Abstract
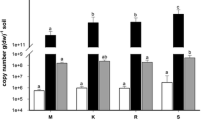
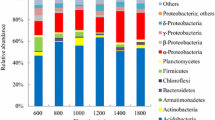
Unraveling the distribution patterns of plants and animals along the elevational gradients has been attracting growing scientific interests of ecologists, whether the microbial communities exhibit similar elevational patterns, however, remains largely less documented. Here, we investigate the biogeographic distribution of soil archaeal and bacterial communities across three vertical climate zones (3,106–4,479 m.a.s.l.) in Mt. Shegyla on the Tibetan Plateau, by combining quantitative PCR and high-throughput barcoded pyrosequencing approaches. Our results found that the ratio of bacterial to archaeal 16S rRNA gene abundance was negatively related with elevation. Acidobacteria dominated in the bacterial communities, Marine benthic group A dominated in the archaeal communities, and the relative abundance of both taxa changed significantly with elevation. At the taxonomic levels of domain, phylum, and class, more bacterial taxa than archaeal exhibited declining trend in diversity along the increasing elevational gradient, as revealed by Shannon and Faith’s phylogenetic diversity indices. Unweighted UniFrac distance clustering showed that the bacterial communities from the mountainous temperate zone clustered together, whereas those from the subalpine cool temperate zone clustered together. However, the partitioning effect of elevational zones on the archaeal community was much weaker compared to that on bacteria. Redundancy analysis revealed that soil geochemical factors explained 58.3 % of the bacterial community variance and 75.4 % of the archaeal community variance. Taken together, we provide evidence that soil bacteria exhibited more apparent elevational zonation feature and decreased diversity pattern than archaea with increasing elevation, and distribution patterns of soil microbes are strongly regulated by soil properties along elevational gradient in this plateau montane ecosystem.




Similar content being viewed by others
References
Körner C (2007) The use of ‘altitude’ in ecological research. Trends Ecol Evol 22:569–574
Gaston KJ (2000) Global patterns in biodiversity. Nature 405:220–227
Parmesan C, Gaines S, Gonzalez L, Kaufman DM, Kingsolver J, Peterson AT, Sagarin R (2005) Empirical perspectives on species borders: from traditional biogeography to global change. Oikos 108:58–75
Kessler M (2000) Altitudinal zonation of Andean cryptogam communities. J Biogeogr 27:275–282
Romdal TS, Rahbek C (2009) Elevational zonation of afrotropical forest bird communities along a homogeneous forest gradient. J Biogeogr 36:327–336
Fierer N, Jackson RB (2005) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci U S A 103:626–631
Alexander JM, Kueffer C, Daehler CC, Edwards PJ, Pauchard AB, Seipel T, MIREN Consortium (2010) Assembly of nonnative floras along elevational gradients explained by directional ecological filtering. Proc Natl Acad Sci U S A 108:656–661
Kozak KH, Wiens JJ (2010) Niche conservatism drives elevational diversity patterns in Appalachian salamanders. Amer Nat 176:40–54
Qiong L, Grytnes JA, Birks HJB (2010) Alpine vegetation and species-richness patterns along two altitudinal gradients in the Gyama Valley, south-central Tibet, China. Plant Ecol Divers 3:3,235–247
Bryant JA, Lamanna C, Morlon H, Kerkhoff AJ, Enquist BJ, Green JL (2008) Microbes on mountainsides: contrasting elevational patterns of bacterial and plant diversity. Proc Natl Acad Sci U S A 105:11505–11511
Fierer N, McCain CM, Meir P, Zimmermann M, Rapp JM, Silman MR, Knight R (2011) Microbes do not follow the elevational diversity patterns of plants and animals. Ecology 92:797–804
Singh D, Takahashi K, Kim M, Chun J, Adams JM (2012) A hump-backed trend in bacterial diversity with elevation on Mount Fuji, Japan. Microbial Ecol 63:429–437
Colwell RK, Lees DC (2000) The mid-domain effect: geometric constraints on the geography of species richness. Trends Ecol Evol 15:70–76
Yang J, Smith HG, Sherratt TN, Wilkinson DM (2010) Is there a size limit for cosmopolitan distribution in free-living microorganisms? A biogeographical analysis of testate amoebae from polar areas. Microbial Ecol 59:635–645
Wang JJ, Soininen J, Zhang Y, Wang BX, Yang XD, Shen J (2011) Contrasting patterns in elevational diversity between microorganisms and macroorganisms. J Biogeogr 38:595–603
Zhang LM, Wang M, Prosser JI, Zheng YM, He JZ (2009) Altitude ammonia-oxidizing bacteria and archaea in soils of Mount Everest. FEMS Microbiol Ecol 70:208–217
Hu HW, Zhang LM, Dai Y, Di HJ, He JZ (2013) pH-dependent distribution of soil ammonia oxidizers across a large geographical scale as revealed by high-throughput pyrosequencing. J Soil Sediment 13:1439–1449
He JZ, Hu HW, Zhang LM (2012) Current insights into the autotrophic thaumarchaeal ammonia oxidation in acidic soils. Soil Biol Biochem 55:146–154
Wu Z, Tang Y, Li X, Wu S, Li H (1981) Dissertations upon the origin, development and regionalization of Xizang flora through the floristic analysis. Proc Symp Qinghai-Xizang Plat 2:1219–1244
Guo P, Liu Q, Li C, Chen X, Jiang K, Wang YZ, Malhotra A (2011) Molecular phylogeography of Jerdon's pitviper (Protobothrops jerdonii): importance of the uplift of the Tibetan Plateau. J Biogeogr 38:2326–2336
Araújo MB, Whittaker RJ, Ladle RJ, Erhard M (2005) Reducing uncertainty in projections of extinction risk from climate change. Glob Ecol Biogeogr 14:529–538
Prosser JI, Bohannan BJM, Curtis TP, Ellis RJ, Firestone MK, Freckleton RP, Green JL, Green LE, Killham K, Lennon JJ, Osborn AM, Solan M, van der Gast CJ, Young JPW (2007) The role of ecological theory in microbial ecology. Nat Rev Microbiol 5:384–392
Cao P, Zhang LM, Shen JP, Zheng YM, Di HJ, He JZ (2012) Distribution and diversity of archaeal communities in selected Chinese soils. FEMS Microbiol Ecol 80:146–158
Suzuki MT, Taylor LT, DeLong EF (2000) Quantitative analysis of small-subunit rRNA genes in mixed microbial populations via 5′-nuclease assays. Appl Environ Microbiol 66:4605–4614
Kemnitz D, Kolb S, Conrad R (2005) Phenotypic characterization of Rice Cluster III archaea without prior isolation by applying quantitative polymerase chain reaction to an enrichment culture. Environ Microbiol 7:553–565
Lane D (1991) 16S/23S rRNA sequencing. Wiley, New York
Caporaso JG, Kuczynski J, Stombaugh J et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Caporaso JG, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R (2010) PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26:266–267
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Faith DP (1992) Conservation evaluation and phylogenetic diversity. Biol Cons 61:1–10
Lozupone CA, Hamady M, Kelley ST, Knight R (2007) Quantitative and qualitative β diversity measures lead to different insights into factors that structure microbial communities. Appl Environ Microbiol 73:1576–1585
R Core Team (2013). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL http://www.R-project.org/
Oksanen J, Kindt R, Legendre P, O’Hara B (2007) vegan: Community Ecology Package. R package version 2.0-6.Available at: http://cran.r-project.org/
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Austral Ecol 26:32–46
O'Malley MA (2007) The nineteenth century roots of ‘everything is everywhere’. Nat Rev Microbiol 5:647–651
Shen C, Xiong J, Zhang H, Feng Y, Lin X, Li X, Liang W, Chu H (2012) Soil pH drives the spatial distribution of bacterial communities along elevation on Changbai Mountain. Soil Biol Biochem 54:204–211
Yang Y, Gao Y, Wang S, Xu D, Yu H, Wu L, Lin Q, Hu Y, Li X, He Z, Deng Y, Zhou J (2013) The microbial gene diversity along an elevation gradient of the Tibetan grassland. ISME J 7:1–11
Kourtev PS, Ehrenfeld JG, Haggblom M (2003) Experimental analysis of the effect of exotic and native plant species on the structure and function of soil microbial communities. Soil Biol Biochem 35:895–905
Zheng YM, Cao P, Fu B, Hughes JM, He JZ (2013) Ecological drivers of biogeographic patterns of soil archaeal community. PLoS One 8:e63375
Kozak KH, Wiens JJ (2007) Climatic zonation drives latitudinal variation in speciation mechanisms. P Roy Soc B-Biol Sci 274:2995–3003
Jenny H (1941) Factors of soil formation: a system of quantitative pedology. McGraw-Hill, New York
Adair KL, Schwartz E (2008) Evidence that ammonia-oxidizing archaea are more abundant than ammonia-oxidizing bacteria in semiarid soils of northern Arizona, USA. Microbial Ecol 56:420–426
Singh D, Takahashi K, Adams JM (2013) Elevational patterns in archaeal diversity on Mt. Fuji PLoS One 7:e44494
Inagaki F, Nunoura T, Nakagawa S, Teske A, Lever M, Lauer A, Suzuki M, Takai K, Delwiche M, Colwell FS (2006) Biogeographical distribution and diversity of microbes in methane hydrate-bearing deep marine sediments on the Pacific Ocean Margin. Proc Natl Acad Sci U S A 103:2815–2820
Wang CS, Zhao XX, Liu ZF, Lippert PC, Graham SA, Coe RS, Yi HS, Zhu LD, Liu S, Li YL (2008) Constraints on the early uplift history of the Tibetan Plateau. Proc Natl Acad Sci U S A 105:4987–4992
Acknowledgments
This work was financially supported by grants from National Science Foundation of China (41230857, 41025004), MOST (2013CB956300), and STSN-21-02. We gratefully acknowledge Drs Mu Wang and Xi Zha from Agricultural and Animal Husbandry College of Tibet for their assistance in soil sampling.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
Spearman’s correlation analysis on environmental variable and diversity index of Shannon (a) and Faith PD (b) (DOC 98 kb)
Rights and permissions
About this article
Cite this article
Wang, JT., Cao, P., Hu, HW. et al. Altitudinal Distribution Patterns of Soil Bacterial and Archaeal Communities Along Mt. Shegyla on the Tibetan Plateau. Microb Ecol 69, 135–145 (2015). https://doi.org/10.1007/s00248-014-0465-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-014-0465-7