Abstract
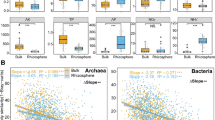
Numerous rare species coexist with a few abundant species in microbial communities and together play an essential role in riparian ecosystems. Relatively little is understood, however, about the nature of assembly processes of these communities and how they respond to a fluctuating environment. In this study, drivers controlling the assembly of abundant and rare subcommunities for bacteria and archaea in a riparian zone were determined, and their resulting patterns on these processes were analyzed. Abundant and rare bacteria and archaea showed a consistent variation in the community structure along the riparian elevation gradient, which was closely associated with flooding frequency. The community assembly of abundant bacteria was not affected by any measured environmental variables, while soil moisture and ratio of submerged time to exposed time were the two most decisive factors determining rare bacterial community. Assembly of abundant archaeal community was also determined by these two factors, whereas rare archaea was significantly associated with soil carbon–nitrogen ratio and total carbon content. The assembly process of abundant and rare bacterial subcommunities was driven respectively by dispersal limitation and variable selection. Undominated processes and dispersal limitation dominated the assembly of abundant archaea, whereas homogeneous selection primarily driven rare archaea. Flooding may therefore play a crucial role in determining the community assembly processes by imposing disturbances and shaping soil niches. Overall, this study reveals the assembly patterns of abundant and rare communities in the riparian zone and provides further insight into the importance of their respective roles in maintaining a stable ecosystem during times of environmental perturbations.





Similar content being viewed by others
Data Availability
Sequences were deposited in the National Center for Biotechnology Information Sequence Read Archive (https://submit.ncbi.nlm.nih.gov/subs/sra/) under accession number PRJNA863415 for bacteria, and PRJNA394057 for archaea.
References
Bao Y, Gao P, He X (2015) The water-level fluctuation zone of Three Gorges Reservoir — a unique geomorphological unit. Earth-Sci Rev 150:14–24. https://doi.org/10.1016/j.earscirev.2015.07.005
Wang S, Wang X, Jiang Y, Han C, Jetten MSM, Schwark L, Zhu G (2021) Abundance and functional importance of complete ammonia oxidizers and other nitrifiers in a riparian ecosystem. Environ Sci Technol 55:4573–4584. https://doi.org/10.1021/acs.est.0c00915
Bo Y, Jägermeyr J, Yin Z, Jiang Y, Xu J, Liang H, Zhou F (2022) Global benefits of non-continuous flooding to reduce greenhouse gases and irrigation water use without rice yield penalty. Glob Change Biol 28:3636–3650. https://doi.org/10.1111/gcb.16132
Kumaragamage D, Concepcion A, Gregory C, Goltz D, Indraratne S, Amarawansha G (2020) Temperature and freezing effects on phosphorus release from soils to overlying floodwater under flooded-anaerobic conditions. J Environ Qual 49:700–711. https://doi.org/10.1002/jeq2.20062
Ran Y, Ma M, Liu Y, Zhou Y, Sun X, Wu S, Huang P (2021) Hydrological stress regimes regulate effects of binding agents on soil aggregate stability in the riparian zones. CATENA 196:104815. https://doi.org/10.1016/j.catena.2020.104815
Ye F, Ma M, Wu S, Jiang Y, Zhu G, Zhang H, Wang Y (2019) Soil properties and distribution in the riparian zone: the effects of fluctuations in water and anthropogenic disturbances. Eur J Soil Sci 70:664–673. https://doi.org/10.1111/ejss.12756
Zhang X, Zhao X, Zhang M (2012) Functional diversity changes of microbial communities along a soil aquifer for reclaimed water recharge. FEMS Microbiol Ecol 80:9–18. https://doi.org/10.1111/j.1574-6941.2011.01263.x
Pedrós-Alió C (2012) The rare bacterial biosphere. Annu Rev Mar Sci 4:449–466. https://doi.org/10.1146/annurev-marine-120710-100948
Liu L, Yang J, Yu Z, Wilkinson DM (2015) The biogeography of abundant and rare bacterioplankton in the lakes and reservoirs of China. ISME J 9:2068–2077. https://doi.org/10.1038/ismej.2015.29
Jia X, Dini-Andreote F, Falcão Salles J (2018) Community assembly processes of the microbial rare biosphere. Trends Microbiol 26:738–747. https://doi.org/10.1016/j.tim.2018.02.011
Shade A, Jones SE, Caporaso JG, Handelsman J, Knight R, Fierer N, Gilbert JA (2014) Conditionally rare taxa disproportionately contribute to temporal changes in microbial diversity. mBio 5:e01371-01314. https://doi.org/10.1128/mBio.01371-14
Lynch MDJ, Neufeld JD (2015) Ecology and exploration of the rare biosphere. Nat Rev Microbiol 13:217–229. https://doi.org/10.1038/nrmicro3400
Jousset A, Bienhold C, Chatzinotas A, Gallien L, Gobet A, Kurm V, Küsel K, Rillig MC, Rivett DW, Salles JF, van der Heijden MGA, Youssef NH, Zhang X, Wei Z, Hol WHG (2017) Where less may be more: How the rare biosphere pulls ecosystems strings. ISME J 11:853–862. https://doi.org/10.1038/ismej.2016.174
Pester M, Bittner N, Deevong P, Wagner M, Loy A (2010) A ‘rare biosphere’ microorganism contributes to sulfate reduction in a peatland. ISME J 4:1591–1602. https://doi.org/10.1038/ismej.2010.75
Hua Z, Han Y, Chen L, Liu J, Hu M, Li S, Kuang J, Chain PSG, Huang L, Shu W (2015) Ecological roles of dominant and rare prokaryotes in acid mine drainage revealed by metagenomics and metatranscriptomics. ISME J 9:1280–1294. https://doi.org/10.1038/ismej.2014.212
Tao R, Zhao X, Zhang H, Hu B, Li J, Chu G (2022) Response and recovery of nosZ abundant and rare subcommunities to organic amendment and nitrification inhibitor disturbances, with implications for N2O emissions. Appl Soil Ecol 173:104386. https://doi.org/10.1016/j.apsoil.2022.104386
Wu W, Logares R, Huang B, Hsieh C-h (2017) Abundant and rare picoeukaryotic sub-communities present contrasting patterns in the epipelagic waters of marginal seas in the northwestern Pacific Ocean. Environ Microbiol 19:287–300. https://doi.org/10.1111/1462-2920.13606
Jiao S, Lu Y (2020) Abundant fungi adapt to broader environmental gradients than rare fungi in agricultural fields. Glob Change Biol 26:4506–4520. https://doi.org/10.1111/gcb.15130
Mo Y, Zhang W, Yang J, Lin Y, Yu Z, Lin S (2018) Biogeographic patterns of abundant and rare bacterioplankton in three subtropical bays resulting from selective and neutral processes. ISME J 12:2198–2210. https://doi.org/10.1038/s41396-018-0153-6
Stegen JC, Lin X, Fredrickson JK, Chen X, Kennedy DW, Murray CJ, Rockhold ML, Konopka A (2013) Quantifying community assembly processes and identifying features that impose them. ISME J 7:2069–2079. https://doi.org/10.1038/ismej.2013.93
Wan W, Grossart HP, He D, Yuan W, Yang Y (2021) Stronger environmental adaptation of rare rather than abundant bacterioplankton in response to dredging in eutrophic Lake Nanhu (Wuhan, China). Water Res 190:116751. https://doi.org/10.1016/j.watres.2020.116751
Stegen JC, Lin X, Fredrickson JK, Konopka AE (2015) Estimating and mapping ecological processes influencing microbial community assembly. Front Microbiol 6:370. https://doi.org/10.3389/fmicb.2015.00370
Zhao Z, Ma Y, Feng T, Kong X, Wang Z, Zheng W, Zhai B (2022) Assembly processes of abundant and rare microbial communities in orchard soil under a cover crop at different periods. Geoderma 406:115543. https://doi.org/10.1016/j.geoderma.2021.115543
Gao GF, Peng D, Tripathi BM, Zhang Y, Chu H (2020) Distinct community assembly processes of abundant and rare soil bacteria in coastal wetlands along an inundation gradient. mSystems 5:e01150-01120. https://doi.org/10.1128/mSystems.01150-20
Jiao S, Chen W, Wei G (2017) Biogeography and ecological diversity patterns of rare and abundant bacteria in oil-contaminated soils. Mol Ecol 26:5305–5317. https://doi.org/10.1111/mec.14218
Ji M, Kong W, Stegen J, Yue L, Wang F, Dong X, Cowan DA, Ferrari BC (2020) Distinct assembly mechanisms underlie similar biogeographical patterns of rare and abundant bacteria in Tibetan Plateau grassland soils. Environ Microbiol 22:2261–2272. https://doi.org/10.1111/1462-2920.14993
Jiao S, Lu Y (2020) Soil pH and temperature regulate assembly processes of abundant and rare bacterial communities in agricultural ecosystems. Environ Microbiol 22:1052–1065. https://doi.org/10.1111/1462-2920.14815
Dini-Andreote F, Stegen JC, van Elsas JD, Salles JF (2015) Disentangling mechanisms that mediate the balance between stochastic and deterministic processes in microbial succession. P Natl Acad Sci USA 112:E1326-1332. https://doi.org/10.1073/pnas.1414261112
Ye F, Wang X, Wang Y, Wu S, Wu J, Hong Y (2021) Different pioneer plant species have similar rhizosphere microbial communities. Plant Soil 464:165–181. https://doi.org/10.1007/s11104-021-04952-7
Ye F, Wu S, Jiang Y, Camp HJMOd, Li Z, Zhu G, Zheng J, Wang Y (2016) Shifts of archaeal community structure in soil along an elevation gradient in a reservoir water level fluctuation zone. J Soil Sediment 16:2728–2739. https://doi.org/10.1007/s11368-016-1485-3
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Logares R, Audic S, Bass D, Bittner L, Boutte C, Christen R, Claverie JM, Decelle J, Dolan JR, Dunthorn M, Edvardsen B, Gobet A, Kooistra WHCF, Mahé F, Not F, Ogata H, Pawlowski J, Pernice MC, Romac S, Shalchian-Tabrizi K, Simon N, Stoeck T, Santini S, Siano R, Wincker P, Zingone A, Richards TA, de Vargas C, Massana R (2014) Patterns of rare and abundant marine microbial eukaryotes. Curr Biol 24:813–821. https://doi.org/10.1016/j.cub.2014.02.050
Dai T, Zhang Y, Tang Y, Bai Y, Tao Y, Huang B, Wen D (2017) Identifying the key taxonomic categories that characterize microbial community diversity using full-scale classification: a case study of microbial communities in the sediments of Hangzhou Bay. FEMS Microbiol Ecol 93:fiw203. https://doi.org/10.1093/femsec/fiw203
Wang Y, Ye F, Wu S, Wu J, Yan J, Xu K, Hong Y (2020) Biogeographic pattern of bacterioplanktonic community and potential function in the Yangtze River: roles of abundant and rare taxa. Sci Total Environ 747:141335. https://doi.org/10.1016/j.scitotenv.2020.141335
Xue Y, Chen H, Yang JR, Liu M, Huang B, Yang J (2018) Distinct patterns and processes of abundant and rare eukaryotic plankton communities following a reservoir cyanobacterial bloom. ISME J 12:2263–2277. https://doi.org/10.1038/s41396-018-0159-0
Ter Braak CJF, Šmilauer P (2012) Canoco reference manual and user's guide: software of ordination, version 5.0. Microcomputer Power, Ithaca, USA
Gaston KJ, Blackburn TM, Greenwood JJD, Gregory RD, Quinn RM, Lawton JH (2000) Abundance–occupancy relationships. J Appl Ecol 37:39–59. https://doi.org/10.1046/j.1365-2664.2000.00485.x
Ren Q, Yuan J, Wang J, Liu X, Ma S, Zhou L, Miao L, Zhang J (2022) Water level has higher influence on soil organic carbon and microbial community in Poyang Lake wetland than vegetation type. Microorganisms 10:131. https://doi.org/10.3390/microorganisms10010131
Chen L, Li C, Feng Q, Wei Y, Zheng H, Zhao Y, Feng Y, Li H (2017) Shifts in soil microbial metabolic activities and community structures along a salinity gradient of irrigation water in a typical arid region of China. Sci Total Environ 598:64–70. https://doi.org/10.1016/j.scitotenv.2017.04.105
Seaton FM, Jones DL, Creer S, George PBL, Smart SM, Lebron I, Barrett G, Emmett BA, Robinson DA (2019) Plant and soil communities are associated with the response of soil water repellency to environmental stress. Sci Total Environ 687:929–938. https://doi.org/10.1016/j.scitotenv.2019.06.052
Campbell BJ, Yu L, Heidelberg JF, Kirchman DL (2011) Activity of abundant and rare bacteria in a coastal ocean. P Natl Acad Sci USA 108:12776–12781. https://doi.org/10.1073/pnas.1101405108
Dimitriu PA, Lee D, Grayston SJ (2010) An evaluation of the functional significance of peat microorganisms using a reciprocal transplant approach. Soil Biol Biochem 42:65–71. https://doi.org/10.1016/j.soilbio.2009.10.001
Morrissey EM, Mau RL, Hayer M, Liu XA, Schwartz E, Dijkstra P, Koch BJ, Allen K, Blazewicz SJ, Hofmockel K, Pett-Ridge J, Hungate BA (2019) Evolutionary history constrains microbial traits across environmental variation. Nat Ecol Evol 3:1064–1069. https://doi.org/10.1038/s41559-019-0918-y
Jia X, Dini-Andreote F, Salles JF (2022) Unravelling the interplay of ecological processes structuring the bacterial rare biosphere. Isme Commun 2:96. https://doi.org/10.1038/s43705-022-00177-6
Powell JR, Karunaratne S, Campbell CD, Yao H, Robinson L, Singh BK (2015) Deterministic processes vary during community assembly for ecologically dissimilar taxa. Nat Commun 6:8444. https://doi.org/10.1038/ncomms9444
Li M, Mi T, He H, Chen Y, Zhen Y, Yu Z (2021) Active bacterial and archaeal communities in coastal sediments: biogeography pattern, assembly process and co-occurrence relationship. Sci Total Environ 750:142252. https://doi.org/10.1016/j.scitotenv.2020.142252
Walters KE, Capocchi JK, Albright MBN, Hao Z, Brodie EL, Martiny JBH (2022) Routes and rates of bacterial dispersal impact surface soil microbiome composition and functioning. ISME J: in press https://doi.org/10.1038/s41396-022-01269-w
McCoy EL, Hagedorn C (1979) Quantitatively tracing bacterial transport in saturated soil systems. Water Air Soil Pollut 11:467–479. https://doi.org/10.1007/BF00283438
Pita R, Mira A, Beja P (2011) Assessing habitat differentiation between coexisting species: the role of spatial scale. Acta Oecol 37:124–132. https://doi.org/10.1016/j.actao.2011.01.006
Calcagno V, Mouquet N, Jarne P, David P (2006) Coexistence in a metacommunity: the competition–colonization trade-off is not dead. Ecol Lett 9:897–907. https://doi.org/10.1111/j.1461-0248.2006.00930.x
Feng Y, Chen R, Stegen JC, Guo Z, Zhang J, Li Z, Lin X (2018) Two key features influencing community assembly processes at regional scale: initial state and degree of change in environmental conditions. Mol Ecol 27:5238–5251. https://doi.org/10.1111/mec.14914
Zhu G, Wang S, Wang W, Wang Y, Zhou L, Jiang B, Op den Camp HJM, Risgaard-Petersen N, Schwark L, Peng Y, Hefting MM, Jetten MSM, Yin C (2013) Hotspots of anaerobic ammonium oxidation at land–freshwater interfaces. Nat Geosci 6:103–107. https://doi.org/10.1038/ngeo1683
Miltner A, Bombach P, Schmidt-Brücken B, Kästner M (2012) SOM genesis: microbial biomass as a significant source. Biogeochem 111:41–55. https://doi.org/10.1007/s10533-011-9658-z
Lennon JT, Jones SE (2011) Microbial seed banks: The ecological and evolutionary implications of dormancy. Nat Rev Microbiol 9:119–130. https://doi.org/10.1038/nrmicro2504
Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (grant numbers 51908145 and 41977153), the Natural Science Foundation of Guangdong Province (grant number 2022A1515010828), and Science and Technology Projects in Guangzhou (grant number 202201020580).
Author information
Authors and Affiliations
Contributions
YW, FY, and YH designed the study. FY and YW performed the field work and laboratory work. FY, ZS, and YW analyzed the data. FY and YW wrote the manuscript. SM, JW, and YH contributed to the writing. All authors have reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ye, F., Sun, Z., Moore, S.S. et al. Discrepant Effects of Flooding on Assembly Processes of Abundant and Rare Communities in Riparian Soils. Microb Ecol 86, 1164–1175 (2023). https://doi.org/10.1007/s00248-022-02152-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-022-02152-z