Abstract
We have studied two different β-peptides in methanol using explicit solvent molecular dynamics simulations and the GROMOS 53A6 force field: a heptapeptide (peptide 1) expected to form a left-handed 314-helix, and a hexapeptide (peptide 2) expected to form a β-hairpin in solution. Our analysis has focused on identifying and analyzing the stability of the dominant secondary structure conformations adopted by the peptides, as well as on comparing the experimental NOE distance upper bounds and 3J-coupling values with their counterparts calculated on the basis of the simulated ensembles. Moreover, we have critically compared the present results with the analogous results obtained with the GROMOS 45A3 (peptide 1) and 43A1 (peptide 2) force fields. We conclude that within the limits of conformational sampling employed here, the GROMOS 53A6 force field satisfactorily reproduces experimental findings regarding the behavior of short β-peptides, with accuracy that is comparable to but not exceeding that of the previous versions of the force field.

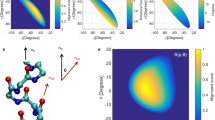
Conformational clustering analysis of the simulated ensemble of a ß-hexapeptide with two different simulation setups (a and b). The central members of all of the clusters populating more than 5% of all of the structures are shown, together with the most dominant hydrogen bonds and the corresponding percentages of cluster members containing them








Similar content being viewed by others
References
Appella DH, Christianson LA, Karle IL, Powell DR, Gellman SH (1997) Polypeptides of cyclic beta-amino acids that form stable helices. Abstr Pap Am Chem Soc 213:328-ORGN
Appella DH, Christianson LA, Karle IL, Powell DR, Gellman SH (1999) Synthesis and characterization of trans-2-aminocyclohexanecarboxylic acid oligomers: an unnatural helical secondary structure and implications for beta-peptide tertiary structure. J Am Chem Soc 121:6206–6212
Berendsen HJC, Postma JP, van Gunsteren WF, DiNola A, Haak JR (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81:3684–3690
Cheng RP (2004) Beyond de novo protein design—de novo design of non-natural folded oligomers. Curr Opin Struct Biol 14:512–520
Cheng RP, Gellman SH, DeGrado WF (2001) Beta-peptides: from structure to function. Chem Rev 101:3219–3232
Cubberley MS, Iverson BL (2001) Models of higher-order structure: foldamers and beyond. Curr Opin Chem Biol 5:650–653
Daura X, van Gunsteren WF, Mark AE (1999) Folding–unfolding thermodynamics of a beta-heptapeptide from equilibrium simulations. Proteins 34:269–280
Daura X, Gademann K, Schäfer H, Jaun B, Seebach D, van Gunsteren WF (2001) The beta-peptide hairpin in solution: conformational study of a beta-hexapeptide in methanol by NMR spectroscopy and MD simulation. J Am Chem Soc 123:2393–2404
DeMarco A, Llinas M, Wüthrich K (1978) H-1-N-15 spin–spin couplings in alumichrome. Biopolymers 17:2727–2742
Glättli A, Daura X, Bindschädler P, Jaun B, Mahajan YR, Mathad RI, Rueping M, Seebach D, van Gunsteren WF (2005) On the influence of charged side chains on the folding–unfolding equilibrium of beta-peptides: a molecular dynamics simulation study. Chemistry 11:7276–7293
Hecht E, Huc I (eds) 2007 Foldamers: structure, properties, and applications. Wiley-VCH, New York
Hintermann T, Seebach D (1997) The biological stability of beta-peptides: no interactions between alpha- and beta-peptidic structures. Chimia 51:244–247
Karplus M (1959) Contact electron-spin coupling of nuclear magnetic moments. J Chem Phys 30:11–15
Oostenbrink C, Villa A, Mark AE, van Gunsteren WF (2004) A biomolecular force field based on the free enthalpy of hydration and solvation: the GROMOS force-field parameter sets 53A5 and 53A6. J Comput Chem 25:1656–1676
Oostenbrink C, Soares TA, van der Vegt NF, van Gunsteren WF (2005) Validation of the 53A6 GROMOS force field. Eur Biophys J 34:273–284
Pardi A, Billeter M, Wüthrich K (1984) Calibration of the angular dependence of the amide proton-C alpha proton coupling constants, 3JHNα, in a globular protein. Use of 3JHNα for identification of helical secondary structure. J Mol Biol 180:741–751
Ryckaert JP, Ciccotti G, Berendsen HJC (1977) Numerical integration of Cartesian equations of motion of a system with constraints—molecular dynamics of n-alkanes. J Comput Phys 23:327–341
Scott WRP, Hünenberger PH, Tironi IG, Mark AE, Billeter SR, Torda AE, Huber T, Krüger P, van Gunsteren WF (1999) The GROMOS biomolecular simulation program package. J Phys Chem A 103:3596–3607
Seebach D, Matthews JL (1997) Beta-peptides: a surprise at every turn. Chem Comm21:2015–2022
Seebach D, Abele S, Schreiber JV, Martinoni B, Nussbaum AK, Schild H, Schulz H, Hennecke H, Woessner R, Bitsch F (1998) Biological and pharmacokinetic studies with beta-peptides. Chimia 52:734–739
Schuler LD, Daura X, Van Gunsteren WF (2001) An improved GROMOS96 force field for aliphatic hydrocarbons in the condensed phase. J Comput Chem 22:1205–1218
van Gunsteren WF, Billeter SR, EisingAA, Hünenberger PH, Krüger P, Mark AE, Scott WRP, Tironi IG (1996) Biomolecular simulation: the GROMOS96 manual and user guide. Biomos, Zurich, Groningen
Acknowledgments
B.Z acknowledges support from an EMBO postdoctoral fellowship. This work was financially supported by grants from the National Center of Competence in Research (NCCR) in Structural Biology of the Swiss National Science Foundation, which is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zagrovic, B., Gattin, Z., Lau, J.KC. et al. Structure and dynamics of two β-peptides in solution from molecular dynamics simulations validated against experiment. Eur Biophys J 37, 903–912 (2008). https://doi.org/10.1007/s00249-008-0307-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00249-008-0307-y