Abstract
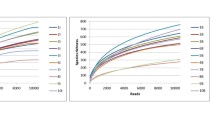
The diversity of the microbial community was identified in two lab-scale, ideally mixed sequencing batch reactors which were run for 115 days. One of the reactors was intermittently aerated (2 h aerobically/2 h anaerobically) whereas the other was consistently aerated. The amount of biomass as dry matter, the degradation of organic carbon determined by chemical oxygen demand and nitrogen-degradation activity were followed over the operation of the two reactors and did not show significant differences between the two approaches at the end of the experiment. At this point, the composition of the microbial community was determined by a terminal restriction fragment length polymorphism approach using multiple restriction enzymes by which organisms were retrieved to the lowest taxonomic level. The microbial composition was then significantly different. The species richness was at least five-fold higher in the intermittently aerated reactor than in the permanently kept aerobic approach which is in line with the observation that ecosystem disturbances result in higher diversity.


Similar content being viewed by others
References
Abwasserverordnung (1997) Verordnung über Anforderungen an das Einleiten von Abwasser in Gewässer, Bundesgesetzblatt, Neufassung vom 17. Juni 2004, I, 1108–1184
Amann RI, Ludwig W, Schleifer KH (1995) Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev 59:143–169
Blackburne R, Vadivelua VM, Yuana Z, Keller J (2007) Kinetic characterization of an enriched Nitrospira culture with comparison to Nitrobacter. Water Res 41:3033–3042
Bottari BB, Ercolini D, Gatti M, Neviani E (2006) Application of FISH technology for microbial analysis: current state and prospects. Appl Microbial Biotech 73:485–494
Bramucci M, Nagarajan V (2006) Bacterial communities in industrial wastewater bioreactors. Curr Opinion Microbiol 9:275–278
Cabezas A, Draper P, Etchebehere C (2009) Fluctuation of microbial activities after influent load variations in a full-scale SBR: recovery of the biomass after starvation. Appl Microbiol Biotech 84:1191–1202
Chen CL, Wu JH, Tseng IC, Liang TM, Liu WT (2009) Characterization of active microbes in a full scale anaerobic fluidized bed reactor treating phenolic wastewater. Microbes Environ 24:144–153
Chudoba J, Grau P, Ottova Y (1973) Control of activated sludge filamentous bulking. I. Effect of hydraulic regime or degree of mixing in an aeration tank. Water Res 7:163–172
Connell JH (1978) Diversity in tropical rain forests and coral reefs. Science 199:1302–1310
Crocetti GR, Hugenholtz P, Bond PL, Schuler A, Keller J, Jenkins D, Blackall LA (2000) Identification of phosphate-accumulating organisms and design of 16S rRNA-directed probes for their detection and quantification. Appl Environ Microbiol 66:1175–1182
Daims H, Taylor MW, Wagner M (2006) Wastewater treatment: a model system for microbial ecology. Trends Biotechnol 24:483–489
Eilmus S, Heil M (2009) Bacterial associates of arboreal ants and their putative functions in an obligate ant-plant mutualism. Appl Environ Microbiol 75:4324–4332
Eilmus S, Rösch C, Bothe H (2007) Prokaryotic life in a potash-polluted marsh with emphasis on N-metabolizing microorganisms. Environ Pollut 146:478–491
Eschenhagen M, Schuppler M, Röske I (2003) Molecular characterization of the microbial community in two activated sludge systems for the advanced treatment of domestic effluents. Water Res 37:3224–3232
Felfoldi T, Szekely AJ, Goral R, Barkacs K, Scheirich D, Andras J, Racz A, Marialigeti K (2010) Polyphasic bacterial community analysis of an aerobic activated sludge removing phenols and thiocyanate from coke plant effluent. Bioresource Technol 101:3406–3410
Forney LF, Liu W-T, Guckert JB, Kumagai Y, Namkung E, Nishihara T, Larson RL (2001) Structure of microbial communities in activated sludge: potential implications for assessing the biodegradability of chemicals. Ecotoxicol Environ Safety, section B 49:40–53
Goel R, Mino T, Satoh H, Matsuo T (1998) Enzyme activities under anaerobic and aerobic conditions in activated sludge sequencing batch reactor. Water Res 32:2081–2088
Hao Y, Winans SC, Glick B, Charles TC (2010) Identification and characterization of new LuxR/LuxI-type quorum sensing systems from metagenomics libraries. Environ Microbiol 12:105–117
Jetten M, Schmid M, Van de Pas-Schoonen K, Sinninghe Damsté JS, Strous S (2005) Anammox organisms: enrichment, cultivation and environmental analysis. Environ Microbiol 397:39–57
Juteau P, Tremblay D, Villemur R, Bisaillon J-G, Beaudet R (2005) Analysis of the bacterial community inhabiting an aerobic thermophilic sequencing batch reactor (AST-SBR) treating swine waste. Environ Biotechn 66:115–122
Kent AD, Smith DJ, Benson BJ, Triplett EW (2003) Web-based phylogenetic assignment tool for analysis of terminal restriction fragment length polymorphism profiles of microbial communities. Appl Environ Microbiol 69:6768–6776
Koenig A, Zhang T, Liu L-H, Fang HHP (2005) Microbial community and biochemistry process in autosulfurotrophic denitrifying biofilm. Chemosphere 58:1041–1047
Kong YH, Xia Y, Nielsen JL, Nielsen PH (2007) Structure and function of the microbial community in a full-scale enhanced biological phosphorus removal plant. Microbiology 153:4061–4073
Li HY, Yang M, Zhang Y, Liu XC, Gao MC, Kamagata Y (2005) Comparison of nitrification performance and microbial community between submerged membrane bioreactor and conventional activated sludge system. Water Sci Technol 51:535–543
Lin CK, Katayama Y, Hosomi M, Murakami A, Okada M (2003) The characteristics of the bacterial community structure and population dynamics for phosphorus removal in SBR activated sludge processes. Water Res 37:2944–2952
Loy A, Daims H, Wagner W (2002) Activated sludge—molecular techniques for determining community composition. In: Bitton G (ed) Encyclopedia of environmental microbiology, vol 1. Wiley, New York, pp 26–43
Marchesi JR, Sato T, Weightman A, Martin TA, Fry JC, Hiom SJ, Wade WG (1998) Design and evaluation of useful bacterium-specific PCR primers that amplify genes coding for bacterial 16S rRNA. Appl Environ Microbiol 64:795–799
Okubo Y, Futamata H, Hiraishi A (2006) Characterization of phototrophic purple nonsulfur bacteria forming colored microbial mats in a swine wastewater ditch. Appl Environ Microbiol 72:6225–6233
Rösch C, Bothe H (2005) Improved assessment of denitrifying, N2-fixing, and total-community bacteria by terminal restriction fragment length polymorphism analysis using multiple restriction enzymes. Appl Environ Microbiol 71:2026–2035
Rösch C, Bothe H (2009) Diversity of total, nitrogen-fixing and denitrifying bacteria in an acid forest soil. Eur J Soil Sciences 60:883–894
Schütte UME, Abdo Z, Bent SJ, Shyu C, Williams CJ, Pierson JD, Forney LJ (2008) Advances in the use of terminal restriction fragment length polymorphism (T-RFLP) analysis of the 16S rRNA genes to characterize microbial communities. Appl Microbiol Biotechnol 80:365–380
Shin SG, Han G, Lim J, Lee C, Hwang S (2010) A comprehensive microbial insight into two-stage anaerobic digestion of food waste-recycling wastewater. Water Research: 44:4838–4849
Stres B, Tiedje JM, Murovec B (2009) BEsTRF: a tool for optimal resolution of terminal-restriction fragment length polymorphism analysis based on user-defined primer-enzyme-sequence-databases. Bioinformatics 26:1556–1558
Tomai CM, Annessini MC, Luberti R, Cento G, Senia A (2003) Kinetics of 4-nitrophenol biodegradation in a sequencing batch reactor. Water Res 37:3803–3814
Tomlinson EJ, Chambers B (1979) Method for prevention of bulking in activated sludge. Water Poll Contr 78:524
Torsvik V, Goksøyr J, Daae FL (1990) High diversity in DNA of soil bacteria. Appl Environ Microbiol 56:782–787
Wagner M, Amann R, Lemmer H, Schleifer K-H (1993) Probing activated sludge with proteobacteria-specific oligonucleotides: inadequacy of culture-dependent methods for describing microbial community structures. Appl Environ Microbiol 59:1520–1525
Wagner M, Amann RI (1996) Die Anwendung von in situ- Hybridisierungssonden zur Aufklärung von Struktur und Dynamik der mikrobiellen Biozönosen in der Abwasserreinigung. In: Lemmer H, Flemming H-C, Griebe T (eds) Ökologie der Abwasserreinigung. Springer, Berlin, pp 93–110
Wagner M, Erhart R, Manz W, Amann R, Lemmer H, Wedi D, Schleifer K-H (1994) Development of a rRNA-targeted oligonucleotide probe for the genus Acinetobacter and its application for in situ monitoring of activated sludge. Appl Environ Microbiol 60:792–800
Wagner M, Loy A (2002) Bacterial community composition and function in sewage treatment plants. Curr Opin Biotechn 13:218–227
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Native Bayesian classifier for rapid assignment of rRNS sequences into bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Wilderer PA, Silverstein JA (1983) Bedeutung periodischer Stress-Bedingungen für die Wirksamkeit biologischer Klärverfahren. GWF Wasser Abwasser 124:546–552
Wilen B-M, Onuki M, Hermansson M, Lumley D, Mino T (2008) Microbial community structure in activated sludge floc analysed by fluorescence in situ hybridization and its relation to floc stability. Water Research 42:2300–2308
Woolard CR, Irvine RL (1995) Treatment of hypersaline wastewater in the sequencing batch reactor. Water Res 29:1159–1168
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Denecke, M., Eilmus, S., Röder, N. et al. Molecular identification of the microbial diversity in two sequencing batch reactors with activated sludge. Appl Microbiol Biotechnol 93, 1725–1734 (2012). https://doi.org/10.1007/s00253-011-3474-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-011-3474-1