Abstract.
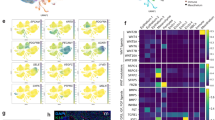
To search for genes that participate in regulatory networks sustaining mouse embryonic T-cell development, we have performed expression profiling using nylon macroarrays. Labeled samples representative of individual developmental stages were utilized, taking advantage of cell homogeneity during early thymus ontogeny. cDNAs revealing differential expression were further selected using labeled samples derived from lymphoid versus non-lymphoid tissues, and from mutant thymi exhibiting T-cell developmental defects. We thus identified clusters of coexpressed genes during T-cell embryogenesis and characterized their sequences through bioinformatics. We compare our results with those from other profiling analyses in the immune system, and discuss their implications for the definition of genes whose products are involved in T-cell development.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Electronic Publication
Rights and permissions
About this article
Cite this article
Ramialison, M., Mohr, E., Nal, B. et al. Expression profiling in mouse fetal thymus reveals clusters of coordinately expressed genes that mark individual stages of T-cell ontogeny. Immunogenetics 54, 469–478 (2002). https://doi.org/10.1007/s00251-002-0488-y
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00251-002-0488-y