Abstract
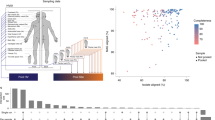
This review article on the skin microbiota was written in response to recent advances that transitioned from culture methods to PCR amplification and sequencing of bacterial and fungal genes as a result of the Human Microbiome Project. This transition enables the investigation of the full diversity of microorganisms inhabiting human skin. The skin provides a range of habitats with different microbiota associated with the three major regions of the skin, namely the moist axilla, perineum, and toe webs; oily or sebaceous head, neck, and trunk; and dry forearms and legs. These new culture-independent tools are revealing the diversity of the human skin microbiota in the different locations of the body and with skin depth. These tools should lead to a better understanding of the state of homeostasis between the microbiota and the host and the overall functionality of that microbiota.
Similar content being viewed by others
References
Noble WC (1981) Microbiology of human skin, 2nd edn. Lloyd-Luke, London, p 443
Hannigan GD, Grice EA (2013) Microbial ecology of skin in the era of metagenomics and molecular microbiology. Cold Spring Harb Perspect Med 3:a015362
Grice EA et al (2009) Topographical and temporal diversity of the human skin microbiome. Science 324:1190–1192
Grice EA, Segre JA (2011) The skin microbiota. Nat Res Microbiol 9:244–253
Fuchs E, Raghaven S (2002) Getting under the skin of epidermal morphogenesis. Nat Rev Genet 3(3):199–209
Segre JA (2006) Epidermal barrier formulation and recovery in skin disorders. J Clin Invest 116:1150–1158
Blanpain C, Horsley V, Fuchs E (2007) Epithelial stem cells: turning over new leaves. Cell 128:445–458
Somerville DA, Noble WC (1973) Micro-colony size of microbes on human skin. J Med Microbiol 6:323
Malcolm SA, Hughes TC (1980) The demonstration of bacteria on and within the stratum corneum using scanning electron microscopy. Br J Dermatol 192(3):267–275
Dominguez-Bello MG et al (2010) Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci U S A 107:11971–119775
Mueller NT et al (2015) The infant microbiome development: mom matters. Trends Mol Med 21(2):109–117
Turnbaugh PJ et al (2009) The Human Microbiome Project. Nature 449:804–810
Fredricks DN et al (2005) Molecular identification of bacteria associated with bacterial vaginosis. N Engl J Med 353:1899–1911
Marples MJ (1965) The ecology of the human skin. Springfield, Illinois Thomas
Marples MJ (1969) Life on the human skin. Sci Am 220:108–115
Savage DC (1977) Microbial ecology of the intestinal tract. Ann Rev Microbiol 31:107–133
Grice EA et al (2008) A diversity profile of the human skin microbiota. Genom Res 18:1043–1050
Gao Z et al (2007) Molecular analysis of the human forearm superficial skin bacterial biota. Proc Natl Acad Sci U S A 104(8):2927–2932
Zhou YJ et al (2013) Biogeography of the ecosystems of the healthy human body. Genom Biol 14:R1
Belkaid Y, Segre JA (2014) Dialogue between skin microbiota and immunity. Science 346(6212):954–959
Findley K, Grice EA (2014) The skin microbiome: a focus on pathogens and their association with skin disease. Plos Pathogens 10(11):31004436
Tan HH (2003) Antibacterial therapy for acne. Am J Clin Dermatol 4(5):307–314
Fitz-Gibbon S et al (2013) Propionibacterium acnes strain populations in the human skin microbiome associated with acne. J Invest Dermatol 133(9):2153–2160
Findley K et al (2013) Human skin fungal diversity. Nature 498(7454):367–370
Gao Z et al (2010) Quantification of major human cutaneous bacterial and fungal populations. J Clin Microbiol 48(10):3575–3581
Fierer N et al (2008) The influence of sex. handedness, and washing on the diversity of hand surface bacteria. Proc Natl Acad Sci U S A 105:17994–17999
Madison KC (2003) Barrier function of the skin: la raison d’être of the epidermis. J Invest Dermatol 121:231–241
Randhawa M et al (2014) Metabolic signature of sun-exposed skin suggests catabolic pathway overweighs anabolic pathway. Plos One 9(3):e90367
Zeeuwen P et al (2012) Microbiome dynamics of human epidermis following skin barrier disruption. Gen Bio 13(R101):1–18
Blaser MJ (2014) Missing microbes: how the overuse of antibiotics is fueling our modern plagues. Henry Holt, New York, p 273
Lemon KP et al (2012) Microbiota—target therapies: an ecological perspective. Sci Transl Med 4(137):1–8
Spinillo A et al (1999) Effect of antibiotic use on the prevalence of symptomatic vulvovaginal candidiasis. Am J Obstet Gynecol 180:14–17
Kelly CP, LaMont JT (2008) Clostridium difficile—more difficult than ever. N Engl J Med 358:1932–1940
Van Nood E et al (2013) Duodenal infusion of donor feces for recurrent Clostridium difficile. N Engl J Med 368(5):407–415
Varga M et al (2012) Efficient transfer of antibiotic resistance plasmids by transduction within methicillin-resistant Staphylococcus aureus USA300 clone. FEMS Microbiol Lett 332:146–152
Holland KT, Bojar RA (2002) Cosmetics: what is their influence on the skin microflora. Am J Clin Dermatol 3:445–449
Kong HH, Segre JA (2012) Skin microbiome: looking back to move forward. J Invest Dermatol 132(3):933–939
Kong HH et al (2012) Temporal shifts in the skin microbiome with disease flares and treatment in children with atopic dermatitis. Genome Res 22:850–859
Chen YE, Tsao H (2013) The skin microbiome: current perspectives and future challenges. J Am Acad Dermatol 69(1):143–155
Costello EK et al (2012) The application of ecological theory towards an understanding of the human microbiome. Science 336(6086):1255–1262
van Rensburg JJ, Lin H et al (2015) The human skin microbiome associates with the outcome of and is influenced by bacterial infection. mBio 6(5):e01315-15
Korte D, Marcelis JH (2014) Platelet concentrates: reducing the risk of transfusion-transmitted bacterial infections. Intern J Clin Trans Med 2:29–37
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cundell, A.M. Microbial Ecology of the Human Skin. Microb Ecol 76, 113–120 (2018). https://doi.org/10.1007/s00248-016-0789-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-016-0789-6