Abstract
Introduction
Hitherto the production of the biopolymer cyanophycin (CGP) using recombinant Escherichia coli strains and cheap mineral salts medium yielded only trace amounts of CGP (<0.5%, w/w) of the cell dry matter (CDM). This was probably due to the instability of the plasmids encoding the cyanophycin synthetase.
Material and methods
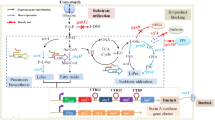
In this study, we developed an anabolism-based media-dependent plasmid addiction system (PAS) to enhance plasmid stability, and we established a process based on a modified mineral salts medium yielding a CGP content of 42% (w/w) at the maximum without the addition of amino acids to the medium for the first time. This PAS is based on different lysine biosynthesis pathways and consists of two components: (1) a knockout of the chromosomal dapE disrupts the native succinylase pathway in E. coli and (2) the complementation by the plasmid-encoded artificial aminotransferase pathway mediated by the dapL gene from Synechocystis sp. PCC 6308, which allows the synthesis of the essential lysine precursor L,L-2,6-diaminopimelate. In addition, this plasmid also harbors cphAC595S, an engineered cyanophycin synthetase gene responsible for CGP production.
Results
Cultivation experiments in Erlenmeyer flask and also in bioreactors in mineral salts medium without antibiotics revealed an at least 4.5-fold enhanced production of CGP in comparison to control cultivations without PAS.
Discussion
Fermentation experiments with culture volume of up to 400 l yielded a maximum of 18% CGP (w/w) and a final cell density of 15.2 g CDM/l. Lactose was used constantly as an effective inducer and carbon source. Thus, we present a convenient option to produce CGP with E. coli at a technical scale without the need to add antibiotics or amino acids using the mineral salts medium designed in this study.






Similar content being viewed by others
References
Aboulmagd E, Oppermann-Sanio FB, Steinbüchel A (2000) Molecular characterization of the cyanophycin synthetase from Synechocystis sp. strain PCC6308. Arch Microbiol 174:297–306
Aboulmagd E, Voss I, Oppermann-Sanio FB, Steinbüchel A (2001) Heterologous expression of cyanophycin synthetase and cyanophycin synthesis in the industrial relevant bacteria Corynebacterium glutamicum and Ralstonia eutropha and in Pseudomonas putida. Biomacromolecules 2:1338–1342
Bukhari AI, Taylor AL (1971) Genetic analysis of diaminopimelic acid- and lysine-requiring mutants of Escherichia coli. J Bacteriol 105:844–854
Diniz SC, Voss I, Steinbüchel A (2006) Optimization of cyanophycin production in recombinant strains of Pseudomonas putida and Ralstonia eutropha employing elementary mode analysis and statistical experimental design. Biotechnol Bioeng 93:698–717
Elbahloul Y, Krehenbrink M, Reichelt R, Steinbüchel A (2005) Physiological conditions conducive to high cyanophycin content in biomass of Acinetobacter calcoaceticus strain ADP1. Appl Environ Microbiol 71:858–866
Frey KM, Oppermann-Sanio FB, Schmidt H, Steinbüchel A (2002) Technical-scale production of cyanophycin with recombinant strains of Escherichia coli. Appl Environ Microbiol 68:3377–3384
Füser G, Steinbüchel A (2007) Analysis of genome sequences for genes of cyanophycin metabolism: identifying putative cyanophycin metabolizing prokaryotes. Macromol Biosci 7:278–296
Hai T, Frey KM, Steinbüchel A (2006) Engineered cyanophycin synthetase (CphA) from Nostoc ellipsosporum confers enhanced CphA activity and cyanophycin accumulation to Escherichia coli. Appl Environ Microbiol 72:7652–7660
Hartree EF (1972) Determination of protein: a modification of the Lowry method that gives a linear photometric response. Anal Biochem 48:422–427
Hudson AO, Singh BK, Leustek T, Gilvarg C (2006) An LL-diaminopimelate aminotransferase defines a novel variant of the lysine biosynthesis pathway in plants. Plant Physiol 140:292–301
Joentgen W, Groth T, Steinbüchel A, Hai T, Oppermann F (1998) Polyasparaginic acid homopolymers and copolymers, biotechnical production and use thereof. Patent application WO 98/39090
Krehenbrink M, Oppermann-Sanio F, Steinbüchel A (2002) Evaluation of non-cyanobacterial genome sequences for occurrence of genes encoding proteins homologous to cyanophycin synthetase and cloning of an active cyanophycin synthetase from Acinetobacter sp. strain DSM 587. Arch Microbiol 177:371–380
Kroll J, Steinle A, Reichelt R, Ewering C, Steinbüchel A (2009) Establishment of a novel anabolism-based addiction system with an artificially introduced mevalonate pathway: complete stabilization of plasmids as universal application in white biotechnology. Metab Eng 11:168–177
Kroll J, Klinter S, Schneider C, Voß I, Steinbüchel A (2010) Plasmid addiction systems: perspectives and applications in biotechnology. Microb Biotechnol. doi:10.1111./j.1751-7915.2010.00170.x
McCoy AJ, Adams NE, Hudson AO, Gilvarg C, Leustek T, Maurelli AT (2006) L, L-diaminopimelate aminotransferase, a trans-kingdom enzyme shared by Chlamydia and plants for synthesis of diaminopimelate/lysine. Proc Natl Acad Sci USA 103:17909–17914
Misono H, Togawa H, Yamamoto T, Soda K (1976) Occurrence of meso-alpha, epsilon-diaminopimelate dehydrogenase in Bacillus sphaericus. Biochem Biophys Res Commun 72:89–93
Mnif B, Vimont S, Boyd A, Bourit E, Picard B, Branger C, Denamur E, Arlet G (2010) Molecular characterization of addiction systems of plasmids encoding extended-spectrum beta-lactamases in Escherichia coli. J Antimicrob Chemother 65:1599–1603
Mooibroek H, Oosterhuis N, Giuseppin M, Toonen M, Franssen H, Scott E, Sanders J, Steinbüchel A (2007) Assessment of technological options and economical feasibility for cyanophycin biopolymer and high-value amino acid production. Appl Microbiol Biotechnol 77:257–267
Neumann K, Stephan DP, Ziegler K, Huhns M, Broer I, Lockau W, Pistorius EK (2005) Production of cyanophycin, a suitable source for the biodegradable polymer polyaspartate, in transgenic plants. Plant Biotechnol J 3:249–258
Oppermann-Sanio FB, Steinbüchel A (2002) Occurrence, functions and biosynthesis of polyamides in microorganisms and biotechnological production. Naturwissenschaften 89:11–22
Pfennig N (1974) Rhodopseudomonas globiformis sp. nov., a new species of the Rhodospirillaceae. Arch Microbiol 100:197–206
Richter R, Hejazi M, Kraft R, Ziegler K, Lockau W (1999) Cyanophycinase, a peptidase degrading the cyanobacterial reserve material multi-l-arginyl-poly-l-aspartic acid (cyanophycin): molecular cloning of the gene of Synechocystis sp. PCC 6803, expression in Escherichia coli, and biochemical characterization of the purified enzyme. Eur J Biochem 263:163–169
Sallam A, Steinbüchel A (2010) Dipeptides in nutrition and therapy: cyanophycin-derived dipeptides as natural alternatives and their biotechnological production. Appl Microbiol Biotechnol 87:815–828
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual 2nd edition. Cold Spring Harbor Laboratory Press, N.Y
Sanders J, Scott E, Weusthuis R, Mooibroek H (2007) Bio-refinery as the bio-inspired process to bulk chemicals. Macromol Biosci 7:105–117
Sanger F, Nicklen FS, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA 74:5463–5467
Schleifer KH, Kandler O (1972) Peptidoglycan types of bacterial cell walls and their taxonomic implications. Bacteriol Rev 36:407–477
Schwamborn M (1998) Chemical synthesis of polyaspartates: a biodegradable alternative to currently used polycarboxylate homo- and copolymers. Polym Degrad Stab 59:39–45
Scott E, Peter F, Sanders J (2007) Biomass in the manufacture of industrial products—the use of proteins and amino acids. Appl Microbiol Biotechnol 75:751–762
Simon RD, Weathers P (1976) Determination of the structure of the novel polypeptide containing aspartic acid and arginine which is found in cyanobacteria. Biochim Biophys Acta 420:165–176
Steinle A, Oppermann-Sanio FB, Reichelt R, Steinbüchel A (2008) Synthesis and accumulation of cyanophycin in transgenic strains of Saccharomyces cerevisiae. Appl Environ Microbiol 74:3410–3418
Terpe K (2006) Overview of bacterial expression systems for heterologous protein production: from molecular and biochemical fundamentals to commercial systems. Appl Microbiol Biotechnol 72:211–222
Velasco AM, Leguina JI, Lazcano A (2002) Molecular evolution of the lysine biosynthetic pathways. J Mol Evol 55:445–459
Voß I, Steinbüchel A (2006) Application of a KDPG-aldolase gene-dependent addiction system for enhanced production of cyanophycin in Ralstonia eutropha strain H16. Metab Eng 8:66–78
Voß I, Diniz SC, Aboulmagd E, Steinbüchel A (2004) Identification of the Anabaena sp. strain PCC7120 cyanophycin synthetase as suitable enzyme for production of cyanophycin in Gram-negative bacteria like Pseudomonas putida and Ralstonia eutropha. Biomacromolecules 5:1588–1595
Wehrmann A, Eggeling L, Sahm H (1994) Analysis of different DNA fragments of Corynebacterium glutamicum complementing dapE of Escherichia coli. Microbiology 140:3349–3356 (SGM)
Weinberger S, Gilvarg C (1970) Bacterial distribution of the use of succinyl and acetyl blocking groups in diaminopimelic acid biosynthesis. J Bacteriol 101:323–324
Ziegler K, Diener A, Herpin C, Richter R, Deutzmann R, Lockau W (1998) Molecular characterization of cyanophycin synthetase, the enzyme catalyzing the biosynthesis of the cyanobacterial reserve material multi-l-arginyl-poly-l-aspartate (cyanophycin). Eur J Biochem 254:154–159
Ziegler K, Deutzmann R, Lockau W (2002) Cyanophycin synthetase-like enzymes of non-cyanobacterial eubacteria: characterization of the polymer produced by a recombinant synthetase of Desulfitobacterium hafniense. Z Naturforsch C 57:522–529
Acknowledgements
The provision of plasmid pET30a::dapL Ss by Prof. Dr. Thomas Leustek is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kroll, J., Klinter, S. & Steinbüchel, A. A novel plasmid addiction system for large-scale production of cyanophycin in Escherichia coli using mineral salts medium. Appl Microbiol Biotechnol 89, 593–604 (2011). https://doi.org/10.1007/s00253-010-2899-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-010-2899-2