Abstract
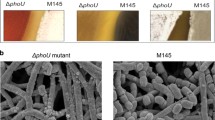
Streptomyces lividans senses and adjusts to a situation of Pi limitation via the expression of genes of the pho regulon controlled by the two-component system PhoR/PhoP. Interestingly, an in silico analysis of the proteins encoded by the six genes located in divergence of phoR/phoP revealed that the latter bear features often found in metalloproteins involved in the sensing/resistance to oxidative stress. We determined whether genes of this region were belonging to the pho regulon and whether the encoded proteins do play a role in the resistance to oxidative stress. For this purpose, a transcriptional analysis of these genes was carried out on the carbon and nitrogen rich medium R2YE and on a minimal medium (MM). On R2YE, the expression of the genes phoU to sco4225 was much higher than on MM and constant throughout growth. On this medium, the expression of phoU was totally PhoP-dependent whereas the expression of sco4227 and sco4226 was partially PhoP-dependent, taking place from the phoU promoter region. In contrast, on MM, the expression of sco4227 and sco4226 was PhoP-independent whereas that of phoU remained PhoP-dependent and showed, as phoR/phoP, a peak of expression at 48 h. sco4225, sco4224, and sco4223 were transcribed from their own promoter independently of PhoP in both media. The mutants of five out of six genes of the region (Δsco4226 mutant could not be obtained) grew poorly in the presence of exogenous oxidants, suggesting a role of the encoded proteins in the resistance to oxidative stress, especially on the rich medium R2YE.






Similar content being viewed by others
References
Alix E, Blanc-Potard AB (2009) Hydrophobic peptides: novel regulators within bacterial membrane. Mol Microbiol 72:5–11
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Apel K, Hirt H (2004) Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 55:373–399
Apel AK, Sola-Landa A, Rodriguez-Garcia A, Martin JF (2007) Phosphate control of phoA, phoC and phoD gene expression in Streptomyces coelicolor reveals significant differences in binding of PhoP to their promoter regions. Microbiology 153:3527–3537
Baek JH, Lee SY (2006) Novel gene members in the Pho regulon of Escherichia coli. FEMS Microbiol Lett 264:104–109
Baltz RH (2008) Renaissance in antibacterial discovery from actinomycetes. Curr Opin Pharmacol 8:557–563
Bentley SD, Chater KF, Cerdeno-Tarraga AM, Challis GL, Thomson NR, James KD, Harris DE, Quail MA, Kieser H, Harper D, Bateman A, Brown S, Chandra G, Chen CW, Collins M, Cronin A, Fraser A, Goble A, Hidalgo J, Hornsby T, Howarth S, Huang CH, Kieser T, Larke L, Murphy L, Oliver K, O'Neil S, Rabbinowitsch E, Rajandream MA, Rutherford K, Rutter S, Seeger K, Saunders D, Sharp S, Squares R, Squares S, Taylor K, Warren T, Wietzorrek A, Woodward J, Barrell BG, Parkhill J, Hopwood DA (2002) Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417:141–147
Blondelet-Rouault MH, Weiser J, Lebrihi A, Branny P, Pernodet JL (1997) Antibiotic resistance gene cassettes derived from the omega interposon for use in E. coli and Streptomyces. Gene 190:315–317
Bogel G, Schrempf H, de Orue O, Lucana D (2009) The heme-binding protein HbpS regulates the activity of the Streptomyces reticuli iron-sensing histidine kinase SenS in a redox-dependent manner. Amino Acids 37:681–691
Buelow DR, Raivio TL (2010) Three (and more) component regulatory systems—auxiliary regulators of bacterial histidine kinases. Mol Microbiol 75:547–566
Capdevila M, Atrian S (2011) Metallothionein protein evolution: a miniassay. J Biol Inorg Chem 16:977–989
Challis GL (2008a) Genome mining for novel natural product discovery. J Med Chem 51:2618–2628
Challis GL (2008b) Mining microbial genomes for new natural products and biosynthetic pathways. Microbiology 154:1555–1569
Church GM, Gilbert W (1984) Genomic sequencing. Proc Natl Acad Sci USA 81:1991–1995
Coyle P, Philcox JC, Carey LC, Rofe AM (2002) Metallothionein: the multipurpose protein. Cell Mol Life Sci 59:627–647
Crepin S, Chekabab SM, Le Bihan G, Bertrand N, Dozois CM, Harel J (2011) The Pho regulon and the pathogenesis of Escherichia coli. Vet Microbiol 153:82–88
den Hengst CD, Buttner MJ (2008) Redox control in actinobacteria. Biochim Biophys Acta 1780:1201–1216
Diaz M, Esteban A, Fernandez-Abalos JM, Santamaria RI (2005) The high-affinity phosphate-binding protein PstS is accumulated under high fructose concentrations and mutation of the corresponding gene affects differentiation in Streptomyces lividans. Microbiology 151:2583–2592
Doumith M, Weingarten P, Wehmeier UF, Salah-Bey K, Benhamou B, Capdevila C, Michel JM, Piepersberg W, Raynal MC (2000) Analysis of genes involved in 6-deoxyhexose biosynthesis and transfer in Saccharopolyspora erythraea. Mol Gen Genet 264:477–485
Dwyer DJ, Kohanski MA, Collins JJ (2009) Role of reactive oxygen species in antibiotic action and resistance. Curr Opin Microbiol 12:482–489
Fenton HJH (1986) Oxidation of tartaric acid in the presence of iron. J Chem Soc 65:899–910
Flardh K, Buttner MJ (2009) Streptomyces morphogenetics: dissecting differentiation in a filamentous bacterium. Nat Rev Microbiol 7:36–49
Ghorbel S, Kormanec J, Artus A, Virolle MJ (2006) Transcriptional studies and regulatory interactions between the phoR-phoP operon and the phoU, mtpA, and ppk genes of Streptomyces lividans TK24. J Bacteriol 188:677–686
Glaser L, Loewy A (1979a) Control of teichoic acid synthesis during phosphate limitation. J Bacteriol 137:327–331
Glaser L, Loewy A (1979b) Regulation of teichoic acid synthesis during phosphate limitation. J Biol Chem 254:2184–2186
Goto Y, Li B, Claesen J, Shi Y, Bibb MJ, van der Donk WA (2010) Discovery of unique lanthionine synthetases reveals new mechanistic and evolutionary insights. PLoS Biol 8:e1000339
Guillemet ML, Moreau PL (2008) Fur-dependent detoxification of organic acids by rpoS mutants during prolonged incubation under aerobic, phosphate starvation conditions. J Bacteriol 190:5567–5575
Gust B, Challis GL, Fowler K, Kieser T, Chater KF (2003) PCR-targeted Streptomyces gene replacement identifies a protein domain needed for biosynthesis of the sesquiterpene soil odor geosmin. Proc Natl Acad Sci USA 100:1541–1546
Hassett DJ, Imlay JA (2007) Bactericidal antibiotics and oxidative stress: a radical proposal. ACS Chem Biol 2:708–710
Hobbs G, Frazer CM, Gardner DCJ, Cullum JA, Oliver SG (1989) Dispersed growth of Streptomyces in liquid culture. Appl Microbiol Biotechnol 31:272–277
Hopwood DA, Kieser T, Wright HM, Bibb MJ (1983) Plasmids, recombination and chromosome mapping in Streptomyces lividans 66. J Gen Microbiol 129:2257–2269
Hsieh YJ, Wanner BL (2010) Global regulation by the seven-component Pi signaling system. Curr Opin Microbiol 13:198–203
Hulett FM (1996) The signal-transduction network for Pho regulation in Bacillus subtilis. Mol Microbiol 19:933–939
Imlay JA (2003) Pathways of oxidative damage. Annu Rev Microbiol 57:395–418
Karray F (2005) Etude de la biosynthèse de l'antibiotique spiramycine par Streptomyces ambofaciens. Université Paris sud 11, Orsay, France
Kehrer JP (2000) The Haber-Weiss reaction and mechanisms of toxicity. Toxicology 149:43–50
Kieser T, Bibb MJ, Buttner MJ, Chater KF, Hopwood DA (2000) Practical Streptomyces Genetics. In: Foundation TJI (ed), Norwich, UK, pp. 248
Kosower NS, Kosower EM (1995) Diamide: an oxidant probe for thiols. Methods Enzymol 251:123–133
Liu J, Lou Y, Yokota H, Adams PD, Kim R, Kim SH (2005) Crystal structure of a PhoU protein homologue: a new class of metalloprotein containing multinuclear iron clusters. J Biol Chem 280:15960–15966
Makino K, Amemura M, Kim SK, Nakata A, Shinagawa H (1993) Role of the sigma 70 subunit of RNA polymerase in transcriptional activation by activator protein PhoB in Escherichia coli. Genes Dev 7:149–160
Marchler-Bauer A, Anderson JB, Cherukuri PF, DeWeese-Scott C, Geer LY, Gwadz M, He S, Hurwitz DI, Jackson JD, Ke Z, Lanczycki CJ, Liebert CA, Liu C, Lu F, Marchler GH, Mullokandov M, Shoemaker BA, Simonyan V, Song JS, Thiessen PA, Yamashita RA, Yin JJ, Zhang D, Bryant SH (2005) CDD: a Conserved Domain Database for protein classification. Nucleic Acids Res 33:D192–D196
Martin JF (2004) Phosphate control of the biosynthesis of antibiotics and other secondary metabolites is mediated by the PhoR-PhoP system: an unfinished story. J Bacteriol 186:5197–5201
Marzan LW, Shimizu K (2011) Metabolic regulation of Escherichia coli and its phoB and phoR genes knockout mutants under phosphate and nitrogen limitations as well as at acidic condition. Microb Cell Fact 10:39
Moreau PL (2004) Diversion of the metabolic flux from pyruvate dehydrogenase to pyruvate oxidase decreases oxidative stress during glucose metabolism in nongrowing Escherichia coli cells incubated under aerobic, phosphate starvation conditions. J Bacteriol 186:7364–7368
Moreau PL, Gerard F, Lutz NW, Cozzone P (2001) Non-growing Escherichia coli cells starved for glucose or phosphate use different mechanisms to survive oxidative stress. Mol Microbiol 39:1048–1060
Muth G, Nussbaumer B, Wohlleben W, Pühler A (1989) A vector system with temperature-sensitive replication for gene disruption and mutational cloning in streptomycetes. Mol Gen Genet 219:341–348
Nguyen HC, Karray F, Lautru S, Gagnat J, Lebrihi A, Huynh TD, Pernodet JL (2010) Glycosylation steps during spiramycin biosynthesis in Streptomyces ambofaciens: involvement of three glycosyltransferases and their interplay with two auxiliary proteins. Antimicrob Agents Chemother 54:2830–2839
Oganesyan V, Oganesyan N, Adams PD, Jancarik J, Yokota HA, Kim R, Kim SH (2005) Crystal structure of the "PhoU-like" phosphate uptake regulator from Aquifex aeolicus. J Bacteriol 187:4238–4244
Ortiz de Orue Lucana D, Zou P, Nierhaus M, Schrempf H (2005) Identification of a novel two-component system SenS/SenR modulating the production of the catalase-peroxidase CpeB and the haem-binding protein HbpS in Streptomyces reticuli. Microbiology 151:3603–3614
Ortiz de Orue Lucana D, Bogel G, Zou P, Groves MR (2009) The oligomeric assembly of the novel haem-degrading protein HbpS is essential for interaction with its cognate two-component sensor kinase. J Mol Biol 386:1108–1122
Ortiz de Orue Lucana D, Groves MR (2009) The three-component signalling system HbpS-SenS-SenR as an example of a redox sensing pathway in bacteria. Amino Acids 37:479–486
Raynal A, Karray F, Tuphile K, Darbon-Rongere E, Pernodet JL (2006) Excisable cassettes: new tools for functional analysis of Streptomyces genomes. Appl Environ Microbiol 72:4839–4844
Redenbach M, Kieser HM, Denapaite D, Eichner A, Cullum J, Kinashi H, Hopwood DA (1996) A set of ordered cosmids and a detailed genetic and physical map for the 8 Mb Streptomyces coelicolor A3(2) chromosome. Mol Microbiol 21:77–96
Rice CD, Pollard JE, Lewis ZT, McCleary WR (2009) Employment of a promoter-swapping technique shows that PhoU modulates the activity of the PstSCAB2 ABC transporter in Escherichia coli. Appl Environ Microbiol 75:573–582
Robinson NJ, Whitehall SK, Cavet JS (2001) Microbial metallothioneins. Adv Microb Physiol 44:183–213
Rodriguez-Garcia A, Barreiro C, Santos-Beneit F, Sola-Landa A, Martin JF (2007) Genome-wide transcriptomic and proteomic analysis of the primary response to phosphate limitation in Streptomyces coelicolor M145 and in a ΔphoP mutant. Proteomics 7:2410–2429
Rodriguez-Garcia A, Sola-Landa A, Apel K, Santos-Beneit F, Martin JF (2009) Phosphate control over nitrogen metabolism in Streptomyces coelicolor: direct and indirect negative control of glnR, glnA, glnII and amtB expression by the response regulator PhoP. Nucleic Acids Res 37:3230–3242
Russwurm M, Koesling D (2004) Guanylyl cyclase: NO hits its target. Biochem Soc Symp: 51–63
Sambrook J, Russel DW 2001 Molecular Cloning: a Laboratory Manual, Third edition edn., pp. Pages.
Sanchez R, Riddle M, Woo J, Momand J (2008) Prediction of reversibly oxidized protein cysteine thiols using protein structure properties. Protein Sci 17:473–481
Santos-Beneit F, Rodriguez-Garcia A, Franco-Dominguez E, Martin JF (2008) Phosphate-dependent regulation of the low- and high-affinity transport systems in the model actinomycete Streptomyces coelicolor. Microbiology 154:2356–2370
Santos-Beneit F, Rodriguez-Garcia A, Apel AK, Martin JF (2009a) Phosphate and carbon source regulation of two PhoP-dependent glycerophosphodiester phosphodiesterase genes of Streptomyces coelicolor. Microbiology 155:1800–1811
Santos-Beneit F, Rodriguez-Garcia A, Sola-Landa A, Martin JF (2009b) Cross-talk between two global regulators in Streptomyces: PhoP and AfsR interact in the control of afsS, pstS and phoRP transcription. Mol Microbiol 72:53–68
Santos-Beneit F, Barriuso-Iglesias M, Fernandez-Martinez LT, Martinez-Castro M, Sola-Landa A, Rodriguez-Garcia A, Martin JF (2011) The RNA polymerase omega factor RpoZ is regulated by PhoP and has an important role in antibiotic biosynthesis and morphological differentiation in Streptomyces coelicolor. Appl Environ Microbiol 77:7586–7594
Sharma RSRT (1996) Preparation of electrocompetent E. coli using salt-free growth medium. Biotechniques 20:42–44
Siedenburg G, Groves MR, de Orue O, Lucana D (2012) Novel redox-sensing modules: accessory protein- and nucleic Acid-mediated signaling. Antioxid Redox Signal 16:668–677
Sola-Landa A, Moura RS, Martin JF (2003) The two-component PhoR-PhoP system controls both primary metabolism and secondary metabolite biosynthesis in Streptomyces lividans. Proc Natl Acad Sci USA 100:6133–6138
Sola-Landa A, Rodriguez-Garcia A, Franco-Dominguez E, Martin JF (2005) Binding of PhoP to promoters of phosphate-regulated genes in Streptomyces coelicolor: identification of PHO boxes. Mol Microbiol 56:1373–1385
Sola-Landa A, Rodriguez-Garcia A, Apel AK, Martin JF (2008) Target genes and structure of the direct repeats in the DNA-binding sequences of the response regulator PhoP in Streptomyces coelicolor. Nucleic Acids Res 36:1358–1368
Steed PM, Wanner BL (1993) Use of the rep technique for allele replacement to construct mutants with deletions of the pstSCAB-phoU operon: evidence of a new role for the PhoU protein in the phosphate regulon. J Bacteriol 175:6797–6809
Taschner NP, Yagil E, Spira B (2004) A differential effect of sigmaS on the expression of the PHO regulon genes of Escherichia coli. Microbiology 150:2985–2992
Thompson CJ, Ward JM, Hopwood DA (1980) DNA cloning in Streptomyces: resistance genes from antibiotic-producing species. Nature 286:525–527
Touati D (2000) Iron and oxidative stress in bacteria. Arch Biochem Biophys 373:1–6
Vazquez-Torres A (2012) Redox active thiol sensors of oxidative and nitrosative stress. Antioxid Redox Signal (in press)
Wang S, He YX, Bao R, Teng YB, Ye BP, Zhou CZ (2008) Cloning, expression, purification, crystallization and preliminary X-ray diffraction analysis of hypothetical protein SCO4226 from Streptomyces coelicolor A3(2). Acta Crystallogr Sect F Struct Biol Cryst Commun 64:847–850
Willey JM, van der Donk WA (2007) Lantibiotics: peptides of diverse structure and function. Annu Rev Microbiol 61:477–501
Yu D, Ellis HM, Lee EC, Jenkins NA, Copeland NG, Court DL (2000) An efficient recombination system for chromosome engineering in Escherichia coli. Proc Natl Acad Sci USA 97:5978–5983
Yuan ZC, Zaheer R, Finan TM (2005) Phosphate limitation induces catalase expression in Sinorhizobium meliloti, Pseudomonas aeruginosa and Agrobacterium tumefaciens. Mol Microbiol 58:877–894
Acknowledgments
This work was supported by the European program ACTINOGEN (http://www.swan.ac.uk/research/actinogen/), the Centre National de la Recherche Scientifique (http://www.cnrs.fr/), the University Paris Sud 11 (http://www.u-psud.fr), and the Pôle de Recherche et d'Enseignement Supérieur UniverSud Paris (http://www.universud-paris.fr). The authors wish to thank Barry Holland for stimulating discussions and for correction of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Emmanuelle Darbon and Cécile Martel contributed equally to the work.
Rights and permissions
About this article
Cite this article
Darbon, E., Martel, C., Nowacka, A. et al. Transcriptional and preliminary functional analysis of the six genes located in divergence of phoR/phoP in Streptomyces lividans . Appl Microbiol Biotechnol 95, 1553–1566 (2012). https://doi.org/10.1007/s00253-012-3995-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-012-3995-2