinomycetes reported from marine habitats between 2007 and mid-2013, including sediments, seawater, symbiotic and mangrove ecosystems. Wet and dry heat treatments, radiations, cold shock, different chemicals, and antibiotics and 1.5 % phenol-treated marine samples combined with selective isolation media can increase the recovery of new and novel genera of rare Actinomycetes in diverse marine samples (Tables 2, 3, 4, 5 and 6). Interestingly, though observed the combination of selective isolation and screening procedures yielded a number of new rare Actinomycetes genera in marine samples, also noticed that a number of new rare Actinomycete species and even novel genera of rare Actinomycetes were successfully isolated without any pretreatment of the marine samples (Tables 3, 4, 5 and 6). Supportively, Qiu et al. (2008) reported that no matter which pretreatment method was applied, different selective media (particularly HVA, ISP-3 agar and DNB agar) always give better isolation of Micromonospora-like colonies than do other media. Furthermore, these authors found that the yield of non-streptomycete colonies increased in all the composite samples. Ongoing research in our group at the University of the South Pacific in Fiji on the isolation of the marine obligate genus Salinispora has shown that the direct plating of air-dried sediments on different complex nutrient media allows the successful isolation of Salinispora spp. (unpublished data). Therefore, it appears that selective media are playing an important role in the isolation of rare marine Actinomycetes. These results clearly reveal that rare or unusual Actinomycetes are widely dispersed in marine environments and that they have enormous novel actinobacterial diversity which can be readily obtained using conventional isolation methods.
Marine sediments are rich in actinobacterial diversity. A total of 38 new rare Actinomycete species belonging to 15 different Actinomycete families have been reported in marine sediments from the period 2007–mid 2013 (Table 3). Among them, nine novel genera such as Actinotalea, Aestuariimicrobium, Demequina, Marinactinospora, Paraoerskovia, Sciscionella, Marisediminicola, Spinactinospora and Miniimonas were reported. The families reported in marine sediments in the period are Nocardioidaceae (four new species), Micrococcineae (suborder) (five new species), Propionibacteriaceae (three new species), Pseudonocardiaceae (five new species), Nocardiopsaceae (two new species), Cellulomonadaceae (one new species), Promicromonosporaceae (two new species), Micromonosporaceae (five new species), Micrococcaceae (two new species), Microbacteriaceae (two new species), Streptosporangiaceae (one new species), Intrasporangiaceae (two new species), Beutenbergiaceae (one new species), Geodermatophilaceae (one new species) and Nocardiaceae (two new species).
The culturability of microorganisms from seawater is considerably lower (0.001–0.10 %) than that from marine sediments (0.25 %) (Amann et al. 1995). Considering the vast volume of seawater in oceans, the extensive microbial diversity for drug discovery efforts should be extended to explore this resource. A total of 11 new rare Actinomycete species belonging to six different Actinomycete families were reported in seawater from the period 2007 to mid-2013 (Table 4). Among them, four novel genera such as Marihabitans, Ponticoccus, Ornithinibacter and Oceanitalea are reported in seawater. The families reported in seawater between 2007 and mid-2013 are Nocardioidaceae (four new species), Intrasporangiaceae (two new species), Propionibacteriaceae (one new species), Micrococcineae (suborder) (one new species), Micrococcaceae (two new species) and Bogoriellaceae (one new species).
Symbiotic microorganisms—especially Actinomycetes (Schneemann et al. 2010; Izumi et al. 2010; Abdelmohsen et al. 2010) from marine invertebrates, plants and animals—are now rapidly emerging for drug discovery programmes (Piel 2009). The symbiotic microbial community is highly novel and diverse, and species composition shows temporal and geographic variation (Webster and Hill 2001). Even so, very little information exists about the taxonomic affiliation of marine symbiotic microorganisms (Friedrich et al. 1999). Most symbionts are as-yet unculturable, although significant advances have been made in the development of cultivation-independent techniques to study such bacteria. Since these methods will likely have a large impact on future chemical studies of symbionts, they will also be discussed because many symbionts remain unidentified (Piel 2009). Interestingly, two novel families such as Iamiaceae (Kurahashi et al. 2009) and Euzebyaceae (Kurahashi et al. 2010) in Actinobacteria were reported from the sea cucumber, Holothuria edulis (Table 5). A total of 17 new rare Actinomycete species belonging to 11 different Actinomycete families have been reported in plants and animals, respectively, between 2007 and mid-2013 (Table 5). Among them, five novel genera Labedella, Phycicola, Iamia, Euzebya and Koreibacter were reported from marine alga and animals. The families reported in marine plants and animals during 2007–mid-2013 are Nocardioidaceae (two new species), Microbacteriaceae (three new species), Micrococcineae (suborder) (three new species), Micrococcaceae (one new species), Tsukamurellaceae (one new species), Pseudonocardiaceae (one new species), Nocardiopsaceae (two new species), Iamiaceae (one new species), Euzebyaceae (one new species), Alteromonadaceae (one new species) and Micromonosporaceae (one new species).
Mangroves are a unique woody plant community of intertidal coasts in tropical and subtropical zones, located at the transition area between the land and the sea (Holguin et al. 2001; Kathiresan and Bingham 2001). They play a very important role as refuge, feeding and breeding areas for many organisms and sustain an extensive food web based on detritus. The mangrove ecosystem is distinguished from other ecosystems by periodic tidal flooding and variable environmental factors such as salinity, tidal gradients and nutrient availability which are believed to be effective selectors for metabolic pathway adaptations that could generate unusual metabolites (Long et al. 2005). This belief has led to increasing exploitation of the mangrove microorganism resources (Alongi 1988; Long et al. 2005; Holguin et al. 2006). A total of 14 new rare Actinomycete species belonging to seven different families have been reported in mangrove sediments from the period 2007–mid-2013 (Table 6). Among them, two novel genera, Ilumatobacter and Lysinimicrobium, were reported from mangrove sediments. The families reported in mangrove sediments between 2007 and mid-2013 are Micromonosporaceae (seven new species), Acidimicrobiaceae (one new species), Micrococcineae (suborder) (one new species), Promicromonosporaceae (one new species), Streptosporangiaceae (two new species), Thermomonosporaceae (one new species) and Demequinaceae (one new species). Interestingly, Hamada et al. (2012) reported a novel family Demequinaceae from mangrove sediments. Mangrove sediments are an abundant source of Actinomycetes population having versatile producers of various enzymes and antimicrobial molecules (Subramani and Narayanasamy 2009).
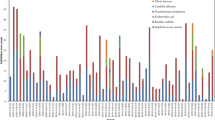
To conclude, a total of 80 new rare Actinomycete species belonging to 23 different rare Actinomycete genera, of which 20 novel genera and 3 novel families, have been reported from marine environments, particularly between 2007 and mid-2013 (Tables 3, 4, 5 and 6; Fig. 1). Furthermore, the family Micromonosporaceae is dominant in marine habitats; genera Nocardioidaceae, Micrococcineae (suborder) and Pseudonocardiaceae are almost as abundant (Fig. 1). The marine environment, representing more than two thirds of the Earth’s surface, is thus a prolific resource for the isolation of less exploited, rare and novel Actinomycetes.
Importance of microbial natural products in novel drug leads
Many of the bacterial pathogens associated with epidemics of human disease have evolved into multidrug-resistant (MDR) forms subsequent to antibiotic use (Davies and Davies 2010). Tuberculosis (TB) is a leading cause of death in the world today and is exacerbated by the prevalence of multi-(MDRTB), extensively (XDR-TB), and totally (TDR-TB) drug-resistant strains. Cancer is the next leading cause of death worldwide. Although more than 30,000 diseases have been clinically described, less than one third of them can be treated symptomatically and fewer can be cured (Schultz and Tsaklakidis 1997). Therefore, the current shortfall in drugs against multidrug-resistant pathogens and other deadly diseases demands urgent attention to develop new antibiotics (Wright and Sutherland 2007). Concern over the paucity of new antibiotics has raised questions regarding the next source of new chemical entities (NCEs) to meet the challenge of continually emerging resistance (Walsh 2003; Macherla et al. 2007). Between 1981 and 2002, the vast majority of NCEs approved for use as antibiotics were natural product derived (Newman et al. 2003), indicating that nature (in particular microorganisms) offers highly relevant scaffolds for developing therapies in the infectious disease arena. While many of the NCEs approved for use at the end of the past century resulted from semi-synthetic modifications to compounds discovered during the ‘Golden Age’ of antibiotics, some recent discoveries indicate that alternative technologies are providing access to new antibiotic scaffolds (Clardy et al. 2006). One approach—which maintains credence in the historic success of microbial-derived NCEs—is to culture new microorganisms from unique natural environments as a source of novel chemistry. A genus previously unexploited from unexplored habitats in the natural product screening collection warrants particular attention, as suggested by Donadio et al. (2002). Recent reports on the isolation and characterization of novel Actinomycetes from poorly researched habitats illustrate the potential of this approach (Bredholt et al. 2008; Eccleston et al. 2008; Okoro et al. 2009). Therefore, screening such organisms and the prospect of discovering new natural products increases which can later be developed as a resource for biotechnology. Despite the vastness of the Earth’s oceans and their inherent biodiversity, the marine environment remains a largely untapped source of new microorganisms, and evidence has emerged that focused exploration of the marine environment will yield unprecedented, chemically prolific species (Fenical and Jensen 2006).
There are more than 22,000 known microbial secondary metabolites, 70 % of which are produced by Actinomycetes, 20 % from fungi, 7 % from Bacillus spp. and 1–2 % by other bacteria. Among the Actinomycetes, the streptomycetes group is economically important because out of the approximately more than 10,000 known antibiotics, 50–55 % are produced by that genus (Berdy 2005; Subramani and Aalbersberg 2012). Actinomycetes are the most economically and biotechnologically useful prokaryotes and hold a prominent position due to their diversity and proven ability to produce novel bioactive compounds (Subramani and Aalbersberg 2012; Blunt et al. 2013). To date, nearly 400 new compounds with cytotoxicity and antimicrobial activity have been isolated from marine Actinomycetes (Proksch and Muller 2006; Fenical and Jensen 2006; Blunt et al. 2009, 2010, 2011). The ecological role of Actinomycetes in the marine ecosystem is largely neglected and various assumptions meant there was little incentive to isolate marine strains for search and discovery of new drugs. The search for and discovery of rare and new Actinomycetes is of significant interest to drug discovery due to a growing need for the development of new and potent therapeutic agents (Subramani and Aalbersberg 2012).
Rare Actinomycetes as a source of new antibiotics
Recently, non-streptomycete Actinomycetes (rare Actinomycetes) have increased significantly up to ~25–30 % share of all known antibiotics (Tishkov 2001; Berdy 2005). Given this, the probability of finding a new compound of economic significance using conventional methodologies of microbial isolation and assay is remote. Efforts to find organisms producing novel antibiotics require either high-throughput screening or specific sampling methods or selections that enrich the unexamined subsets of Actinomycetes (Tiwari and Gupta 2012b). Tiwari and Gupta (2012b) recently reviewed bioactive compounds reported from different genera of rare Actinomycetes obtained from various natural habitats. They conclude that many of the successful antimicrobial agents currently available in the market are produced by rare Actinomycetes, like rifamycins by Amycolatopsis mediterranei, erythromycin by Saccharopolyspora erythraea, teicoplanin by Actinoplanes teichomyceticus, vancomycin by Amycolatopsis orientalis, gentamicin from Micromonopsora purpurea and a chronological sequence of antibiotic compounds discovered as products of Micromonospora spp., Actinoplanes spp. and Streptosporangium spp. (Cooper et al. 1990; Lancini and Lorenzetti 1993; Lazzarini et al. 2000; Pfefferle et al. 2000). Among the available rare Actinomycetes genera, Amycolatopsis, Saccharopolyspora, Actinoplanes and Micromonopsora have been exploited as a prolific source of novel secondary metabolites (Geok et al. 2007; Murakami et al. 2007; Renu et al. 2008; Zhuge et al. 2008; Igarashi et al. 2008; Berdnikova et al. 2009; Beth et al. 2009; Liras and Demain 2009; Zhang et al. 2009; Dharmendra et al. 2010; Dasari et al. 2012); however, lesser exploited rare genera such as Actinomadura, Nocardiopsis, Dactylosporangium, Kibdelosporangium, Microbispora, Kitasatospora, Planomonospora, Planobispora, Salinispora, Marinispora, Serinicoccus and Verrucosispora are now drawing attention. These impacts emphasize the need to continue research in this area and the investments in rare Actinomycetes can be considered as being completely warranted.
Novel/new metabolites from marine rare Actinomycetes
This review also tried to update the information on rare Actinomycetes obtained from marine habitats and the antibiotic compounds identified from other groups of marine rare Actinomycetes during 2007–mid-2013. Table 7 shows some examples of new bioactive metabolites isolated from marine rare Actinomycetes from 2007 to mid-2013. This is by no means an exhaustive search of all novel secondary metabolites produced by marine rare Actinomycetes genera during this 6-year period; nevertheless, this list is impressive and illustrates the many different diverse structures with biological activities reported. Among them, a few compounds such as groups of abyssomicins, proximicins, thiocoralines and gifhornenolones produced by Verrucosispora spp. and lipoxazolidinones, lynamicins and marinisporolides produced by Marinispora spp. (Figs. 2, 3 and 4) are of particular interest due to their rarity, potency and diverse bioactivity. The recently isolated rare and first marine obligate genus Salinispora produced an array of novel metabolites which have previously been discussed (Subramani and Aalbersberg 2012).
Now, emphasizing another interesting rare Actinomycete genus Verrucosispora is quite limited presumably due to its limited distribution in the marine environment. Recently, Verrucosispora spp. produced an array of new and novel abyssomicins (Fig. 2), a new class of unique polycyclic natural products with potent antibacterial, antitubercular, antitumor and anti-Bacille Calmette Guerin activity (Keller et al. 2007a, b; Wang et al. 2013b). Abyssomicins are of great significance since these molecules are the first to inhibit biosynthesis of para-aminobenzoic acid biosynthetic pathway, a pathway essential for many microorganisms but absent in humans (Riedlinger et al. 2004; Keller et al. 2007b). Ongoing interest in the synthesis, biosynthesis and pharmacology of the abyssomicins has fuelled further exploration of this interesting class of compounds and perhaps may lead to related derivatives with better biological profiles (Wang et al. 2013b). The recent first complete genome sequence of Verrucosispora sp. increased the expectancy from this group of strains in novel biodiscovery efforts (Roh et al. 2011).
Proximicins (Fig. 3), novel aminofuran antibiotics also produced by Verrucosispora spp., bear the hitherto unknown γ-amino acid 4-aminofuran-2-carboxylic acid moeity, which adds a new element of structural diversity to the previously described heterocyclic antibiotics (Fiedler et al. 2008; Schneider et al. 2008). The biological activity of proximicins did not show appreciable antibacterial activity against drug-resistant human pathogens. However, they displayed potent antitumor activity against a range of human tumor cell lines.
Gifhornenolones A and B (Fig. 2) are new terpenoids isolated from the marine ascidian-associated Verrucosispora gifhornensis. The biological activity of gifhornenolone A showed potent inhibitory activity to the androgen receptor (Shirai et al. 2010).
Thiochondrillines (Fig. 3), analogs of thiocoraline, are potent cytotoxic thiodepsipeptides isolated from the sponge-associated Verrucosispora sp. (Wyche et al. 2011). The marine environment, which harbours over 20 million microbes (Qui 2010), has provided several microbial-derived compounds, such as salinosporamide A (Feling et al. 2003), TZT-1027 (Kobayashi et al. 1997) and ILX-651 (Mita et al. 2006) that are currently in clinical trials (Mayer et al. 2010). Among the list of microbial-derived marine natural products with therapeutic relevance is thiocoraline, a potential candidate for clinical trials (Faircloth et al. 1997). Thiocoraline and its analogs have potent cytotoxic properties against a wide range of human cancer cell lines (Romero et al. 1997; Erba et al. 1999; Negri et al. 2007; Wyche et al. 2011).
Lipoxazolidinones A–C (Fig. 4) are novel 2-alkylidene-5-alkyl-4-oxazolidinones isolated from novel and rare genus Marinispora (Macherla et al. 2007). The biological activity of lipoxazolidinones exhibited broad spectrum antimicrobial activity similar to that of the commercial antibiotic linezolid (Zyvox), a 2-oxazolidinone (Macherla et al. 2007). Hydrolysis of the amide bond of the 4-oxazolidinone ring of lipoxazolidinone A resulted in loss of antibacterial activity. The 2-alkylidene-4-oxazolidinone represents a new antibiotic pharmacophore and is unprecedented in nature.
Lynamicins A–E (Fig. 4) are chlorinated bisindole pyrroles isolated from the rare Actinomycete Marinispora sp. (McArthur et al. 2008). The antimicrobial spectrum of lynamicins was evaluated against a panel of 11 pathogens, which demonstrated that these substances possess broad spectrum activity against both Gram-positive and Gram-negative pathogens. Significantly, lynamicins were active against drug-resistant pathogens such as methicillin-resistant Staphylococcus aureus and vancomycin-resistant Enterococcus faecium (McArthur et al. 2008).
In addition, marinisporolides A and B (Fig. 4) are polyene-polyol macrolides also isolated from Marinispora sp. (Kwon et al. 2009). The marinisporolides are 34-membered macrolides composed of a conjugated pentaene and several pairs of 1,3-dihydroxyl functionalities and show interesting photoreactivity and chiroptical properties. Marinisporolide A contains a bicyclic spiro-bis-tetrahydropyran ketal functionality, while marinisporolide B is the corresponding hemiketal.
These highlighted structures, chemical diversity, biological properties and discovery of these new compounds (Table 7; Figs. 2, 3 and 4) continue to indicate that rare and new/novel Actinomycetes of the genera will be a significant resource for structurally/biologically interesting molecules.
Conclusions
Over the past three decades, the marine environment has continuously been providing a number of new/novel Actinomycetes and bioactive compounds, but the potential of this area still remains virtually unexplored. Until recently, microbiologists were greatly limited in their study of natural microbial ecosystems due to an inability to cultivate most naturally occurring microorganisms (Cragg and Newman 2005). The marine environment is huge and harbours an enormous hidden microbial diversity. As-yet undiscovered and unusual or rare microorganisms may contain possible cures for diseases demanding new antibiotics to combat the multidrug-resistant human pathogens and emerging deadly diseases. Application of selective isolation and enriched methods can lead to the discovery of new/novel and rare bioactive Actinobacteria from marine ecological niches having the potential to biosynthesize novel bioactive compounds. As summarized in this review, a combination of different pretreatment techniques along with suitable selective isolation media, enrichment culture supplemented with specific antibiotics, enabled the isolation of rare and novel Actinomycetes and the production of unusual bioactive metabolites.
Furthermore as reviewed above, the marine environment contains a myriad of new and rare Actinobacteria providing novel structural diversity waiting to be discovered and used in the biotechnological and pharmaceutical industries. Even so, the study on marine rare Actinobacteria is just beginning. Researchers are in the early stages of a renaissance in natural product discovery from marine Actinobacteria. It is now known that new Actinomycete taxa occur in the ocean and that some display specific adaptations for their life in the marine environment (Mincer et al. 2002; Jensen et al. 2005a, 2007). These taxa include the chemically prolific genera Salinispora and Marinispora which produce exciting new and novel structural classes of secondary metabolites. In this line, another rare Actinomycete genus, Verrucosispora, is also proving to be a productive source of new metabolites such as the abyssomicins. In addition, the rare Actinomycetes obtained from marine sediments are metabolically active and produce interesting bioactive molecules (Dai et al. 2010; Goodfellow et al. 2012a, b; Tian et al. 2013). These results provide clear evidence that targeting rare and new/novel marine Actinomycete genera and species will lead to the discovery of new chemotypes with significant biological activity and the potential to become leads for drug discovery.
References
Abdel-Mageed WM, Milne BF, Wagner M, Schumacher M, Sandor P, Pathom-aree W, Goodfellow M, Bull AT, Horikoshi K, Ebel R, Diederich M, Fiedler HP, Jaspars M (2010) Dermacozines, a new phenazine family from deep-sea Dermacocci isolated from a Mariana Trench sediment. Org Biomol Chem 8:2352–2362
Abdelmohsen UR, Pimentel-Elardo SM, Hanora A, Radwan M, Abou-El-Ela SH, Ahmed S, Hentschel U (2010) Isolation, phylogenetic analysis and anti-infective activity screening of marine sponge-associated Actinomycetes. Mar Drugs 8:399–412
Alferova IV, Terekhova LP (1988) Use of the method of enriching of soil samples with calcium carbonate for isolation of Actinomyces. Antibiot Khimioter 33:888–890
Alongi DM (1988) Bacterial productivity and microbial biomass in tropical mangrove sediments. Microb Ecol 15:59–79
Amann RI, Ludwig W, Schleifer KH (1995) Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev 59:143–169
Asolkar RN, Kirkland TN, Jensen PR, Fenical W (2010) Arenimycin, an antibiotic effective against rifampin- and methicillin-resistant Staphylococcus aureus from the marine Actinomycete Salinispora arenicola. J Antibiot (Tokyo) 63:37–39
Bala M, Kaur C, Kaur I, Khan F, Mayilraj S (2011) Kocuria sediminis sp. nov., isolated from a marine sediment sample. Antonie Van Leeuwenhoek 101:469–478
Baltz RH (2006) Marcel Faber Roundtable: is our antibiotic pipeline unproductive because of starvation, constipation or lack of inspiration? J Ind Microbiol Biotechnol 33:507–513
Baltz RH (2007) A call to arms. Nat Rev Drug Discov 6:8–12
Baltz RH (2011) Strain improvement in Actinomycetes in the postgenomic era. J Ind Microbiol Biotechnol 38:657–666
Barke J, Seipke RF, Grüschow S, Heavens D, Drou N, Bibb MJ, Goss RJM, Yu DW, Hutchings MI (2010) A mixed community of Actinomycetes produce multiple antibiotics for the fungus farming ant Acromyrmex octospinosus. BMC Biol 8:109–118
Baskaran R, Vijayakumar R, Mohan PM (2011) Enrichment method for the isolation of bioactive Actinomycetes from mangrove sediments of Andaman Islands, India. Malaysian J Microbiol 7:26–32
Bentley SD, Chater KF, Cerdeño-Tárraga AM, Challis GL, Thomson NR, James KD, Harris DE, Quail MA, Kieser H, Harper D, Bateman A, Brown S, Chandra G, Chen CW, Collins M, Cronin A, Fraser A, Goble A, Hidalgo J, Hornsby T, Howarth S, Huang CH, Kieser T, Larke L, Murphy L, Oliver K, O’Neil S, Rabbinowitsch E, Rajandream MA, Rutherford K, Rutter S, Seeger K, Saunders D, Sharp S, Squares R, Squares S, Taylor K, Warren T, Wietzorrek A, Woodward J, Barrell BG, Parkhill J, Hopwood DA (2002) Complete genome sequence of the model Actinomycete Streptomyces coelicolor A3(2). Nature 417:141–147
Berdnikova TF, Shashkov AS, Katrukha GS, Lapchinskaia OA, Iurkevich NV, Grachev AA, Nifantev NE (2009) The structure of antibiotic Eremomycin B. Bioorg Khim 35:550–556
Berdy J (2005) Bioactive microbial metabolites. J Antibiot (Tokyo) 58:1–26
Beth J, Andre W, Michelle H, Mike L, Jens C, Neal C (2009) Actinomycetes scale-up for the production of the antibacterial, Nocathiacin. Biotechnol Prog 25:176–188
Bian J, Li Y, Wang J, Song FH, Liu M, Dai HQ, Ren B, Gao H, Hu X, Liu ZH, Li WJ, Zhang LX (2009) Amycolatopsis marina sp. nov., an Actinomycete isolated from an ocean sediment. Int J Syst Evol Microbiol 59:477–481
Blunt JW, Copp BR, Hu WP, Munro MH, Northcote PT, Prinsep MR (2009) Marine natural products. Nat Prod Rep 26:170–244
Blunt JW, Copp BR, Munro MH, Northcote PT, Prinsep MR (2010) Marine natural products. Nat Prod Rep 27:165–237
Blunt JW, Copp BR, Munro MH, Northcote PT, Prinsep MR (2011) Marine natural products. Nat Prod Rep 28:196–268
Blunt JW, Copp BR, Keyzers RA, Munro MHG, Prinsep MR (2013) Marine natural products. Nat Prod Rep 30:237–323
Bot ANM, Ortius-Lechner D, Finster K, Maile R, Boomsma JJ (2002) Variable sensitivity of fungi and bacteria to compounds produced by the metapleural glands of leaf-cutting ants. Insect Soc 49:363–370
Bredholdt H, Galatenko OA, Engelhardt K, Fjaervik E, Terekhova LP, Zotchev SB (2007) Rare Actinomycete bacteria from the shallow water sediments of the Trondheim fjord, Norway: isolation, diversity and biological activity. Environ Microbiol 9:2756–2764
Bredholt H, Fjaervik E, Johnsen G, Zotchev SB (2008) Actinomycetes from sediments in the Trondheim Fjord, Norway: diversity and biological activity. Mar Drugs 6:12–24
Bulina TI, Alferova IV, Terekhova LP (1997) A novel approach to isolation of Actinomycetes involving irradiation of soil samples with microwaves. Microbiology 66:231–234
Bull AT, Stach JE, Ward AC, Goodfellow M (2005) Marine Actinobacteria: perspectives, challenges, future directions. Antonie Van Leeuwenhoek 87:65–79
Butman CA, Carlton JT (1995) Marine biological diversity—some important issues, opportunities and critical research needs. Rev Geophys 33:1201–1209
Cai Y, Xue Q, Chen Z, Zhang R (2009) Classification and salt-tolerance of Actinomycetes in the Qinghai lake water and lakeside saline soil. J Sustainable Dev 2:107–110
Carlson S, Marler L, Nam SJ, Santarsiero BD, Pezzuto JM, Murphy BT (2013) Potential chemopreventive activity of a new macrolide antibiotic from a marine-derived Micromonospora sp. Mar Drugs 11:1152–1161
Chang XB, Liu WZ, Zhang XH (2011) Spinactinospora alkalitolerans gen. nov. sp. nov., an Actinomycete isolated from marine sediment. Int J Syst Evol Microbiol 61:2805–2810
Charusanti P, Fong NL, Nagarajan H, Pereira AR, Li HJ, Abate EA, Su Y, Gerwick WH, Palsson BO (2012) Exploiting adaptive laboratory evolution of Streptomyces clavuligerus for antibiotic discovery and overproduction. PLoS One 7:e33727
Chen YG, Tang SK, Zhang YQ, Li ZY, Yi LB, Wang YX, Li WJ, Cui XL (2009a) Arthrobacter halodurans sp. nov., a new halotolerant bacterium isolated from sea water. Antonie Van Leeuwenhoek 96:63–70
Chen YG, Wang YX, Zhang YQ, Tang SK, Liu ZX, Xiao HD, Xu LH, Cui XL, Li WJ (2009b) Nocardiopsis litoralis sp. nov., a halophilic marine Actinomycete isolated from a sea anemone. Int J Syst Evol Microbiol 59:2708–2713
Cho SH, Hwang CW, Chung HK, Yang CS (1994) A new medium for the selective isolation of soil Actinomycetes. K J Appl Microbiol Biotechnol 22:561–563
Cho JY, Williams PG, Kwon HC, Jensen PR, Fenical W (2007) Lucentamycins A-D, cytotoxic peptides from the marine-derived Actinomycete Nocardiopsis lucentensis. J Nat Prod 70:1321–1328
Choi DH, Kim HM, Noh JH, Cho BC (2007) Nocardioides marinus sp. nov. Int J Syst Evol Microbiol 57:775–779
Clardy J, Fischbach MA, Walsh CT (2006) New antibiotics from bacterial natural products. Nat Biotechnol 24:1541–1550
Clardy J, Fischbach MA, Currie CR (2009) The natural history of antibiotics. Curr Biol 19:R437–R441
Cooper R, Das P, Federbush C, Mierzwa R, Patel M, Pramanik B, Truumees I (1990) Characterization of peptidyl-nucleoside antifungal antibiotics from fermentation broth. J Ind Microbiol 5:1–8
Cragg GM, Newman DJ (2005) Biodiversity: a continuing source of novel drug leads. Pure Appl Chem 77:7–24
Cross T (1981) Aquatic Actinomycetes: a critical survey of the occurrence, growth and role of Actinomycetes in aquatic habitats. J Appl Bacteriol 50:379–423
Currie CR, Scott JA, Summerbell RC, Malloch D (1999) Fungus-growing ants use antibiotic-producing bacteria to control garden parasites. Nature 398:701–705
D’Onofrio A, Crawford JM, Stewart EJ, Witt K, Gavrish E, Epstein S, Clardy J, Lewis K (2010) Siderophores from neighboring organisms promote the growth of uncultured bacteria. Chem Biol 17:254–264
Dai HQ, Wang J, Xin YH, Pei G, Tang SK, Ren B, Ward A, Ruan JS, Li WJ, Zhang LX (2010) Verrucosispora sediminis sp. nov., a cyclodipeptide-producing Actinomycete from deep-sea sediment. Int J Syst Evol Microbiol 60:1807–1812
Dasari VR, Muthyala MK, Nikku MY, Donthireddy SR (2012) Novel pyridinium compound from marine Actinomycete, Amycolatopsis alba var. nov. DVR D4 showing antimicrobial and cytotoxic activities in vitro. Microbiol Res 167:346–351
Davies J, Davies D (2010) Origins and evolution of antibiotic resistance. Microbiol Mol Biol Rev 74:417–433
Demain AL, Sanchez S (2009) Microbial drug discovery: 80 years of progress. J Antibiot 62:5–16
Dharmendra SD, Jesus MF, Gary TMH (2010) Influence of cultivation conditions on the production of a thermostable extracellular lipase from Amycolatopsis mediterranei DSM 43304. J Ind Microbiol Biotechnol 37:1–17
Donadio S, Monciardini P, Alduina R, Mazza P, Chiocchini C, Cavaletti L, Sosio M, Puglia AM (2002) Microbial technologies for the discovery of novel bioactive metabolites. J Biotechnol 99:187–198
Du ZJ, Lv GQ, Rooney AP, Miao TT, Xu QQ, Chen GJ (2011) Agarivorans gilvus sp. nov. isolated from seaweed. Int J Syst Evol Microbiol 61:493–496
Eccleston GP, Brooks PR, Kurtböke DI (2008) The occurrence of bioactive Micromonosporae in aquatic habitats of the Sunshine Coast in Australia. Mar Drugs 6:243–261
El-Gendy MM, Hawas UW, Jaspars M (2008) Novel bioactive metabolites from a marine derived bacterium Nocardia sp. ALAA 2000. J Antibiot (Tokyo) 61:379–386
Emery KO (1969) The continental shelves. Sci Am 221:106–122
Engelhardt K, Degnes KF, Kemmler M, Bredholt H, Fjaervik E, Klinkenberg G, Sletta H, Ellingsen TE, Zotchev SB (2010) Production of a new thiopeptide antibiotic, TP-1161, by a marine Nocardiopsis species. Appl Environ Microbiol 76:4969–4976
Erba E, Bergamaschi D, Ronzoni S, Faretta M, Taverna S, Bonfanti M, Catapano CV, Faircloth G, Jimeno J, D’Incalci M (1999) Mode of action of thiocoraline, a natural marine compound with anti-tumour activity. Br J Cancer 80:971–980
Eustaquio AS, Nam SJ, Penn K, Lechner A, Wilson MC, Fenical W, Jensen PR, Moore BS (2011) The discovery of salinosporamide K from the marine bacterium Salinispora pacifica by genome mining gives insight into pathway evolution. ChemBioChem 12:61–64
Faircloth G, Jimeno J, D’Incalci M (1997) Biological activity of thiocoraline, a novel marine depsipeptide. Eur J Cancer 33:S175
Feling RH, Buchanan GO, Mincer TJ, Kauffman CA, Jensen PR, Fenical W (2003) Salinosporamide A: a highly cytotoxic proteasome inhibitor from a novel microbial source, a marine bacterium of the new genus Salinospora. Angew Chem Int Ed Engl 42:355–357
Fenical W, Jensen PR (2006) Developing a new resource for drug discovery: marine Actinomycete bacteria. Nat Chem Biol 2:666–673
Ferriss RS (1984) Effects of microwave oven treatment on microorganisms in soil. Phytopathology 74:121–126
Fiedler HP, Bruntner C, Riedlinger J, Bull AT, Knutsen G, Goodfellow M, Jones AL, Maldonado L, Pathom-aree W, Beil W, Schneider K, Keller S, Sussmuth RD (2008) Proximicin A, B and C, novel aminofuran antibiotic and anticancer compounds isolated from marine strains of the Actinomycete Verrucosispora. J Antibiot 61:158–163
Finster KW, Cockell CS, Voytek MA, Gronstal AL, Kjeldsen KU (2009) Description of Tessaracoccus profundi sp. nov., a deep-subsurface Actinobacterium isolated from a Chesapeake impact crater drill core (940 m depth). Antonie Van Leeuwenhoek 96:515–526
Fischbach MA, Walsh CT (2009) Antibiotics for emerging pathogens. Science 325:1089–1093
Friedrich AB, Merkert H, Fendert T, Hacker J, Proksch P, Hentschel U (1999) Microbial diversity in the marine sponge Aplysina cavernicola (formerly Verongia cavernicola) analyzed by fluorescence in situ hybridization (FISH). Mar Biol 134:461–470
Fu P, Wang S, Hong K, Li X, Liu P, Wang Y, Zhu W (2011) Cytotoxic bipyridines from the marine-derived Actinomycete Actinoalloteichus cyanogriseus WH1-2216-6. J Nat Prod 74:1751–1756
Fu Y, Li Q, Liu K, Xu Y, Wang Y, Jiao N (2012) Oceanitalea nanhaiensis gen. nov., sp. nov., an Actinobacterium isolated from seawater. Int J Syst Evol Microbiol 62:2490–2494
Gärtner A, Ohlendorf B, Schulz D, Zinecker H, Wiese J, Imhoff JF (2011) Levantilides A and B, 20-membered macrolides from a Micromonospora strain isolated from the mediterranean deep sea sediment. Mar Drugs 9:98–108
Gavrish E, Bollmann A, Epstein S, Lewis K (2008) A trap for in situ cultivation of filamentous Actinobacteria. J Microbiol Methods 72:257–262
Geok YAT, Stuart R, Ernest L, Roselyn B, Wonyong K, Michael G (2007) Amycolatopsis regifaucium sp. nov., a novel Actinomycete that produces Kigamicin. Int J Syst Bacteriol 57:2562–2567
Gochnauer MB, Leppard GG, Komaratat P, Kates M, Novitsky T, Kushner DJ (1975) Isolation and characterization of Actinopolyspora halophila, gen. et sp. nov., an extremely halophilic Actinomycete. Can J Microbiol 21:1500–1511
Gontang EA, Fenical W, Jensen PR (2007) Phylogenetic diversity of Gram-positive bacteria cultured from marine sediments. Appl Environ Microbiol 73:3272–3282
Goodfellow M (2010) Selective isolation of Actinobacteria. In: Baltz RH, Davies J, Demain AL (eds) Manual of industrial microbiology and biotechnology. Section 1: isolation and screening of secondary metabolites and enzymes. Bull AT, Davies JE (section eds). ASM, Washington, 3:13-27
Goodfellow M, Fiedler HP (2010) A guide to successful bioprospecting: informed by actinobacterial systematics. Antonie Van Leeuwenhoek 98:119–142
Goodfellow M, Haynes JA (1984) Actinomycetes in marine sediments. In: Ortiz-Ortiz L (ed) Biological, biochemical and biomedical aspects of Actinomycetes. Academic, New York, pp 453–472
Goodfellow M, O’Donnel AG (1989) Search and discovery of industrially significant Actinomycetes. In: Baumberg S, Hunter IS, Rhodes PM (eds) Microbial products. New approaches, society for general microbiology symposium No. 44. Cambridge University Press, Cambridge, pp 343–383
Goodfellow M, Williams E (1986) New strategies for the selective isolation of industrially important bacteria. Biotechnol Genet Eng Rev 4:213–262
Goodfellow M, Brown R, Ahmed L, Pathom-Aree W, Bull AT, Jones AL, Stach JE, Zucchi TD, Zhang L, Wang J (2012a) Verrucosispora fiedleri sp. nov., an Actinomycete isolated from a fjord sediment which synthesizes proximicins. Antonie Van Leeuwenhoek 103:493–502
Goodfellow M, Stach JE, Brown R, Bonda AN, Jones AL, Mexson J, Fiedler HP, Zucchi TD, Bull AT (2012b) Verrucosispora maris sp. nov., a novel deep-sea Actinomycete isolated from a marine sediment which produces abyssomicins. Antonie Van Leeuwenhoek 101:185–193
Groth I, Schumann P, Schuetze B, Augsten K, Kramer I, Stackebrandt E (1999) Beutenbergia cavernae gen. nov., sp. nov., an L-lysine-containing Actinomycete isolated from a cave. Int J Syst Bacteriol 49:1733–1740
Haeder S, Wirth R, Herz H, Spiteller D (2009) Candicidin-producing Streptomyces support leaf-cutting ants to protect their fungus garden against the pathogenic fungus Escovopsis. Proc Natl Acad Sci U S A 106:4742–4746
Hamada M, Tamura T, Yamamura H, Suzuki K, Hayakawa M (2012) Lysinimicrobium mangrovi gen. nov., sp. nov., an Actinobacterium isolated from the rhizosphere of a mangrove. Int J Syst Evol Microbiol 62:1731–1735
Hamada M, Tamura T, Yamamura H, Suzuki K, Hayakawa M (2013) Demequina flava sp. nov. and Demequina sediminicola sp. nov., isolated from sea sediment. Int J Syst Evol Microbiol 63:249–253
Hamaki T, Suzuki M, Fudou R, Jojima Y, Kajiura T, Tabuchi A, Sen K, Shibai H (2005) Isolation of novel bacteria and Actinomycetes using soil-extract agar medium. J Biosci Bioeng 99:485–492
Hayakawa M (1990) Studies on selective isolation methods and distribution of soil Actinomycetes. Actinomycetol 4:103–112
Hayakawa M (1994) Selective isolation methods for rare Actinomycetes (in Japanese). In: Hotta K, Horinouchi S (eds) Bioscience and Actinomycetes. Igakusyuppan Center, Tokyo, pp 219–229
Hayakawa M (2003) Selective isolation of rare Actinomycetes genera using pretreatment techniques. In: Kurtböke I (ed) Selective isolation of rare Actinomycetes. Queensland Complete Printing Service, Nambour, pp 55–81
Hayakawa M (2008) Studies on the isolation and distribution of rare Actinomycetes in soil. Actinomycetol 22:12–19
Hayakawa M, Nonomura H (1987a) Humic acid-vitamin agar, a new medium for the selective isolation of soil Actinomycetes. J Ferment Technol 65:501–509
Hayakawa M, Nonomura H (1987b) Efficacy of artificial humic acid as a selective nutrient in HV agar used for the isolation of soil Actinomycetes. J Ferment Technol 65:609–616
Hayakawa M, Nonomura H (1993) In: Beppu T, Seino A, Hotta K (eds) Selective isolation methods for soil Actinomycetes (in Japanese). Japan Scientific Societies, Tokyo
Hayakawa M, Ishizawa K, Nonomura H (1988) Distribution of rare Actinomycetes in Japanese soils. J Ferment Technol 66:367–373
Hayakawa M, Kajiura T, Nonomura H (1991a) New methods for the highly selective isolation of Streptosporangium and Dactylosporangium from soil. J Ferment Bioeng 72:327–333
Hayakawa M, Sadakata T, Kajiura T, Nonomura H (1991b) New methods for the highly selective isolation of Micromonospora and Microbispora from soil. J Ferment Bioeng 72:320–326
Hayakawa M, Tamura T, Iino H, Nonomura H (1991c) Pollen-baiting and drying method for highly selective isolation of Actinoplanes spp. from soil. J Ferment Bioeng 72:433–438
Hayakawa M, Tamura T, Nonomura H (1991d) Selective isolation of Actinoplanes and Dactylosporangium from soil by using γ-collidine as the chemoattractant. J Ferment Bioeng 72:426–432
Hayakawa M, Otoguro M, Takeuchi T, Yamazaki T, Iimura Y (2000) Application of a method incorporating differential centrifugation for selective isolation of motile Actinomycetes in soil and plant litter. Antonie Van Leeuwenhoek 78:171–185
Hayakawa M, Yoshida Y, Iimura Y (2004) Selection of bioactive soil Actinomycetes belonging to the Streptomyces violaceusniger phenotypic cluster. J Gen Appl Microbiol 96:973–981
He J, Xu Y, Sahu MK, Tian XP, Nie GX, Xie Q, Zhang S, Sivakumar K, Li WJ (2012) Actinomadura sediminis sp. nov., a marine Actinomycete isolated from mangrove sediment. Int J Syst Evol Microbiol 62:1110–1116
Heidelberg KB, Gilbert JA, Joint I (2010) Marine genomics: at the interface of marine microbial ecology and biodiscovery. Microb Biotechnol 3:531–543
Holguin G, Vazquez P, Bashan Y (2001) The role of sediment microorganisms in the productivity, conservation, and rehabilitation of mangrove ecosystems: an overview. Biol Fertil Soils 33:265–278
Holguin G, Gonzalez-Zamorano P, de-Bashan LE, Mendoza R, Amador E, Bashan Y (2006) Mangrove health in an arid environment encroached by urban development—a case study. Sci Total Environ 363:260–274
Hong K, Gao AH, Xie QY, Gao H, Zhuang L, Lin HP, Yu HP, Li J, Yao XS, Goodfellow M, Ruan JS (2009) Actinomycetes for marine drug discovery isolated from mangrove soils and plants in China. Mar Drugs 7:24–44
Huang H, Lv J, Hu Y, Fang Z, Zhang K, Bao S (2008) Micromonospora rifamycinica sp. nov., a novel Actinomycete from mangrove sediment. Int J Syst Evol Microbiol 58:17–20
Huang H, Wu X, Yi S, Zhou Z, Zhu J, Fang Z, Yue J, Bao S (2009) Rifamycin S and its geometric isomer produced by a newly found Actinomycete, Micromonospora rifamycinica. Antonie Van Leeuwenhoek 95:143–148
Igarashi M, Sawa R, Kinoshita N, Hashizume H, Nakagawa N, Homma Y, Nishimura Y, Akamatsu Y (2008) Pargamicin A, a novel cyclic peptide antibiotic from Amycolatopsis sp. J Antibiot 61:387–393
Ikeda H, Ishikawa J, Hanamoto A, Shinose M, Kikuchi H, Shiba T, Sakaki Y, Hattori M, Omura S (2003) Complete genome sequence and comparative analysis of the industrial microorganism Streptomyces avermitilis. Nat Biotechnol 21:526–531
Inahashi Y, Matsumoto A, Omura S, Takahashi Y (2011) Streptosporangium oxazolinicum sp. nov., a novel endophytic Actinomycete producing new antitrypanosomal antibiotics, spoxazomicins. J Antibiot 64:297–302
Istianto Y, Koesoemowidodo RSA, Saputra H, Watanabe Y, Pranamuda H, Marwoto B (2012) Application of phenol pretreatment for the isolation of rare Actinomycetes from Indonesian soil. Microbiology (Indonesia) 6:42–47
Izumi H, Gauthier MEA, Degnan BM, Ng YK, Hewavitharana AK, Shaw PN, Fuerst JA (2010) Diversity of Mycobacterium species from marine sponges and their sensitivity to antagonism by sponge-derived rifamycin synthesizing Actinobacterium in the genus Salinispora. FEMS Microbiol Lett 313:33–40
Janso JE, Carter GT (2010) Biosynthetic potential of phylogenetically unique endophytic Actinomycetes from tropical plants. Appl Environ Microbiol 76:4377–4386
Jensen PR, Gontang E, Mafnas C, Mincer TJ, Fenical W (2005a) Culturable marine Actinomycetes diversity from tropical Pacific Ocean sediments. Environ Microbiol 7:1039–1048
Jensen PR, Mincer TJ, Williams PG, Fenical W (2005b) Marine Actinomycete diversity and natural product discovery. Antonie Van Leeuwenhoek 87:43–48
Jensen PR, Williams PG, Oh DC, Zeigler L, Fenical W (2007) Species-specific secondary metabolite production in marine Actinomycetes of the genus Salinispora. Appl Environ Microbiol 73:1146–1152
Jiang CL, Xu LH (1996) Diversity of soil Actinomycetes in Yunnan, China. Appl Environ Microbiol 62:244–248
Jiang Y, Li WJ, Xu P, Tang SK, Xu LH (2006) Study on Actinomycete diversity under salt and alkaline environments. Wei Sheng Wu Xue Bao 46:191–195
Jiang Y, Wiese J, Cao YR, Xu LH, Imhoff JF, Jiang CL (2009) Promicromonospora flava sp. nov., isolated from sediment of the Baltic Sea. Int J Syst Evol Microbiol 59:1599–1602
Jiang Y, Cao Y, Zhao L, Wang Q, Jin R, He W, Xue Q (2010) Ultrasonic treatment of soil samples for Actinomycete isolation. Wei Sheng Wu Xue Bao 50:1094–1097
Jung SY, Kim HS, Song JJ, Lee SG, Oh TK, Yoon JH (2007) Aestuariimicrobium kwangyangense gen. nov., sp. nov., an LL-diaminopimelic acid-containing bacterium isolated from tidal flat sediment. Int J Syst Evol Microbiol 57:2114–2118
Jurado V, Groth I, Gonzalez JM, Laiz L, Saiz-Jimenez C (2005a) Agromyces salentinus sp. nov. and Agromyces neolithicus sp. nov. Int J Syst Evol Microbiol 55:153–157
Jurado V, Groth I, Gonzalez JM, Laiz L, Saiz-Jimenez C (2005b) Agromyces subbeticus sp. nov., isolated from a cave in southern Spain. Int J Syst Evol Microbiol 55:1897–1901
Kaeberlein T, Lewis K, Epstein SS (2002) Isolating “uncultivable” microorganisms in pure culture in a simulated natural environment. Science 296:1127–1129
Kageyama A, Haga T, Kasai H, Shizuri Y, Omura S, Takahashi Y (2008) Marihabitans asiaticum gen. nov., sp. nov., a meso-diaminopimelic acid-containing member of the family Intrasporangiaceae. Int J Syst Evol Microbiol 58:2429–2432
Kaltenpoth M (2009) Actinobacteria as mutualists: general healthcare for insects? Trends Microbiol 17:529–535
Kämpfer P, Lodders N, Grün-Wollny I, Martin K, Busse HJ (2012) Nocardia grenadensis sp. nov., isolated from sand of the Caribbean Sea. Int J Syst Evol Microbiol 62:693–697
Kathiresan K, Bingham BL (2001) Biology of mangroves and mangrove ecosystems. Adv Mar Biol 40:81–251
Kavitha A, Prabhakar P, Vijayalakshmi M, Venkateswarlu Y (2009) Production of bioactive metabolites by Nocardia levis MK-VL_113. Lett Appl Microbiol 49:484–490
Keller S, Nicholson G, Drahl C, Sorensen E, Fiedler HP, Süssmuth RD (2007a) Abyssomicins G and H and atrop-abyssomicin C from the marine Verrucosispora strain AB-18-032. J Antibiot (Tokyo) 60:391–394
Keller S, Schadt HS, Ortel I, Süssmuth RD (2007b) Action of atrop-abyssomicin C as an inhibitor of 4-amino-4-deoxychorismate synthase PabB. Angew Chem Int Ed Engl 46:8284–8286
Khamna S, Yokota A, Lumyong S (2009) Actinomycetes isolated from medicinal plant rhizosphere soils: diversity and screening of antifungal compounds, indole-3-acetic acid and siderophore production. World J Microbiol Biotechnol 25:649–655
Khan ST, Harayama S, Tamura T, Ando K, Takagi M, Kazuo SY (2009) Paraoerskovia marina gen. nov., sp. nov., an Actinobacterium isolated from marine sediment. Int J Syst Evol Microbiol 59:2094–2098
Kim CJ, Lee KH, Shimazu A, Kwon OS, Park DJ (1995) Isolation of rare Actinomycetes in various types of soil. Korean J Appl Microbiol Biotechnol 23:36–42
Kim HM, Choi DH, Hwang CY, Cho BC (2008) Nocardioides salarius sp. nov., isolated from seawater enriched with zooplankton. Int J Syst Evol Microbiol 58:2056–2064
Kim SH, Yang HO, Sohn YC, Kwon HC (2010) Aeromicrobium halocynthiae sp. nov., a taurocholic acid-producing bacterium isolated from the marine ascidian Halocynthia roretzi. Int J Syst Evol Microbiol 60:2793–2798
Knight V, Sanglier JJ, DiTullio D, Braccili S, Bonner P, Waters J, Hughes D, Zhang L (2003) Diversifying microbial natural products for drug discovery. Appl Microbiol Biotechnol 62:446–458
Kobayashi M, Natsume T, Tamaoki S, Watanabe J, Asano H, Mikami T, Miyasaka K, Miyazaki K, Gondo M, Sakakibara K, Tsukagoshi S (1997) Antitumor activity of TZT-1027, a novel dolastatin 10 derivative. Jpn J Cancer Res 88:316–327
Kokare CR, Mahadik KR, Kadam SS (2004) Isolation of bioactive marine Actinomycetes from sediments isolated from Goa and Maharashtra coastlines (west coast of India). Indian J Mar Sci 33:248–256
Kumar V, Bharti A, Gupta VK, Gusain O, Bisht GS (2012) Actinomycetes from solitary wasp mud nest and swallow bird mud nest: isolation and screening for their antibacterial activity. World J Microbiol Biotechnol 28:871–880
Kurahashi M, Fukunaga Y, Sakiyama Y, Harayama S, Yokota A (2009) Iamia majanohamensis gen. nov., sp. nov., an Actinobacterium isolated from sea cucumber Holothuria edulis, and proposal of Iamiaceae fam. nov. Int J Syst Evol Microbiol 59:869–873
Kurahashi M, Fukunaga Y, Sakiyama Y, Harayama S, Yokota A (2010) Euzebya tangerina gen. nov., sp. nov., a deeply branching marine Actinobacterium isolated from the sea cucumber Holothuria edulis, and proposal of Euzebyaceae fam. nov., Euzebyales ord. nov. and Nitriliruptoridae subclassis nov. Int J Syst Evol Microbiol 60:2314–2319
Kurosawa K, Ghiviriga I, Sambandan TG, Lessard PA, Barbara JE, Rha C, Sinskey AJ (2008) Rhodostreptomycins, antibiotics biosynthesized following horizontal gene transfer from Streptomyces padanus to Rhodococcus fascians. J Am Chem Soc 130:1126–1127
Kurtböke Dİ (2012) Biodiscovery from rare Actinomycetes: an eco-taxonomical perspective. Appl Microbiol Biotechnol 93:1843–1852
Kwon HC, Kauffman CA, Jensen PR, Fenical W (2009) Marinisporolides, polyene-polyol macrolides from a marine Actinomycetes of the new genus Marinispora. J Org Chem 74:675–684
Lam KS (2006) Discovery of novel metabolites from marine Actinomycetes. Curr Opin Microbiol 9:245–251
Lancini G, Lorenzetti R (1993) Biotechnology of antibiotics and other bioactive microbial metabolites. Plenum, New York, pp 49–57
Laureti L, Song L, Huang S, Corre C, Leblond P, Challis GL, Aigle B (2011) Identification of a bioactive 51-membered macrolide complex by activation of a silent polyketide synthase in Streptomyces ambofaciens. Proc Natl Acad Sci U S A 108:6258–6263
Lazzarini A, Cavaletti L, Toppo G, Marinelli F (2000) Rare genera of Actinomycetes as potential producers of new antibiotics. Antonie Van Leeuwenhoek 78:399–405
Lee SD (2006a) Actinocorallia cavernae sp. nov., isolated from a natural cave in Jeju, Korea. Int J Syst Evol Microbiol 56:1085–1088
Lee SD (2006b) Amycolatopsis jejuensis sp. nov. and Amycolatopsis halotolerans sp. nov., novel Actinomycetes isolated from a natural cave. Int J Syst Evol Microbiol 56:549–553
Lee SD (2006c) Nocardia jejuensis sp. nov., a novel Actinomycetes isolated from a natural cave on Jeju Island, Republic of Korea. Int J Syst Evol Microbiol 56:559–562
Lee SD (2007a) Labedella gwakjiensis gen. nov., sp. nov., a novel Actinomycete of the family Microbacteriaceae. Int J Syst Evol Microbiol 57:2498–2502
Lee SD (2007b) Marmoricola aequoreus sp. nov., a novel Actinobacterium isolated from marine sediment. Int J Syst Evol Microbiol 57:1391–1395
Lee SD (2007c) Nocardioides furvisabuli sp. nov., isolated from black sand. Int J Syst Evol Microbiol 57:35–39
Lee SD (2008a) Agrococcus jejuensis sp. nov., isolated from dried seaweed. Int J Syst Evol Microbiol 58:2297–2300
Lee SD (2008b) Brevibacterium marinum sp. nov., isolated from seawater. Int J Syst Evol Microbiol 58:500–504
Lee SD, Kim SJ (2007) Aeromicrobium tamlense sp. nov., isolated from dried seaweed. Int J Syst Evol Microbiol 57:337–341
Lee DW, Lee SD (2008a) Tessaracoccus flavescens sp. nov., isolated from marine sediment. Int J Syst Evol Microbiol 58:785–789
Lee DW, Lee SD (2008b) Aeromicrobium ponti sp. nov., isolated from seawater. Int J Syst Evol Microbiol 58:987–991
Lee DW, Lee SD (2008c) Ponticoccus gilvus gen. nov., sp. nov., a novel member of the family Propionibacteriaceae from seawater. J Microbiol 46:508–512
Lee DW, Lee SD (2010) Koreibacter algae gen. nov., sp. nov., isolated from seaweed. Int J Syst Evol Microbiol 60:1510–1515
Lee SD, Kang SO, Hah YC (2000a) Catellatospora koreensis sp. nov., a novel Actinomycete isolated from goldmine cave. Int J Syst Evol Microbiol 50:1103–1111
Lee SD, Kim ES, Roe JH, Kim J, Kang SO, Hah C (2000b) Saccharothrix violacea sp. nov., isolated from a gold mine cave, and Saccharothrix albidocapillata comb. nov. Int J Syst Evol Microbiol 50:1315–1323
Lee SD, Kim ES, Min KL, Lee WY, Kang SO, Hah YC (2001) Pseudonocardia kongjuensis sp. nov., isolated from a gold mine cave. Int J Syst Evol Microbiol 51:1505–1510
Lee DW, Hyun CG, Lee SD (2007) Nocardioides marinisabuli sp. nov., a novel Actinobacterium isolated from beach sand. Int J Syst Evol Microbiol 57:2960–2963
Lee DW, Lee JM, Seo JP, Schumann P, Kim SJ, Lee SD (2008a) Phycicola gilvus gen. nov., sp. nov., an Actinobacterium isolated from living seaweed. Int J Syst Evol Microbiol 58:1318–1323
Lee JE, Seo JP, Lee DW, Ko YH, Lee SD (2008b) Terrabacter lapilli sp. nov., an Actinomycete isolated from stone. Int J Syst Evol Microbiol 58:1084–1088
Lee SD, Lee DW, Kim JS (2008c) Nocardioides hwasunensis sp. nov. Int J Syst Evol Microbiol 58:278–281
Lewis K, Epstein S, D’Onofrio A, Ling LL (2010) Uncultured microorganisms as a source of secondary metabolites. J Antibiot (Tokyo) 63:468–476
Li IV, Terekhova LP, Alferova IV, Galatenko OA, Gapochka MG (2003) The application of succession analysis in combination with extremely high-frequency irradiation to the selective isolation of Actinomycetes from soil. Mikrobiologiia 72:131–135
Li HR, Yu Y, Luo W, Zeng YX (2010) Marisediminicola antarctica gen. nov., sp. nov., an Actinobacterium isolated from the Antarctic. Int J Syst Evol Microbiol 60:2535–2539
Li S, Tian X, Niu S, Zhang W, Chen Y, Zhang H, Yang X, Zhang W, Li W, Zhang S, Ju J, Zhang C (2011) Pseudonocardians A-C, new diazaanthraquinone derivatives from a deap-sea Actinomycete Pseudonocardia sp. SCSIO 01299. Mar Drugs 9:1428–1439
Li J, Yang J, Zhu WY, He J, Tian XP, Xie Q, Zhang S, Li WJ (2012a) Nocardiopsis coralliicola sp. nov., isolated from the gorgonian coral, Menella praelonga. Int J Syst Evol Microbiol 62:1653–1658
Li J, Zhao GZ, Long LJ, Wang FZ, Tian XP, Zhang S, Li WJ (2012b) Rhodococcus nanhaiensis sp. nov., an Actinobacterium isolated from marine sediment. Int J Syst Evol Microbiol 62:2517–2521
Liao ZL, Tang SK, Guo L, Zhang YQ, Tian XP, Jiang CL, Xu LH, Li WJ (2009) Verrucosispora lutea sp. nov., isolated from a mangrove sediment sample. Int J Syst Evol Microbiol 59:2269–2273
Liras P, Demain AL (2009) Enzymology of beta-lactam compounds with cephem structure produced by Actinomycete. Methods Enzymol 458:401–429
Little AEF, Murakami T, Mueller UG, Currie CR (2006) Defending against parasites: fungus-growing ants combine specialized behaviours and microbial symbionts to protect their fungus gardens. Biol Lett 2:12–16
Liu Z, Li Y, Zheng LQ, Huang YJ, Li WJ (2010) Saccharomonospora marina sp. nov., isolated from an ocean sediment of the East China Sea. Int J Syst Evol Microbiol 60:1854–1857
Long H, Xiang W, Zhuang T, Lin P (2005) Microorganism resource of mangrove ecosystems. Chinese J Ecol 24:696–702
Macherla VR, Liu J, Sunga M, White DJ, Grodberg J, Teisan S, Lam KS, Potts BC (2007) Lipoxazolidinones A, B, and C: antibacterial 4-oxazolidinones from a marine Actinomycete isolated from a Guam marine sediment. J Nat Prod 70:1454–1457
Maldonado LA, Fenical W, Jensen PR, Kauffman CA, Mincer TJ, Ward AC, Bull AT, Goodfellow M (2005) Salinispora arenicola gen. nov., sp. nov. and Salinispora tropica sp. nov., obligate marine Actinomycetes belonging to the family Micromonosporaceae. Int J Syst Evol Microbiol 55:1759–1766
Maloney KN, MacMillan JB, Kauffman CA, Jensen PR, DiPasquale AG, Rheingold AL, Fenical W (2009) Lodopyridone, a structurally unprecedented alkaloid from a marine Actinomycete. Org Lett 11:5422–5424
Mangamuri UK, Muvva V, Poda S, Kamma S (2012) Isolation, identification and molecular characterization of rare Actinomycetes from mangrove ecosystem of Nizampatnam. Malaysian J Microbiol 8:83–91
Matsumoto A, Takahashi Y, Mochizuki M, Seino A, Iwai Y, Omura S (1998) Characterization of Actinomycetes isolated from fallen leaves. Actinomycetologica 12:46–48
Matsumoto A, Kasai H, Matsuo Y, Omura S, Shizuri Y, Takahashi Y (2009) Ilumatobacter fluminis gen. nov., sp. nov., a novel Actinobacterium isolated from the sediment of an estuary. J Gen Appl Microbiol 55:201–205
Matsumoto A, Nakai K, Morisaki K, Omura S, Takahashi Y (2010) Demequina salsinemoris sp. nov., isolated on agar media supplemented with ascorbic acid or rutin. Int J Syst Evol Microbiol 60:1206–1209
Mayer AM, Glaser KB, Cuevas C, Jacobs RS, Kem W, Little RD, McIntosh JM, Newman DJ, Potts BC, Shuster DE (2010) The odyssey of marine pharmaceuticals: a current pipeline perspective. Trends Pharmacol Sci 31:255–265
McArthur KA, Mitchell SS, Tsueng G, Rheingold A, White DJ, Grodberg J, Lam KS, Potts BC (2008) Lynamicins A-E, chlorinated bisindole pyrrole antibiotics from a novel marine Actinomycete. J Nat Prod 71:1732–1737
McInerney MJ, Struchtemeyer CG, Sieber J, Mouttaki H, Stams AJ, Schink B, Rohlin L, Gunsalus RP (2008) Physiology, ecology, phylogeny, and genomics of microorganisms capable of syntrophic metabolism. Ann NY Acad Sci 1125:58–72
Medema MH, Trefzer A, Kovalchuk A, van den Berg M, Müller U, Heijne W, Wu L, Alam MT, Ronning CM, Nierman WC, Bovenberg RA, Breitling R, Takano E (2010) The sequence of a 1.8-mb bacterial linear plasmid reveals a rich evolutionary reservoir of secondary metabolic pathways. Genome Biol Evol 2:212–224
Mehbub MF, Amin AKMR (2012) Isolation and identification of Actinobacteria from two south Australian marine sponges Aplysilla rosea and Aplysina sp. Bangladesh Res Publ J 7:345–354
Meklat A, Sabaou N, Zitouni A, Mathieu F, Lebrihi A (2011) Isolation, taxonomy, and antagonistic properties of halophilic Actinomycetes in Saharan soils of Algeria. Appl Environ Microbiol 77:6710–6714
Miguélez EM, Martín C, Hardisson C, Manzanal MB (1993) Synchronous germination of Streptomyces antibioticus spores: tool for the analysis of hyphal growth in liquid cultures. FEMS Microbiol Lett 109:123–129
Mincer TJ, Jensen PR, Kauffman CA, Fenical W (2002) Widespread and persistent populations of a major new marine Actinomycete taxon in ocean sediments. Appl Environ Microbiol 68:5005–5011
Mita AC, Hammond LA, Bonate PL, Weiss G, McCreery H, Syed S, Garrison M, Chu QS, DeBono JS, Jones CB, Weitman S, Rowinsky EK (2006) Phase I and pharmacokinetic study of tasidotin hydrochloride (ILX651), a third-generation dolastatin-15 analogue, administered weekly for 3 weeks every 28 days in patients with advanced solid tumors. Clin Cancer Res 12:5207–5215
Mueller UG, Dash D, Rabeling C, Rodrigues A (2008) Coevolution between attine ants and Actinomycete bacteria: a reevaluation. Evolution 62:2894–2912
Murakami R, Fujita Y, Kizuka M, Kagawa T, Muramatsu Y, Miyakoshi S, Takatsu T, Inukai M (2007) A-102395, a new inhibitor of bacterial Translocase I, produced by Amycolatopsis sp. SANK 60206. J Antibiot 60:690–695
Murphy BT, Narender T, Kauffman CA, Woolery M, Jensen PR, Fenical W (2010) Saliniquinones A-F, new members of the highly cytotoxic anthraquinone-c-pyrones from the marine Actinomycete Salinispora arenicola. Aust J Chem 63:929–934
Mutka SC, Carney JR, Liu Y, Kennedy J (2006) Heterologous production of epothilone C and D in Escherichia coli. Biochemistry 45:1321–1330
Naikpatil SV, Rathod JL (2011) Selective isolation and antimicrobial activity of rare Actinomycetes from mangrove sediment of Karwar. J Ecobiotechnol 3:48–53
Nakaew N, Pathom-aree W, Lumyong S (2009) Generic diversity of rare Actinomycetes from Thai cave soils and their possible use as new bioactive compounds. Actinomycetologica 23:21–26
Nam SJ, Gaudencio SP, Kauffman CA, Jensen PR, Kondratyuk TP, Marler LE, Pezzuto JM, Fenical W (2010) Fijiolides A and B, inhibitors of TNF-alpha-induced NFkappaB activation, from a marine-derived sediment bacterium of the genus Nocardiopsis. J Nat Prod 73:1080–1086
Natsume M, Yasui K, Marumo S (1989) Calcium ion regulates aerial mycelium formation in Actinomycetes. J Antibiot 42:440–447
Negri A, Marco E, García-Hernández V, Domingo A, Llamas-Saiz AL, Porto-Sandá S, Riguera R, Laine W, David-Cordonnier MH, Bailly C, García-Fernández LF, Vaquero JJ, Gago F (2007) Antitumor activity, X-ray crystal structure, and DNA binding properties of thiocoraline A, a natural bisintercalating thiodepsipeptide. J Med Chem 50:3322–3333
Newman DJ, Cragg GM (2007) Natural products as sources of new drugs over the last 25 years. J Nat Prod 70:461–477
Newman DJ, Cragg GM, Snader KM (2003) Natural products as sources of new drugs over the period 1981–2002. J Nat Prod 66:1022–1037
Nichols D, Lewis K, Orjala J, Mo S, Ortenberg R, O’Connor P, Zhao C, Vouros P, Kaeberlein T, Epstein SS (2008) Short peptide induces an “uncultivable” microorganism to grow in vitro. Appl Environ Microbiol 74:4889–4897
Niyomvong N, Pathom-aree W, Thamchaipenet A, Duangmal K (2012) Actinomycetes from tropical limestone Caves. Chiang Mai J Sci 39:373–388
Nolan RD, Cross T (1988) Isolation and screening of Actinomycetes. In: Goodfellow M, Williams ST, Mordarski M (eds) Actinomycetes in biotechnology. Academic, London, pp 2–8
Nonomura H (1988) Isolation, taxonomy and ecology of soil Actinomycetes. Actinomycetologia 3:45–54
Nonomura H, Hayakawa M (1988) New methods for the selective isolation of soil Actinomycetes. In: Okami Y, Beppu T, Ogawara H (eds) Biology of Actinomycetes. Japan Scientific Societies, Tokyo, pp 288–293
Oh DC, Gontang EA, Kauffman CA, Jensen PR, Fenical W (2008) Salinipyrones and pacificanones, mixed-precursor polyketides from the marine Actinomycete Salinispora pacifica. J Nat Prod 71:570–575
Oh DC, Poulsen M, Currie CR, Clardy J (2009) Dentigerumycin: a bacterial mediator of an ant–fungus symbiosis. Nat Chem Biol 5:391–393
Ohnishi Y, Ishikawa J, Hara H, Suzuki H, Ikenoya M, Ikeda H, Yamashita A, Hattori M, Horinouchi S (2008) Genome sequence of the streptomycin-producing microorganism Streptomyces griseus IFO 13350. J Bacteriol 190:4050–4060
Okami B, Hotta AK (1988) Search and discovery of new antibiotics. In: Goodfellow M, Williams ST, Mordarski M (eds) Actinomycetes in biotechnology. Pergamon, Oxford, pp 33–67
Okoro CK, Brown R, Jones AL, Andrews BA, Asenjo JA, Goodfellow M, Bull AT (2009) Diversity of culturable Actinomycetes in hyper-arid soils of the Atacama Desert, Chile. Antonie Van Leeuwenhoek 95:121–133
Olson JB, Harmody DK, Bej AK, McCarthy PJ (2007) Tsukamurella spongiae sp. nov., a novel Actinomycete isolated from a deep-water marine sponge. Int J Syst Evol Microbiol 57:1478–1481
Otoguro M, Hayakawa M, Yamazaki T, Iimura Y (2001) An integrated method for the enrichment and selective isolation of Actiniokineospora spp. in soil and plant litter. J Appl Microbiol 91:118–130
Palomo S, González I, de la Cruz M, Martín J, Tormo JR, Anderson M, Hill RT, Vicente F, Reyes F, Genilloud O (2013) Sponge-derived Kocuria and Micrococcus spp. as sources of the new thiazolyl peptide antibiotic kocurin. Mar Drugs 11:1071–7086
Park SC, Baik KS, Kim MS, Chun J, Seong CN (2008) Nocardioides dokdonensis sp. nov., an Actinomycete isolated from sand sediment. Int J Syst Evol Microbiol 58:2619–2623
Pathom-Aree W, Stach JE, Ward AC, Horikoshi K, Bull AT, Goodfellow M (2006) Diversity of Actinomycetes isolated from Challenger Deep sediment (10,898 m) from the Mariana Trench. Extremophiles 10:181–189
Peiru S, Menzella HG, Rodriguez E, Carney J, Gramajo H (2005) Production of the potent antibacterial polyketide erythromycin C in Escherichia coli. Appl Environ Microbiol 71:2539–2547
Pettit GR, Tan R, Pettit RK, Smith TH, Feng S, Doubek DL, Richert L, Hamblin J, Weber C, Chapuis JC (2007) Antineoplastic agents 560. Isolation and structure of kitastatin 1 from an Alaskan Kitasatospora sp. J Nat Prod 70:1069–1072
Pfefferle C, Theobald U, Gürtler H, Fiedler HP (2000) Improved secondary metabolite production in the genus Streptosporangium by optimization of the fermentation conditions. J Biotechnol 80:135–142
Piel J (2009) Metabolites from symbiotic bacteria. Nat Prod Rep 26:338–362
Pimentel-Elardo SM, Tiro LP, Grozdanov L, Hentschel U (2008) Saccharopolyspora cebuensis sp. nov., a novel Actinomycete isolated from a Philippine sponge (Porifera). Int J Syst Evol Microbiol 58:628–632
Pindi PK, Manorama R, Begum Z, Shivaji S (2010) Arthrobacter antarcticus sp. nov., isolated from an Antarctic marine sediment. Int J Syst Evol Microbiol 60:2263–2266
Proksch P, Muller WEG (2006) Frontiers in marine biotechnology. Routledge, Norfolk, pp 225–288
Prudhomme J, McDaniel E, Ponts N, Bertani S, Fenical W, Jensen P, Roch KL (2008) Marine Actinomycetes: a new source of compounds against the human malaria parasite. PLoS One 3:e2335
Qin S, Li J, Chen HH, Zhao GZ, Zhu WY, Jiang CL, Xu LH, Li WJ (2009) Isolation, diversity, and antimicrobial activity of rare Actinobacteria from medicinal plants of tropical rain forests in Xishuangbanna, China. Appl Environ Microbiol 75:6176–6186
Qiu D, Ruan J, Huang Y (2008) Selective isolation and rapid identification of members of the genus Micromonospora. Appl Environ Microbiol 74:5593–5597
Qui J (2010) Nature News. doi:10.1038/news.2010.190. http://www.nature.com/news/2010/100418/full/news.2010.190.html. Accessed 18 April 2010
Raju R, Piggott AM, Conte M, Tnimov Z, Alexandrov K, Capon RJ (2010) Nocardiopsins: new FKBP12-binding macrolide polyketides from an Australian marine-derived Actinomycete, Nocardiopsis sp. Chemistry 16:3194–3200
Renu S, Monisha K, Rup L (2008) Bioactive compounds from marine Actinomycetes. Indian J Microbiol 48:410–431
Riedlinger J, Reicke A, Zähner H, Krismer B, Bull AT, Maldonado LA, Ward AC, Goodfellow M, Bister B, Bischoff D, Süssmuth RD, Fiedler HP (2004) Abyssomicins, inhibitors of the para-aminobenzoic acid pathway produced by the marine Verrucosispora strain AB-18-032. J Antibiot (Tokyo) 57:271–279
Roh H, Uguru GC, Ko HJ, Kim S, Kim BY, Goodfellow M, Bull AT, Kim KH, Bibb MJ, Choi IG, Stach JEM (2011) Genome sequence of the abyssomicin- and proximicin producing marine Actinomycete Verrucosispora maris AB-18-032. J Bacteriol 193:3391–3392
Romero F, Espliego F, Pérez Baz J, García De Quesada T, Grávalos D, de la Calle F, Fernández-Puentes JL (1997) Thiocoraline, a new depsipeptide with antitumor activity produced by a marine Micromonospora. I. Taxonomy, fermentation, isolation, and biological activities. J Antibiot (Tokyo) 50:734–737
Sánchez S, Chávez A, Forero A, García-Huante Y, Romero A, Sánchez M, Rocha D, Sánchez B, Avalos M, Guzmán-Trampe S, Rodríguez-Sanoja R, Langley E, Ruiz B (2010) Carbon source regulation of antibiotic production. J Antibiot (Tokyo) 63:442–459
Schabereiter-Gurtner C, Saiz-Jimenez C, Pinar G, Lubitz W, Rolleke S (2002) Altamira Cave Paleolithic paintings harbour partly unknown bacterial communities. FEMS Microbiol Lett 211:7–11
Schneemann I, Nagel K, Kajahn I, Labes A, Wiese J, Imhoff JF (2010) Comprehensive investigation of marine Actinobacteria associated with the sponge Halichondria panicea. Appl Environ Microbiol 76:3702–3714
Schneider K, Keller S, Wolter FE, Roglin L, Beil W, Seitz O, Nicholson G, Bruntner C, Riedlinger J, Fiedler HP, Sussmuth RD (2008) Proximicins A, B and C antitumor furan analogues of netropsin from the marine Actinomycetes Verrucosispora induced up regulation of p53 and the cyclin kinase inhibitor p21. Angew Chem Int Ed 47:3258–3261
Schultz M, Tsaklakidis C (1997) Nach Chem Tech Lab 45:159–165
Sen R, Ishak HD, Estrada D, Dowd SE, Hong E, Mueller UG (2009) Generalized antifungal activity and 454-screening of Pseudonocardia and Amycolatopsis bacteria in nests of fungus-growing ants. Proc Natl Acad Sci U S A 106:17805–17810
Seo YB, Kim DE, Kim GD, Kim HW, Nam SW, Kim YT, Lee JH (2009) Kocuria gwangalliensis sp. nov., an Actinobacterium isolated from seawater. Int J Syst Evol Microbiol 59:2769–2772
Seong CN, Choi JH, Baik KS (2001) An improved selective isolation of rare Actinomycetes from forest soil. J Microbiol 39:17–23
Shirai M, Okuda M, Motohashi K, Inoto M, Furihata K, Matsuo Y, Shizuri Y, Seto H (2010) Terpenoids produced by Actinomycetes: isolation, structural elucidation and biosynthesis of new diterpenes: gifhornenolones A and B from Verrucosispora gifhornensis YM28-088. J Antibiot 63:245–250
Simmons L, Kaufmann K, Garcia R, Schwär G, Huch V, Müller R (2011) Bendigoles D-F, bioactive sterols from the marine sponge-derived Actinomadura sp. SBMs009. Bioorg Med Chem 19:6570–6575
Sogin ML, Morrison HG, Huber JA, Mark WD, Huse SM, Neal PR, Arrieta JM, Herndl GJ (2006) Microbial diversity in the deep sea and the underexplored “rare biosphere”. Proc Natl Acad Sci U S A 103:12115–12120
Solano G, Rojas-Jiménez K, Jaspars M, Tamayo-Castillo G (2009) Study of the diversity of culturable Actinomycetes in the North Pacific and Caribbean coasts of Costa Rica. Antonie Van Leeuwenhoek 96:71–78
Song JY, Jeong H, Yu DS, Fischbach MA, Park HS, Kim JJ, Seo JS, Jensen SE, Oh TK, Lee KJ, Kim JF (2010) Draft genome sequence of Streptomyces clavuligerus NRRL 3585, a producer of diverse secondary metabolites. J Bacteriol 192:6317–6318
Songsumanus A, Tanasupawat S, Igarashi Y, Kudo T (2013) Micromonospora maritima sp. nov., isolated from mangrove soil. Int J Syst Evol Microbiol 63:554–559
Sousa TS, Jimenez PC, Ferreira EG, Silveira ER, Braz-Filho R, Pessoa OD, Costa-Lotufo LV (2012) Anthracyclinones from Micromonospora sp. J Nat Prod 75:489–493
Stach J (2010) Antimicrobials: treasures from the oceans. Microbiol Today 105:1–3
Stach JE, Bull AT (2005) Estimating and comparing the diversity of marine Actinobacteria. Antonie Van Leeuwenhoek 87:3–9
Stach JEM, Maldonado LA, Ward AC, Goodfellow M, Bull AT (2003) New primers for the class Actinobacteria: application to marine and terrestrial environments. Environ Microbiol 5:828–841
Stewart EJ (2012) Growing unculturable bacteria. J Bacteriol 194:4151–4160
Subramani R, Aalbersberg W (2012) Marine Actinomycetes: an ongoing source of novel bioactive metabolites. Microbiol Res 167:571–580
Subramani R, Narayanasamy M (2009) Screening of marine Actinomycetes isolated from the Bay of Bengal, India for antimicrobial activity and industrial enzymes. World J Microbiol Biotechnol 25:2103–2111
Supong K, Suriyachadkun C, Tanasupawat S, Suwanborirux K, Pittayakhajonwut P, Kudo T, Thawai C (2013) Micromonospora sediminicola sp. nov., isolated from marine sediment. Int J Syst Evol Microbiol 63:570–575
Taechowisan T, Peberdy JF, Lumyong S (2003) Isolation of endophytic Actinomycetes from selected plants and their antifungal activity. World J Microbiol Biotechnol 19:381–385
Takahashi Y, Matsumoto A, Seino A, Iwai Y, Omura S (1996) Rare Actinomycetes isolated from desert soils. Actinomycetologica 10:91–97
Takizawa M, Colwell RR, Hill RT (1997) Isolation and diversity of Actinomycetes in the Chesapeake Bay. Appl Environ Microbiol 59:997–1002
Tan GY, Ward AC, Goodfellow M (2006) Exploration of Amycolatopsis diversity in soil using genus-specific primers and novel selective media. Syst Appl Microbiol 29:557–569
Tanasupawat S, Jongrungruangchok S, Kudo T (2010) Micromonospora marina sp. nov., isolated from sea sand. Int J Syst Evol Microbiol 60:648–652
Terahara T, Kobayashi T, Imada C (2013) An effective method based on wet-heat treatment for the selective isolation of Micromonospora from estuarine sediments. World J Microbiol Biotechnol 29:1677–1684
Thawai C, Tanasupawat S, Kudo T (2008) Micromonospora pattaloongensis sp. nov., isolated from a Thai mangrove forest. Int J Syst Evol Microbiol 58:1516–1521
Tian XP, Tang SK, Dong JD, Zhang YQ, Xu LH, Zhang S, Li WJ (2009a) Marinactinospora thermotolerans gen. nov., sp. nov., a marine Actinomycete isolated from a sediment in the northern South China Sea. Int J Syst Evol Microbiol 59:948–952
Tian XP, Zhi XY, Qiu YQ, Zhang YQ, Tang SK, Xu LH, Zhang S, Li WJ (2009b) Sciscionella marina gen. nov., sp. nov., a marine Actinomycete isolated from a sediment in the northern South China Sea. Int J Syst Evol Microbiol 59:222–228
Tian XP, Xu Y, Zhang J, Li J, Chen Z, Kim CJ, Li WJ, Zhang CS, Zhang S (2012) Streptomyces oceani sp. nov., a new obligate marine Actinomycete isolated from a deep-sea sample of seep authigenic carbonate nodule in South China Sea. Antonie Van Leeuwenhoek 102:335–343
Tian XP, Long LJ, Li SM, Zhang J, Xu Y, He J, Li J, Wang FZ, Li WJ, Zhang CS, Zhang S (2013) Pseudonocardia antitumoralis sp. nov., a deoxynyboquinone-producing Actinomycete isolated from a deep-sea sediment. Int J Syst Evol Microbiol 63:893–899
Tishkov S (2001) Bioactive products from Actinomycetes—antibiotics, enzyme inhibitors, immunomodulators. In: Moncheva P, Tishkov S, Chipeva V (eds) Innovative aspects in biotechnology of prokaryotes. National bank for industrial microorganisms and cell cultures, Sofia, pp 111–138
Tiwari K, Gupta RK (2012a) Diversity and isolation of rare Actinomycetes: an overview. Crit Rev Microbiol 39:256–294
Tiwari K, Gupta RK (2012b) Rare Actinomycetes: a potential storehouse for novel antibiotics. Crit Rev Biotechnol 32:108–132
Traiwan J, Park MK, Kim W (2011) Serinicoccus chungangensis sp. nov., isolated from tidal flat sediment, and emended description of the genus Serinicoccus. Int J Syst Evol Microbiol 61:1299–1303
Tsao PH, Leben C, Keitt GW (1960) An enrichment method for isolating Actinomycetes that produce diffusible antifungal antibiotics. Phytopathology 50:88–89
Tseng M, Liao HC, Chiang WP, Yuan GF (2011) Isoptericola chiayiensis sp. nov., isolated from mangrove soil. Int J Syst Evol Microbiol 61:1667–1670
Ue H, Matsuo Y, Kasai H, Yokota A (2011a) Demequina globuliformis sp. nov., Demequina oxidasica sp. nov. and Demequina aurantiaca sp. nov., Actinobacteria isolated from marine environments, and proposal of Demequinaceae fam. nov. Int J Syst Evol Microbiol 61:1322–1329
Ue H, Matsuo Y, Kasai H, Yokota A (2011b) Miniimonas arenae gen. nov., sp. nov., an Actinobacterium isolated from sea sand. Int J Syst Evol Microbiol 61:123–127
Vartoukian SR, Palmer RM, Wade WG (2010) Strategies for culture of ‘unculturable’ bacteria. FEMS Microbiol Lett 309:1–7
Walsh C (2003) Where will new antibiotics come from? Nat Rev Microbiol 1:65–70
Wang Y, Mueller UG, Clardy J (1999) Antifungal diketopiperazines from symbiotic fungus of fungus growing ant Cyphomyrmex minutus. J Chem Ecol 25:935–941
Wang F, Gai Y, Chen M, Xiao X (2009) Arthrobacter psychrochitiniphilus sp. nov., a psychrotrophic bacterium isolated from Antarctica. Int J Syst Evol Microbiol 59:2759–2762
Wang J, Li Y, Bian J, Tang SK, Ren B, Chen M, Li WJ, Zhang LX (2010) Prauserella marina sp. nov., isolated from ocean sediment of the South China Sea. Int J Syst Evol Microbiol 60:985–989
Wang C, Xu XX, Qu Z, Wang HL, Lin HP, Xie QY, Ruan JS, Hong K (2011a) Micromonospora rhizosphaerae sp. nov., isolated from mangrove rhizosphere soil. Int J Syst Evol Microbiol 61:320–324
Wang F, Xu XX, Qu Z, Wang C, Lin HP, Xie QY, Ruan JS, Sun M, Hong K (2011b) Nonomuraea wenchangensis sp. nov., isolated from mangrove rhizosphere soil. Int J Syst Evol Microbiol 61:1304–1308
Wang DS, Xue QH, Zhu WJ, Zhao J, Duan JL, Shen GH (2013a) Microwave irradiation is a useful tool for improving isolation of Actinomycetes from soil. Microbiology 82:102–110
Wang Q, Song F, Xiao X, Huang P, Li L, Monte A, Abdel-Mageed WM, Wang J, Guo H, He W, Xie F, Dai H, Liu M, Chen C, Xu H, Liu M, Piggott AM, Liu X, Capon RJ, Zhang L (2013b) Abyssomicins from the South China Sea deep-sea sediment Verrucosispora sp.: natural thioether Michael addition adducts as antitubercular prodrugs. Angew Chem Int Ed Engl 52:1231–1234
Webster NS, Hill RT (2001) The culturable microbial community of the Great Barrier Reef sponge Rhopaloeides odorabile is dominated by an α-Proteobacterium. Mar Biol 138:843–851
Williams ST, Wellington EMH (1982) Principles and problems of selective isolation of microbes. In: Bullock JD, Nisbet LJ, Winstanley DJ (eds) Bioactive microbial products I: search and discovery. Academic, London, pp 9–26
Williams P, Winzer K, Chan WC, Camara M (2007a) Look who’s talking: communication and quorum sensing in the bacterial world. Philos Trans R Soc Lond B Biol Sci 362:1119–1134
Williams PG, Miller ED, Asolkar RN, Jensen PR, Fenical W (2007b) Arenicolides A-C, 26- membered ring macrolides from the marine Actinomycete Salinispora arenicola. J Org Chem 72:5025–5034
Wright GD, Sutherland AD (2007) New strategies for combating multidrug-resistant bacteria. Trends Mol Med 13:260–267
Wu Y, Li WJ, Tian W, Zhang LP, Xu L, Shen QR, Shen B (2010) Isoptericola jiangsuensis sp. nov., a chitin-degrading bacterium. Int J Syst Evol Microbiol 60:904–908
Wu H, Chen W, Wang G, Dai S, Zhou D, Zhao H, Guo Y, Ouyang YC, Li X (2012) Culture-dependent diversity of Actinobacteria associated with seagrass (Thalassia hemprichii). African J Microbiol Res 6:87–94
Wyche TP, Hou Y, Braun D, Cohen HC, Xiong MP, Bugni TS (2011) First natural analogs of the cytotoxic thiodepsipeptide thiocoraline A from a marine Verrucosispora sp. J Org Chem 76:6542–6547
Xi L, Zhang L, Ruan J, Huang Y (2011a) Description of Verrucosispora qiuiae sp. nov., isolated from mangrove swamp sediment, and emended description of the genus Verrucosispora. Int J Syst Evol Microbiol 62:1564–1569
Xi L, Zhang L, Ruan J, Huang Y (2011b) Nonomuraea maritima sp. nov., isolated from coastal sediment. Int J Syst Evol Microbiol 61:2740–2744
Xiao C, Huang H, Ye J, Wu X, Zhu J, Zhan B, Bao S (2011a) Ornithinibacter aureus gen. nov., sp. nov., a novel member of the family Intrasporangiaceae. Int J Syst Evol Microbiol 61:659–664
Xiao J, Luo Y, Xie S, Xu J (2011b) Serinicoccus profundi sp. nov., an Actinomycete isolated from deep-sea sediment, and emended description of the genus Serinicoccus. Int J Syst Evol Microbiol 61:16–19
Xiao J, Luo Y, Xu J, Xie S, Xu J (2011c) Modestobacter marinus sp. nov., a psychrotolerant Actinobacterium from deep-sea sediment, and emended description of the genus Modestobacter. Int J Syst Evol Microbiol 61:1710–1714
Xu LH, Jiang CL (1996) A study on diversity of aquatic Actinomycetes in lakes of the middle plateau, Yunnan, China. Appl Environ Microbiol 62:249–253
Xu LH, Li WJ, Liu ZH, Jiang CL (2007) Actinomycete taxonomy. Acedemic, Beijing, pp 202–208
Xu XX, Qu Z, Wang H, Lin HP, Wang C, Xie QY, Ruan JS, Hong K (2011) Asanoa hainanensis sp. nov., isolated from rhizosphere soil of Acrostichum speciosum in a mangrove, and emended description of the genus Asanoa. Int J Syst Evol Microbiol 61:2384–2388
Xu XX, Wang HL, Lin HP, Wang C, Qu Z, Xie QY, Ruan JS, Hong K (2012) Microbispora hainanensis sp. nov., isolated from rhizosphere soil of Excoecaria agallocha in a mangrove. Int J Syst Evol Microbiol 62:2430–2434
Xue Q, Dua CM, Wang LN, Lin YB (2010) The influence of microwave irradiation to the isolation effect of soil Actinomycetes. Chinese J Microbiol 3:19–24
Yang B, Xue QH, Chen ZQ, Zhou YQ, Zhang XL, Xu YJ, Guo ZY (2008) Effects of microwave irradiation on isolation of soil Actinomycetes. Ying Yong Sheng Tai Xue Bao 5:1091–1098
Yang XW, Zhang GY, Ying JX, Yang B, Zhou XF, Steinmetz A, Liu YH, Wang N (2013) Isolation, characterization, and bioactivity evaluation of 3-((6-methylpyrazin-2-yl)methyl)-1H-indole, a new alkaloid from a deep-sea-derived Actinomycete Serinicoccus profundi sp. nov. Mar Drugs 11:33–39
Yi H, Schumann P, Chun J (2007) Demequina aestuarii gen. nov., sp. nov., a novel Actinomycete of the suborder Micrococcineae, and reclassification of Cellulomonas fermentans Bagnara et al. 1985 as Actinotalea fermentans gen. nov., comb. nov. Int J Syst Evol Microbiol 57:151–156
Yi-lei N, Ru L, Yang X, Hong J (2009) Isolation, purification and structure determination of antifungal antibiotic FW03-1149 produced by marine Micromonospora sp. FIM03-1149. Chinese J Mar Drugs 01
Young P (1997) Major microbial diversity initiative recommended. ASM News 63:417–421
Yu L, Lai Q, Yi Z, Zhang L, Huang Y, Gu L, Tang X (2013) Microbacterium sediminis sp. nov., a psychrotolerant, thermotolerant, halotolerant and alkalitolerant Actinomycete isolated from deep-sea sediment. Int J Syst Evol Microbiol 63:25–30
Zhang L (2005) Integrated approaches for discovering novel drugs from microbial natural products. In: Zhang L, Demain AL (eds) Natural products. Drug discovery and therapeutic medicine. Humana, Totowa, pp 33–35
Zhang L, An R, Wang J, Sun N, Zhang S, Hu J, Kuai J (2005) Exploring novel bioactive compounds from marine microbes. Curr Opin Microbiol 8:276–281
Zhang C, Herath K, Jayasuriya H, Ondeyka JG, Zink DL, Occi J, Birdsall G, Venugopal J, Ushio M, Burgess B, Masurekar P, Barrett JF, Singh SB (2009) Thiazomycins, thiazolyl peptide antibiotics from Amycolatopsis fastidiosa. J Nat Prod 72:841–847
Zhang L, Xi L, Ruan J, Huang Y (2012) Micromonospora yangpuensis sp. nov., isolated from a sponge. Int J Syst Evol Microbiol 62:272–278
Zhang Q, Li S, Chen Y, Tian X, Zhang H, Zhang G, Zhu Y, Zhang S, Zhang W, Zhang C (2013) New diketopiperazine derivatives from a deep-sea-derived Nocardiopsis alba SCSIO 03039. J Antibiot (Tokyo) 66:31–36
Zhuge B, Fang HY, Yu H, Rao ZM, Shen W, Song J, Zhuge J (2008) Bioconversion of Lovastatin to a novel statin, Wuxistatin by Amycolatopsis sp. Appl Microbiol Biotechnol 79:209–216
Zobell CE (1946) Marine microbiology. Chronica Botanica Co, Waltham
Acknowledgments
This work was supported by the US National Institutes of Health’s International Cooperative Biodiversity Groups program (grant NIH ICBG U01-TW007401). The authors thank Dr. Brad Carte and Dr. Patricia Kailola for critically reading the manuscript, their helpful comments and encouraging remarks.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Subramani, R., Aalbersberg, W. Culturable rare Actinomycetes: diversity, isolation and marine natural product discovery. Appl Microbiol Biotechnol 97, 9291–9321 (2013). https://doi.org/10.1007/s00253-013-5229-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-013-5229-7