Abstract
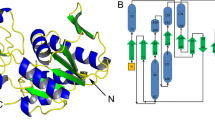
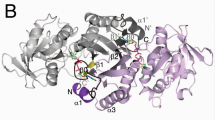
Phosphopantetheine adenylyltransferase (EC. 2.7.7.3, PPAT) catalyzes the penultimate step of the multistep reaction in the coenzyme A (CoA) biosynthesis pathway. In this step, an adenylyl group from adenosine triphosphate (ATP) is transferred to 4′-phosphopantetheine (PNS) yielding 3′-dephospho-coenzyme A (dpCoA) and pyrophosphate (PPi). PPAT from strain C3 of Klebsiella pneumoniae (KpPPAT) was cloned, expressed and purified. It was crystallized using 0.1 M HEPES buffer and PEG10000 at pH 7.5. The crystals belonged to tetragonal space group P41212 with cell dimensions of a = b = 72.82 Å and c = 200.37 Å. The structure was determined using the molecular replacement method and refined to values of 0.208 and 0.255 for Rcryst and Rfree factors, respectively. The structure determination showed the presence of three crystallographically independent molecules A, B and C in the asymmetric unit. The molecules A and B are observed in the form of a dimer in the asymmetric unit while molecule C belongs to the second dimer whose partner is related by crystallographic twofold symmetry. The polypeptide chain of KpPPAT folds into a β/α structure. The conformations of the side chains of several residues in the substrate binding site in KpPPAT are significantly different from those reported in other PPATs. As a result, the modes of binding of substrates, phosphopantetheine (PNS) and adenosine triphosphate (ATP) differ considerably. The binding studies using fluorescence spectroscopy indicated a KD value of 3.45 × 10−4 M for ATP which is significantly lower than the corresponding values reported for PPAT from other species.








Similar content being viewed by others
Data availability
The coordinates and structure factors for KpPPAT have been deposited in the Protein Data Bank, under accession numbers 8I8I. All data other than X-ray crystallography supporting the findings of this study are available within the paper and its Supplementary Information.
References
Aghajanian S, Worrall DM (2002) Identification and characterization of the gene encoding the human phosphopantetheine adenylyltransferase and dephospho-CoA kinase bifunctional enzyme (CoA synthase). Bioche J 365:13–18. https://doi.org/10.1042/bj20020569
Agirre J, Atanasova M, Bagdonas H et al (2023) The CCP 4 suite: integrative software for macromolecular crystallography. Acta Crystallogr D Struct Biol 79:449–461. https://doi.org/10.1107/S2059798323003595
Badger J, Sauder JM, Adams JM et al (2005) Structural analysis of a set of proteins resulting from a bacterial genomics project. Proteins: Struct Funct Bioinform 60:787–796. https://doi.org/10.1002/prot.20541
Bork P, Holm L, Koonin EV, Sander C (1995) The cytidylyltransferase superfamily: identification of the nucleotide-binding site and fold prediction. Proteins: Struct Funct Genet 22:259–266. https://doi.org/10.1002/prot.340220306
Brüser T, Selmer T, Dahl C (2000) “ADP sulfurylase” from Thiobacillus denitrificansis an adenylylsulfate: phosphate adenylyltransferase and belongs to a new family of nucleotidyltransferases. J Biol Chem 275:1691–1698. https://doi.org/10.1074/jbc.275.3.1691
Chatterjee R, Mondal A, Basu A, Datta S (2016) Transition of phosphopantetheine adenylyltransferase from catalytic to allosteric state is characterized by ternary complex formation in Pseudomonas aeruginosa. Biochimica Et Biophysica Acta (BBA) - Proteins Proteom 1864:773–786. https://doi.org/10.1016/j.bbapap.2016.03.018
Cusack S (1997) Aminoacyl-tRNA synthetases. Curr Opin Struct Biol 7:881–889. https://doi.org/10.1016/S0959-440X(97)80161-3
D’Angelo I, Raffaelli N, Dabusti V et al (2000) Structure of nicotinamide mononucleotide adenylyltransferase: a key enzyme in NAD+ biosynthesis. Structure 8:993–1004. https://doi.org/10.1016/S0969-2126(00)00190-8
Delarue M (1995) Aminoacyl-tRNA synthetases. Curr Opin Struct Biol 5:48–55. https://doi.org/10.1016/0959-440X(95)80008-O
Edwards TE, Leibly DJ, Bhandari J et al (2011) Structures of phosphopantetheine adenylyltransferase from Burkholderia pseudomallei. Acta Crystallogr Sect F Struct Biol Cryst Commun 67:1032–1037. https://doi.org/10.1107/S1744309111004349
Emsley P, Cowtan K (2004) Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr 60:2126–2132. https://doi.org/10.1107/S0907444904019158
Franklin MC, Cheung J, Rudolph MJ et al (2015) Structural genomics for drug design against the pathogen Coxiella burnetii. Proteins: Struct Funct Bioinform 83:2124–2136. https://doi.org/10.1002/prot.24841
Gerdes SY, Scholle MD, D’Souza M et al (2002) From genetic footprinting to antimicrobial drug targets: examples in cofactor biosynthetic pathways. J Bacteriol 184:4555–4572. https://doi.org/10.1128/JB.184.16.4555-4572.2002
Gupta A, Singh PK, Iqbal N et al (2019) Structural and binding studies of phosphopantetheine adenylyl transferase from Acinetobacter baumannii. Biochimica Et Biophysica Acta (BBA) - Proteins Proteom 1867:537–547. https://doi.org/10.1016/j.bbapap.2019.03.002
Izard T (2002) The crystal structures of phosphopantetheine adenylyltransferase with bound substrates reveal the enzyme’s catalytic mechanism 1 1Edited by K. Nagai J Mol Biol 315:487–495. https://doi.org/10.1006/jmbi.2001.5272
Izard T (2003) A novel adenylate binding site confers phosphopantetheine adenylyltransferase interactions with coenzyme A. J Bacteriol 185:4074–4080. https://doi.org/10.1128/JB.185.14.4074-4080.2003
Kabsch W (2010) XDS. Acta Crystallogr D Biol Crystallogr 66:125–132. https://doi.org/10.1107/S0907444909047337
Krissinel E (2010) Crystal contacts as nature’s docking solutions. J Comput Chem 31:133–143. https://doi.org/10.1002/jcc.21303
Krissinel E, Henrick K (2005) Detection of protein assemblies in crystals. In: Berthold MR, Glen RC, Diederichs K, Kohlbacher O, Fischer I (eds) Computational life sciences. Lecture Notes in Computer Science, vol 3695, pp 163–174. https://doi.org/10.1007/11560500_15
Krissinel E, Henrick K (2007) Inference of macromolecular assemblies from crystalline state. J Mol Biol 372:774–797. https://doi.org/10.1016/j.jmb.2007.05.022
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26:283–291. https://doi.org/10.1107/S0021889892009944
Lee HH, Yoon H-J, Kang JY et al (2009) The structure of Staphylococcus aureus phosphopantetheine adenylyltransferase in complex with 3′-phosphoadenosine 5′-phosphosulfate reveals a new ligand-binding mode. Acta Crystallogr Sect F Struct Biol Cryst Commun 65:987–991. https://doi.org/10.1107/S1744309109036616
Magill SS, Edwards JR, Bamberg W et al (2014) Multistate point-prevalence survey of health care-associated infections. N Engl J Med 370:1198–1208. https://doi.org/10.1056/NEJMoa1306801
Murshudov GN, Vagin AA, Dodson EJ (1997) Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr D Biol Crystallogr 53:240–255. https://doi.org/10.1107/S0907444996012255
Park YS, Gee P, Sanker S et al (1997) Identification of functional conserved residues of CTP:glycerol-3-phosphate cytidylyltransferase. J Biol Chem 272:15161–15166. https://doi.org/10.1074/jbc.272.24.15161
Perona JJ, Rould MA, Steitz TA (1993) Structural basis for transfer RNA aminoacylation by Escherichia coli glutaminyl-tRNA synthetase. Biochemistry 32:8758–8771. https://doi.org/10.1021/bi00085a006
Podschun R, Ullmann U (1998) Klebsiella spp. as nosocomial pathogens: epidemiology, taxonomy, typing methods, and pathogenicity factors. Clin Microbiol Rev 11:589–603. https://doi.org/10.1128/CMR.11.4.589
Potterton E, Briggs P, Turkenburg M, Dodson E (2003) A graphical user interface to the CCP 4 program suite. Acta Crystallogr D Biol Crystallogr 59:1131–1137. https://doi.org/10.1107/S0907444903008126
Proudfoot A, Frank AO, Ruggiu F et al (2016) Facilitating unambiguous NMR assignments and enabling higher probe density through selective labeling of all methyl containing amino acids. J Biomol NMR 65:15–27. https://doi.org/10.1007/s10858-016-0032-2
Ramachandran GN, Sasisekharan V (1968) Conformation of polypeptides and proteins. Adv Protein Chem 23:283–438. https://doi.org/10.1016/s0065-3233(08)60402-7
Read RJ, Adams PD, Arendall WB et al (2011) A new generation of crystallographic validation tools for the Protein Data Bank. Structure. https://doi.org/10.1016/j.str.2011.08.006
Robishaw JD, Neely JR (1985) Coenzyme A metabolism. Am J Physiol-Endocrinol Metab 248:E1–E9. https://doi.org/10.1152/ajpendo.1985.248.1.E1
Rossman MG, Liljas A, Branden CI (1975) The enzymes, 3rd edn. Academic Press, New York
Spry C, Kirk K, Saliba KJ (2008) Coenzyme A biosynthesis: an antimicrobial drug target. FEMS Microbiol Rev 32:56–106. https://doi.org/10.1111/j.1574-6976.2007.00093.x
Takahashi H, Inagaki E, Fujimoto Y et al (2004) Structure and implications for the thermal stability of phosphopantetheine adenylyltransferase from Thermus thermophilus. Acta Crystallogr D Biol Crystallogr 60:97–104. https://doi.org/10.1107/S0907444903025319
Thomas SE, Mendes V, Kim SY et al (2017) Structural biology and the design of new therapeutics: from HIV and cancer to mycobacterial infections. J Mol Biol 429:2677–2693. https://doi.org/10.1016/j.jmb.2017.06.014
Timofeev V, Smirnova E, Chupova L et al (2012) X-ray study of the conformational changes in the molecule of phosphopantetheine adenylyltransferase from Mycobacterium tuberculosis during the catalyzed reaction. Acta Crystallogr D Biol Crystallogr 68:1660–1670. https://doi.org/10.1107/S0907444912040206
Vagin A, Teplyakov A (2010) Molecular replacement with MOLREP. Acta Crystallogr D Biol Crystallogr 66:22–25. https://doi.org/10.1107/S0907444909042589
Venkatachalam KV, Fuda H, Koonin EV, Strott CA (1999) Site-selected mutagenesis of a conserved nucleotide binding HXGH motif located in the ATP sulfurylase domain of human bifunctional 3′-phosphoadenosine 5′-phosphosulfate synthase. J Biol Chem 274:2601–2604. https://doi.org/10.1074/jbc.274.5.2601
von Delft F, Lewendon A, Dhanaraj V et al (2001) The crystal structure of E. coli pantothenate synthetase confirms it as a member of the cytidylyltransferase superfamily. Structure 9:439–450. https://doi.org/10.1016/S0969-2126(01)00604-9
Weber CH, Park YS, Sanker S et al (1999) A prototypical cytidylyltransferase: CTP: glycerol-3-phosphate cytidylyltransferase from Bacillus subtilis. Structure 7:1113–1124. https://doi.org/10.1016/S0969-2126(99)80178-6
Wubben TJ, Mesecar AD (2010) Kinetic, thermodynamic, and structural insight into the mechanism of phosphopantetheine adenylyltransferase from mycobacterium tuberculosis. J Mol Biol 404:202–219. https://doi.org/10.1016/j.jmb.2010.09.002
Yasutake Y, Ota H, Hino E et al (2011) Structures of Burkholderia thailandensis nucleoside kinase: implications for the catalytic mechanism and nucleoside selectivity. Acta Crystallogr D Biol Crystallogr 67:945–956. https://doi.org/10.1107/S0907444911038777
Yoon H-J, Kang JY, Mikami B et al (2011) Crystal structure of phosphopantetheine adenylyltransferase from Enterococcus faecalis in the ligand-unbound state and in complex with ATP and pantetheine. Mol Cells 32:431–435. https://doi.org/10.1007/s10059-011-0102-y
Acknowledgements
Authors gratefully acknowledge the support from the Department of Biotechnology (DBT), Ministry of Science and Technology, Government of India, New Delhi for sponsoring the Synchrotron beamlines at the European Synchrotron Radiation Facility (ESRF), Grenoble, France and the Indian Council of Medical Research (ICMR), New Delhi for a grant to SS. TPS thanks the Science and Engineering Research Board (SERB), New Delhi for the grant of a Distinguished Fellow position to him under the National Science Chair Program of SERB. NA thanks the University Grants Commission, New Delhi for the award of a fellowship to him.
Funding
This work was supported by a project (Grant No. I-1251) from the Indian Council of Medical Research (ICMR), New Delhi to SS and a grant of a Distinguished Fellow position under the National Science Chair (Grant No. SB/DF/002/2019) to TPS from the Science and Engineering Research Board (SERB), New Delhi of the Government of India.
Author information
Authors and Affiliations
Contributions
Conceptualization: SS, NA, TPS; Methodology: NA, PS, SS, TPS; Formal analysis and investigation: NA, SS, TPS; Writing-original draft preparation: TPS, SS, NA; Writing-review and editing: NA, SS, TPS; Funding acquisition: SS, TPS; Resources: SS, TPS, NA, PS; Supervision: SS, TPS.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ahmad, N., Sharma, P., Sharma, S. et al. Structure of a novel form of phosphopantetheine adenylyltransferase from Klebsiella pneumoniae at 2.59 Å resolution. Eur Biophys J 53, 147–157 (2024). https://doi.org/10.1007/s00249-024-01703-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00249-024-01703-1