Abstract
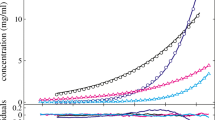
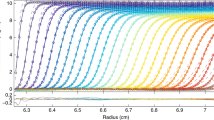
A recent investigation was aimed at obtaining structural information on a highly extended protein via SEC-MALS-SAXS. Significantly broadened elution peaks were observed, reminiscent of a phenomenon known as viscous fingering. This phenomenon is usually observed above 50 mg/mL for proteins like bovine serum albumin (BSA). Interestingly, the highly extended protein (Brpt5.5) showed viscous fingering at concentrations lower than 5 mg/mL. The current study explores this and other non-ideal behavior, emphasizing the presence of these effects at relatively low concentrations for extended proteins. BSA, Brpt5.5, and a truncated form of Brpt5.5 referred to as Brpt1.5 are studied systematically using size-exclusion chromatography (SEC), sedimentation velocity analytical ultracentrifugation (AUC), and viscosity. The viscous fingering effect is quantified using two approaches and is found to correlate well with the intrinsic viscosity of the proteins—Brpt5.5 exhibits the most severe effect and is the most extended protein tested in the study. By AUC, the hydrodynamic non-ideality was measured for each protein via global analysis of a concentration series. Compared to BSA, both Brpt1.5 and Brpt5.5 showed significant non-ideality that could be easily visualized at concentrations at or below 5 mg/mL and 1 mg/mL, respectively. A variety of relationships were examined for their ability to differentiate the proteins by shape using information from AUC and/or viscosity. Furthermore, these relationships were also tested in the context of hydrodynamic modeling. The importance of considering non-ideality when investigating the structure of extended macromolecules is discussed.





Similar content being viewed by others
Data availability
Data is available from the authors upon request.
References
Brautigam CA (2015) Calculations and publication-quality illustrations for analytical ultracentrifugation data. Methods Enzymol 562:109–133. https://doi.org/10.1016/bs.mie.2015.05.001
Brautigam CA, Tso S-C, Deka RK, Liu WZ, Norgard MV (2020) Using modern approaches to sedimentation velocity to detect conformational changes in proteins. Eur Biophys J 49:729–743. https://doi.org/10.1007/s00249-020-01453-w
Brookes E, Demeler B, Rosano C, Rocco M (2010) The implementation of somo (solution modeller) in the ultrascan analytical ultracentrifugation data analysis suite: Enhanced capabilities allow the reliable hydrodynamic modeling of virtually any kind of biomacromolecule. Eur Biophys J 39:423–435. https://doi.org/10.1007/s00249-009-0418-0
Bujacz A (2012) Structures of bovine, equine and leporine serum albumin. Acta Crystallogr D Biol Crystallogr 68:1278–1289. https://doi.org/10.1107/s0907444912027047
Byron O (1997) Construction of hydrodynamic bead models from high-resolution X-ray crystallographic or nuclear magnetic resonance data. Biophys J 72:408–415. https://doi.org/10.1016/S0006-3495(97)78681-8
Chaton CT, Herr AB (2015) Elucidating complicated assembling systems in biology using size-and-shape analysis of sedimentation velocity data. Methods Enzymol 562:187–204. https://doi.org/10.1016/bs.mie.2015.04.004
Chaton CT, Herr AB (2017) Defining the metal specificity of a multifunctional biofilm adhesion protein. Protein Sci 26:1964–1973. https://doi.org/10.1002/pro.3232
Comper WD, Preston BN (1992) The analytical ultracentrifuge as a tool for diffusion measurements. Cross diffusion effects in ternary polymer: polymer: solvent systems. In: Harding SE, Rowe AJ, Horton JC (eds) Analytical ultracentrifugation in biochemisty and polymer science. CRC Press, London, pp 428–442
Conrady DG, Brescia CC, Horii K, Weiss AA, Hassett DJ, Herr AB (2008) A zinc-dependent adhesion module is responsible for intercellular adhesion in staphylococcal biofilms. Proc Natl Acad Sci USA 105:19456–19461. https://doi.org/10.1073/pnas.0807717105
Conrady DG, Wilson JJ, Herr AB (2013) Structural basis for Zn2+-dependent intercellular adhesion in staphylococcal biofilms. Proc Natl Acad Sci USA 110:E202-211. https://doi.org/10.1073/pnas.1208134110
Correia JJ, Wright RT, Sherwood PJ, Stafford WF (2020) Analysis of nonideality: Insights from high concentration simulations of sedimentation velocity data. Eur Biophys J. https://doi.org/10.1007/s00249-020-01474-5
Correia JJ, Stafford WF (2015) Chapter three-sedimentation velocity: a classical perspective. In: James LC (ed) Methods in enzymology, vol 562. Academic Press, London, pp 49–80. https://doi.org/10.1016/bs.mie.2015.06.042
Correia JJ, Lyons DF, Sherwood P, Stafford WF (2016) Techniques for dissecting the johnston-ogston effect. In: Uchiyama S, Arisaka F, Stafford WF, Laue T (eds) Analytical ultracentrifugation: Instrumentation, software, and applications. Springer, Tokyo, pp 483–498. https://doi.org/10.1007/978-4-431-55985-6_24
Creeth JM, Knight CG (1965) On the estimation of the shape of macromolecules from sedimentation and viscosity measurements. Biochim Biophys Acta (BBA) Biophys Including Photosynthesis 102:549–558. https://doi.org/10.1016/0926-6585(65)90145-7
Du Z et al (2021) The trrosetta server for fast and accurate protein structure prediction. Nat Protoc 16:5634–5651. https://doi.org/10.1038/s41596-021-00628-9
Einstein A (1906) Zur theorie der brownschen bewegung. Ann Physik 19:289–305
Einstein A (1911) Bemerkung zu dem gesetz von eötvös. Ann Physik 34:591–593
Elez K, Bonvin AMJJ, Vangone A (2018) Distinguishing crystallographic from biological interfaces in protein complexes: role of intermolecular contacts and energetics for classification. BMC Bioinform 19:438. https://doi.org/10.1186/s12859-018-2414-9
Fleming PJ, Fleming KG (2018) Hullrad: fast calculations of folded and disordered protein and nucleic acid hydrodynamic properties. Biophys J 114:856–869. https://doi.org/10.1016/j.bpj.2018.01.002
Fleming PJ, Correia JJ, Fleming KG (2023) Revisiting macromolecular hydration with hullradsas. Eur Biophys J. https://doi.org/10.1007/s00249-022-01627-8
Flodin P (1961) Methodological aspects of gel filtration with special reference to desalting operations. J Chromatogr A 5:103–115. https://doi.org/10.1016/S0021-9673(01)92827-4
Ghirlando R, Zhao H, Balbo A, Piszczek G, Curth U, Brautigam CA, Schuck P (2014) Measurement of the temperature of the resting rotor in analytical ultracentrifugation. Anal Biochem 458:37–39. https://doi.org/10.1016/j.ab.2014.04.029
Harding SE (1995) On the hydrodynamic analysis of macromolecular conformation. Biophys Chem 55:69–93. https://doi.org/10.1016/0301-4622(94)00143-8
Harding SE (1997) The intrinsic viscosity of biological macromolecules. Progress in measurement, interpretation and application to structure in dilute solution. Prog Biophys Mol Biol 68:207–262. https://doi.org/10.1016/S0079-6107(97)00027-8
Harding SE, Horton JC, Cölfen H (1997) The ellips suite of macromolecular conformation algorithms. Eur Biophys J 25:347–359. https://doi.org/10.1007/s002490050048
Hayes DB, Stafford WF (2010) Sedview, real-time sedimentation analysis. Macromol Biosci 10:731–735. https://doi.org/10.1002/mabi.201000075
Huggins ML (1942) The viscosity of dilute solutions of long-chain molecules. IV. Dependence on concentration. J Am Chem Soc 64:2716–2718. https://doi.org/10.1021/ja01263a056
Jeansonne MS, Foley JP (1991) Review of the exponentially modified Gaussian (EMG) function since 1983. J Chromatogr Sci 29:258–266. https://doi.org/10.1093/chromsci/29.6.258
Juba D, Audus DJ, Mascagni M, Douglas JF, Keyrouz W (2017) Zeno: software for calculating hydrodynamic, electrical, and shape properties of polymer and particle suspensions. J Res Natl Inst Stand Technol 122:1–2. https://doi.org/10.6028/jres.122.020
Jumper J et al (2021) Highly accurate protein structure prediction with alphafold. Nature 596:583–589. https://doi.org/10.1038/s41586-021-03819-2
Kang EH, Mansfield ML, Douglas JF (2004) Numerical path integration technique for the calculation of transport properties of proteins. Phys Rev E Stat Nonlin Soft Matter Phys 69:031918. https://doi.org/10.1103/PhysRevE.69.031918
Khasa H, Kilby G, Chen X, Wang C (2021) Analytical band centrifugation for the separation and quantification of empty and full AAV particles. Mol Therapy Methods Clin Dev 21:585–591. https://doi.org/10.1016/j.omtm.2021.04.008
Kraemer EO (1938) Molecular weights of celluloses and cellulose derivates. Ind Eng Chem 30:1200–1203. https://doi.org/10.1021/ie50346a023
Laue TM, Shah BD, Ridgeway TM, Pelletier SL (1992) Computer-aided interpretation of analytical sedimentation data for proteins. In: Harding SE, Rowe AJ, Horton JC (eds) Analytical ultracentrifugation in biochemistry and polymer science. Royal Society of Chemistry, London, pp 90–125
Luo J, Liu Z, Guo Y, Li M (2015) A structural dissection of large protein-protein crystal packing contacts. Sci Rep 5:14214. https://doi.org/10.1038/srep14214
Marx DC, Leblanc MJ, Plummer AM, Krueger S, Fleming KG (2020) Domain interactions determine the conformational ensemble of the periplasmic chaperone sura. Protein Sci. https://doi.org/10.1002/pro.3924
Monsen RC, Chakravarthy S, Dean WL, Chaires JB, Trent JO (2021) The solution structures of higher-order human telomere g-quadruplex multimers. Nucl Acids Res 49:1749–1768. https://doi.org/10.1093/nar/gkaa1285
Nishihara T, Doty P (1958) The sonic fragmentation of collagen macromolecules. Proc Natl Acad Sci USA 44:411–417. https://doi.org/10.1073/pnas.44.5.411
Philo JS (2000) A method for directly fitting the time derivative of sedimentation velocity data and an alternative algorithm for calculating sedimentation coefficient distribution functions. Anal Biochem 279:151–163. https://doi.org/10.1006/abio.2000.4480
Philo JS (2023) Sednterp: a calculation and database utility to aid interpretation of analytical ultracentrifugation and light scattering data. Eur Biophys J. https://doi.org/10.1007/s00249-023-01629-0
Plante LD, Romano PM, Fernandez EJ (1994) Viscous fingering in chromatography visualized via magnetic resonance imaging. Chem Eng Sci 49:2229–2241. https://doi.org/10.1016/0009-2509(94)E0046-S
Rocco M, Byron O (2015a) Computing translational diffusion and sedimentation coefficients: an evaluation of experimental data and programs. Eur Biophys J 44:417–431. https://doi.org/10.1007/s00249-015-1042-9
Rocco M, Byron O (2015b) Hydrodynamic modeling and its application in AUC. Methods Enzymol 562:81–108. https://doi.org/10.1016/bs.mie.2015.04.010
Schneider CM, Cölfen H (2018) Analytical band centrifugation revisited. Eur Biophys J 47:799–807. https://doi.org/10.1007/s00249-018-1315-1
Schneider CM, Haffke D, Cölfen H (2018) Band sedimentation experiment in analytical ultracentrifugation revisited. Anal Chem 90:10659–10663. https://doi.org/10.1021/acs.analchem.8b02768
Schuck P (2000) Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and lamm equation modeling. Biophys J 78:1606–1619. https://doi.org/10.1016/s0006-3495(00)76713-0
Shelton CL, Conrady DG, Herr AB (2017) Functional consequences of b-repeat sequence variation in the staphylococcal biofilm protein Aap: deciphering the assembly code. Biochem J 474:427–443. https://doi.org/10.1042/bcj20160675
Sherwood PJ, Stafford WF (2016) Sedanal: model-dependent and model-independent analysis of sedimentation data. In: Uchiyama S, Arisaka F, Stafford WF, Laue T (eds) Analytical ultracentrifugation: instrumentation, software, and applications. Springer, Tokyo, pp 81–102. https://doi.org/10.1007/978-4-431-55985-6_6
Solomon OF, Ciutǎ IZ (1962) Détermination de la viscosité intrinsèque de solutions de polymères par une simple détermination de la viscosité. J Appl Polym Sci 6:683–686. https://doi.org/10.1002/app.1962.070062414
Stafford WF (1992) Boundary analysis in sedimentation transport experiments: a procedure for obtaining sedimentation coefficient distributions using the time derivative of the concentration profile. Anal Biochem 203:295–301. https://doi.org/10.1016/0003-2697(92)90316-Y
Stafford WF, Braswell EH (2004) Sedimentation velocity, multi-speed method for analyzing polydisperse solutions. Biophys Chem 108:273–279. https://doi.org/10.1016/j.bpc.2003.10.027
Stafford WF, Sherwood PJ (2004) Analysis of heterologous interacting systems by sedimentation velocity: curve fitting algorithms for estimation of sedimentation coefficients, equilibrium and kinetic constants. Biophys Chem 108:231–243. https://doi.org/10.1016/j.bpc.2003.10.028
Stafford WF (2016) Analysis of nonideal, interacting, and noninteracting systems by sedimentation velocity analytical ultracentrifugation. In: Uchiyama S, Arisaka F, Stafford WF, Laue T (eds) Analytical ultracentrifugation: instrumentation, software, and applications. Springer, Tokyo, pp 463–482. https://doi.org/10.1007/978-4-431-55985-6_23
Svedberg T, Fåhraeus R (1926) A new method for the determination of the molecular weight of the proteins. J Am Chem Soc 48:430–438. https://doi.org/10.1021/ja01413a019
Svedberg T, Lewis NB (1928) The molecular weights of phycoerythrin and of phycocyan. J Am Chem Soc 50:525–536. https://doi.org/10.1021/ja01389a042
Svedberg T, Nichols JB (1926) The molecular weight of egg albumin I. In electrolyte-free condition. J Am Chem Soc 48:3081–3092. https://doi.org/10.1021/ja01691a008
Svedberg T, Nichols JB (1927) The application of the oil turbine type of ultracentrifuge to the study of the stability region of carbon monoxide-hemoglobin. J Am Chem Soc 49:2920–2934. https://doi.org/10.1021/ja01410a046
Vinograd J, Bruner R, Kent R, Weigle J (1963) Band-centrifugation of macromolecules and viruses in self-generating density gradients. Proc Natl Acad Sci USA 49:902–910. https://doi.org/10.1073/pnas.49.6.902
Winzor DJ, Dinu V, Scott DJ, Harding SE (2021) Quantifying the concentration dependence of sedimentation coefficients for globular macromolecules: a continuing age-old problem. Biophys Rev 13:273–288. https://doi.org/10.1007/s12551-021-00793-x
Wright RT, Hayes D, Sherwood PJ, Stafford WF, Correia JJ (2018) AUC measurements of diffusion coefficients of monoclonal antibodies in the presence of human serum proteins. Eur Biophys J 47:709–722. https://doi.org/10.1007/s00249-018-1319-x
Yarawsky AE, Herr AB (2020) The staphylococcal biofilm protein Aap forms a tetrameric species as a necessary intermediate before amyloidogenesis. J Biol Chem 295:12840–12850. https://doi.org/10.1074/jbc.RA120.013936
Yarawsky AE, Johns SL, Schuck P, Herr AB (2020) The biofilm adhesion protein Aap from staphylococcus epidermidis forms zinc-dependent amyloid fibers. J Biol Chem. https://doi.org/10.1074/jbc.RA119.010874
Yarawsky AE, Hopkins JB, Chatzimagas L, Hub JS, Herr AB (2022) Solution structural studies of pre-amyloid oligomer states of the biofilm protein Aap. J Mol Biol 434:167708. https://doi.org/10.1016/j.jmb.2022.167708
Funding
Work was performed using National Institutes of Health funding from the National Institute of General Medical Sciences (R01-GM094363) awarded to A.B.H.
Author information
Authors and Affiliations
Contributions
AEY conceived the project, designed experiments, performed experiments, analyzed data, and wrote the manuscript. VD designed experiments, performed experiments, and analyzed data. SEH guided analysis and reviewed the manuscript. ABH designed experiments and provided funding. All authors reviewed the manuscript and contributed significantly to the final product.
Corresponding author
Ethics declarations
Conflict of interest
A.B.H. serves as a Scientific Advisory Board member for Hoth Therapeutics, Inc., holds equity in Hoth Therapeutics and Chelexa BioSciences, LLC, and was a co-inventor on seven patents broadly related to proteins described in this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Special Issue: Analytical Ultracentrifugation 2022.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yarawsky, A.E., Dinu, V., Harding, S.E. et al. Strong non-ideality effects at low protein concentrations: considerations for elongated proteins. Eur Biophys J 52, 427–438 (2023). https://doi.org/10.1007/s00249-023-01648-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00249-023-01648-x