Abstract
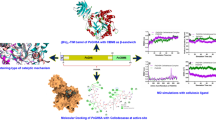
The stability of Debaryomyces nepalensis NCYC 3413 xylose reductase, a homodimeric enzyme recombinantly expressed and purified from E. coli Rosetta cells, was studied at different pH ranging from 5.0 to 10.0. Deactivation kinetics at different pH were studied by analyzing residual activity of the recombinant enzyme over time at 40 °C whereas conformational changes and stability dependence were investigated by using circular dichroism and differential scanning calorimetry. Four osmolytes viz. glycerol, sucrose, trehalose and sorbitol were explored for their effect on the deactivation and melting temperatures of the enzyme under neutral and extreme pH conditions. The enzyme was found to be catalytically and structurally stable at pH 7.0 with half-life of 250 min and a melting temperature of 50 °C. It was found that alteration in both secondary and tertiary structures caused enzyme deactivation in acidic pH while increased deactivation rates at alkaline pH was attributed to the variation of tertiary structure over time. Estimated thermodynamic parameters also showed that the enzyme stability was highest at neutral pH with ΔH of 348 kcal/mole and ΔG40 of 9.53 kcal/mole. All four osmolytes were effective in enhancing enzyme stability by several folds at extreme pH with sorbitol being the most efficient, which increased enzyme half-life by 11-fold at pH 10.0 and 8-fold at pH 5.0.







Similar content being viewed by others
References
Ahmad E, Fatima S, Khan MM, Khan RH (2010) More stable structure of wheat germ lipase at low pH than its native state. Biochimie 92:885–893. https://doi.org/10.1016/j.biochi.2010.03.023
Balejčíková L, Garamus VM, Avdeev MV, Petrenko VI, Almásy L, Kopčanský P (2017) The effect of solution pH on the structural stability of magnetoferritin. Colloids Surfaces B Biointerfaces 156:375–381. https://doi.org/10.1016/j.colsurfb.2017.05.036
Barroca M, Santos G, Johansson B, Gillotin F, Feller G, Collins T (2017) Deciphering the factors defining the pH-dependence of a commercial glycoside hydrolase family 8 enzyme. Enzyme Microb Technol 96:163–169. https://doi.org/10.1016/j.enzmictec.2016.10.011
Chen G, Zhang Q, Lu Q, Feng B (2019) Protection effect of polyols on Rhizopus chinensis lipase counteracting the deactivation from high pressure and high temperature treatment. Int J Biol Macromol 127:555–562. https://doi.org/10.1016/j.ijbiomac.2019.01.082
De Freitas BR, Dos Santos JC, Da Silva SS (2011) A novel use for sugarcane bagasse hemicellulosic fraction: Xylitol enzymatic production. Biomass Bioenerg 35:3241–3246. https://doi.org/10.1016/j.biombioe.2011.02.014
Delgado Arcaño Y, Valmaña García OD, Mandelli D, Carvalho WA, Magalhães Pontes LA (2018) Xylitol: a review on the progress and challenges of its production by chemical route. Catal Today. https://doi.org/10.1016/j.cattod.2018.07.060
Drago GA, Gibson TD (2001) Enzyme stability and stabilisation: applications and case studies. Eng Manuf Biotechnol. https://doi.org/10.1007/0-306-46889-1_24
Gill P, Moghadam TT, Ranjbar B (2010) Differential scanning calorimetry techniques: applications in biology and nanoscience. J Biomol Tech 21:167–193
Huisman GW, Liang J, Krebber A (2010) Practical chiral alcohol manufacture using ketoreductases. Curr Opin Chem Biol 14:122–129. https://doi.org/10.1016/j.cbpa.2009.12.003
Kosjek B, Nti-gyabaah J, Telari K, Dunne L, Moore JC (2008) Preparative asymmetric synthesis of 4,4-dimethoxytetrahydro-2H-pyran-3-ol with a ketone reductase and in situ cofactor recycling using glucose dehydrogenase. Org Process Res Dev 12:584–588
Kumar S, Gummadi SN (2011) Purification and biochemical characterization of a moderately halotolerant NADPH dependent xylose reductase from Debaryomyces nepalensis NCYC 3413. Bioresour Technol 102:9710–9717. https://doi.org/10.1016/j.biortech.2011.07.030
Louis-Jeune C, Andrade-Navarro MA, Perez-Iratxeta C (2012) Prediction of protein secondary structure from circular dichroism using theoretically derived spectra. Proteins Struct Funct Bioinform 80:374–381. https://doi.org/10.1002/prot.23188
Malla S, Gummadi SN (2018) Thermal stability of xylose reductase from Debaryomyces nepalensis NCYC 3413: deactivation kinetics and structural studies. Process Biochem 67:71–79. https://doi.org/10.1016/j.procbio.2018.01.010
Miyawaki O, Dozen M, Hirota K (2016) Cooperative hydration effect causes thermal unfolding of proteins and water activity plays a key role in protein stability in solutions. J Biosci Bioeng 122:203–207. https://doi.org/10.1016/j.jbiosc.2016.01.005
Neuhauser W, Steininger M, Haltrich D, Kulbe KD, Nidetzky B (1998) A pH-controlled fed-batch process can overcome inhibition by formate in NADH-dependent enzymatic reductions using formate dehydrogenase-catalyzed coenzyme regeneration. Biotechnol Bioeng 60:277–282
Nidetzky B, Neuhauser W, Haltrich D, Kulbe KD (1996) Continuous enzymatic production of xylitol with simultaneous coenzyme regeneration in a charged membrane reactor. Biotechnol Bioeng 52:387–396. https://doi.org/10.1002/(SICI)1097-0290(19961105)52:3%3c387:AID-BIT4%3e3.0.CO;2-G
Nirmal NP, Laxman RS (2014) Enhanced thermostability of a fungal alkaline protease by different additives. Enzyme Res. https://doi.org/10.1155/2014/109303
Ohtake S, Kita Y, Arakawa T (2011) Interactions of formulation excipients with proteins in solution and in the dried state. Adv Drug Deliv Rev 63:1053–1073. https://doi.org/10.1016/j.addr.2011.06.011
Paidimuddala B, Krishna Aradhyam G, Gummadi SN (2017a) A halotolerant aldose reductase from Debaryomyces nepalensis: gene isolation, overexpression and biochemical characterization. RSC Adv 7:20384–20393. https://doi.org/10.1039/C7RA01697B
Paidimuddala B, Rathod A, Gummadi SN (2017b) Inhibition of Debaryomyces nepalensis xylose reductase by lignocellulose derived by-products. Biochem Eng J 121:73–82. https://doi.org/10.1016/j.bej.2017.01.019
Paidimuddala B, Mohapatra SB, Gummadi SN, Manoj N (2018) Crystal structure of yeast xylose reductase in complex with a novel NADP-DTT adduct provides insights into substrate recognition and catalysis. FEBS J. https://doi.org/10.1111/febs.14667
Rafiqul ISM, Sakinah AMM, Zularisam AW (2015) Evaluation of sawdust hemicellulosic hydrolysate for bioproduction of xylitol by enzyme xylose reductase. Food Bioprod Process 94:82–89. https://doi.org/10.1016/j.fbp.2015.01.005
Sadana A (1995) Biocatalysis: fundamentals of enzyme deactvation kinetics. Prentice-Hall, Inc., Englewood Cliffs
Sanli G, Dudley JI, Blaber M (2003) Structural biology of the Aldo-Keto reductase. Cell Biochem Biophys 38:79–101
Singh R, Kumar M, Mittal A, Mehta PK (2016) Microbial enzymes: industrial progress in 21st century. 3Biotech 6:1–15. https://doi.org/10.1007/s13205-016-0485-8
Smith PK, Krohn RI, Hermanson GT, Mallia AK, Gartner FH, Provenzano MD, Fujimoto EK, Goeke NM, Olson BJ, Klenk DC (1985) Measurement of protein using bicinchoninic acid. Anal Chem 150:76–85
Talley K, Emil A (2011) On the pH-optimum of activity and stability of proteins. Proteins 78:2699–2706. https://doi.org/10.1002/prot.22786
Woodyer R, Simurdiak M, Van Der Donk W, Zhao H (2005) Heterologous expression, purification, and characterization of a highly active Xylose reductase from Neurospora crassa. Society 71:1642–1647. https://doi.org/10.1128/AEM.71.3.1642-1647.2005
Wright TA, Stewart JM, Page RC, Konkolewicz D (2017) Extraction of thermodynamic parameters of protein unfolding using parallelized differential scanning fluorimetry. J Phys Chem Lett 8:553–558. https://doi.org/10.1021/acs.jpclett.6b02894
Acknowledgements
Authors acknowledge Department of Biotechnology, Government of India for funding, and IIT Madras for the research facilities as well as DST-FIST facility for CD and DSC instruments. SM acknowledges MHRD, Government of India for the fellowship. Authors acknowledge Dr. Swati Dash for her kind suggestions in improving manuscript.
Funding
No specific grant was received for the present work from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
SNG devised and designed the studies and SM performed the experiments. SNG framed the outline of manuscript and suggested corrections whereas SM prepared the manuscript. Both the authors read and agreed the complete manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Malla, S., Gummadi, S.N. Counteraction of osmolytes on pH-induced unfolding of xylose reductase from Debaryomyces nepalensis NCYC 3413. Eur Biophys J 49, 267–277 (2020). https://doi.org/10.1007/s00249-020-01432-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00249-020-01432-1