Abstract
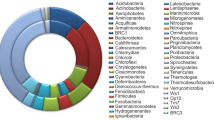
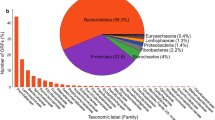
Fungus-growing termites, their symbiotic fungi, and microbiota inhibiting their intestinal tract comprise a highly efficient cellulose-hydrolyzing system; however, little is known about the role of gut microbiota in this system. Twelve fosmid clones with β-glucosidase activity were previously obtained by functionally screening a metagenomic library of a fungus-growing termite, Macrotermes annandalei. Ten contigs containing putative β-glucosidase genes (bgl1–10) were assembled by sequencing data of these fosmid clones. All these contigs were binned to Bacteroidetes, and all these β-glucosidase genes were phylogenetically closed to those from Bacteroides or Dysgonomonas. Six out of 10 β-glucosidase genes had predicted signal peptides, indicating a transmembrane capability of these enzymes to mediate cellulose hydrolysis within the gut of the termites. To confirm the activities of these β-glucosidase genes, three genes (bgl5, bgl7, and bgl9) were successfully expressed and purified. The optimal temperature and pH of these enzymes largely resembled the environment of the host’s gut. The gut microbiota composition of the fungus-growing termite was also determined by 454 pyrosequencing, showing that Bacteroidetes was the most dominant phylum. The diversity and the enzyme properties of β-glucosidases revealed in this study suggested that Bacteroidetes as the major member in fungus-growing termites contributed to cello-oligomer degradation in cellulose-hydrolyzing process and represented a rich source for β-glucosidase genes.




Similar content being viewed by others
References
Breznak JA (1994) Role of microorganisms in the digestion of lignocellulose by termites. Annu Rev Entomol 39:453–487
Abo-Khatwa AN (1989) Termitomyces: a new source of potent cellulases. JKing Abulaziz Univ Science 1:51–59
Breznak JA (1982) Intestinal microbiota of termites and other xylophagous insects. Annu Rev Microbiol 36:323–343
Wood TG, Thomas RJ (1989) The mutualistic association between Macrotermitinae and Termitomyces. In: Wilding N, Collins NM, Hammond PM, Webber JF (eds) Insect-fungus interaction. Academic, London, pp 69–92
Hongoh Y, Ekpornprasit L, Inoue T, Moriya S, Trakulnaleamsai S, Ohkuma M, Noparatnaraporn N, Kudo T (2006) Intracolony variation of bacterial gut microbiota among castes and ages in the fungus-growing termite Macrotermes gilvus. Mol Ecol 15(2):505–516
Mackenzie LM, Muigai AT, Osir EO, Lwande W, Keller M, Yoledo G, Boga HI (2007) Bacterial diversity in the intestinal tract of the fungus-cultivating termite Macrotermes michaelseni (Sjostedt). African Journal of Biotechnology 6(6):658–667
Hongoh Y, Deevong P, Inoue T, Moriya S, Trakulnaleamsai S, Ohkuma M, Vongkaluang C, Noparatnaraporn N, Kudo T (2005) Intra- and interspecific comparisons of bacterial diversity and community structure support coevolution of gut microbiota and termite host. Appl Environ Microbiol 71(11):6590–6599
Shinzato N, Muramatsu M, Matsui T, Watanabe Y (2007) Phylogenetic analysis of the gut bacterial microflora of the fungus-growing termite Odontotermes formosanus. Biosci Biotechnol Biochem 71(4):906–915
Long YH, Xie L, Liu N, Yan X, Li MH, Fan MZ, Wang QA (2010) Comparison of gut-associated and nest-associated microbial communities of a fungus-growing termite (Odontotermes yunnanensis). Insect Sci 17(3):265–276
Rouland-Lefevre C, Inoue T, Johjima T (2006) Termitomyces/termite interactions. In: Konig H, Varma A (eds) Intestinal microorganisms of soil invertebrates. Springer, Berlin, pp 335–350
Mathew GM, Ju YM, Lai CY, Mathew DC, Huang CC (2012) Microbial community analysis in the termite gut and fungus comb of Odontotermes formosanus: the implication of Bacillus as mutualists. FEMS Microbiol Ecol 79(2):504–517
Tokuda G, Lo N, Watanabe H, Arakawa G, Matsumoto T, Noda H (2004) Major alteration of the expression site of endogenous cellulases in members of an apical termite lineage. Molecular Ecology 13(10):3219–3228
Wu Y, Chi S, Yun C, Shen Y, Tokuda G, Ni J (2012) Molecular cloning and characterization of an endogenous digestive beta-glucosidase from the midgut of the fungus-growing termite Macrotermes barneyi. Insect Mol Biol 21(6):604–614
Hyodo F, Inoue T, Azuma JI, Tayasu I, Tabe T (2000) Role of the mutualistic fungus in lignin degradation in the fungus-growing termite Macrotermes gilvus (Isoptera; Macrotermitinae). Soil Biol Biochem 32:653–658
Hyodo F, Tayasu I, Inoue T, Azuma JI, Kudo T, Abe T (2003) Differential role of symbiotic fungi in lignin degradation and food provision for fungus-growing termites (Macrotermitinae: Isoptera). Functional Ecology 17:186–193
Johjima T, Taprab Y, Noparatnaraporn N, Kudo T, Ohkuma M (2006) Large-scale identification of transcripts expressed in a symbiotic fungus (Termitomyces) during plant biomass degradation. Appl Microbiol Biotechnol 73(1):195–203
Martin M, Martin J (1978) Cellulose digestion in the midgut of the fungus-growing termite Macrotermes natalensis: the role of acquired digestive enzyme. Science 199:1453–1455
Matoub M, Rouland C (1995) Purification and properties of the xylanases from the termite Macrotermes bellicosus and its symbiotic fungus Termitomyces sp. Comp Biochem Physiol B Biochem Mol Biol 112(4):629–635
Liu N, Yan X, Zhang M, Xie L, Wang Q, Huang Y, Zhou X, Wang S, Zhou Z (2011) Microbiome of fungus-growing termites: a new reservoir for lignocellulase genes. Appl Environ Microbiol 77(1):48–56
Liu N, Zhang L, Zhou H, Zhang M, Yan X, Wang Q, Long Y, Xie L, Wang S, Huang Y, Zhou Z (2013) Metagenomic insights into metabolic capacities of the gut microbiota in a fungus-cultivating termite (Odontotermes yunnanensis). PLoS One 8(7):e69184
Bastien G, Arnal G, Bozonnet A, Laguerre S, Ferreir F, GFaure R, Henrissat B, Lefevre G, Robe P, Bouchez O, Noirot C (2013) Mining for hemicellulases in the fungus-growing termite Pseudacanthotermes militaris using functional metagenomics. Biotechnol Biofuels 6:78. doi:10.1186/1754-6834-1186-1178
DeSantis TZ, Jr., Hugenholtz P, Keller K, Brodie EL, Larsen N, Piceno YM, Phan R, Andersen GL (2006) NAST: a multiple sequence alignment server for comparative analysis of 16S rRNA genes. Nucleic Acids Res 34(Web Server issue):W394-399
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72(7):5069–5072
Huber T, Faulkner G, Hugenholtz P (2004) Bellerophon: a program to detect chimeric sequences in multiple sequence alignments. Bioinformatics 20(14):2317–2319
Schloss PD, Handelsman J (2005) Introducing DOTUR, a computer program for defining operational taxonomic units and estimating species richness. Appl Environ Microbiol 71(3):1501–1506
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73(16):5261–5267
Huson DH, Auch AF, Qi J, Schuster SC (2007) Megan analysis of metagenomic data. Genome Res 17:377–386
Mitra S, Klar B, Huson DH (2009) Visual and statistical comparison of metagenomes. Bioinformatics 25:1849
Katoh K, Toh H (2008) Recent developments in the MAFFT multiple sequence alignment program. Brief Bioinformatics 9:286–298
Mackenzie LM, Muigai AT, Osir EO, Lwande W, Keller M, Toledo G, Hi B (2007) Bacterial diversity in the intestinal tract of the fungus-cultivating termite Macrotermes michaelseni (Sjostedt). African Journal of Biotechnology 6:658–667
Kohler T, Dietrich C, Scheffrahn RH, Brune A (2012) High-resolution analysis of gut environment and bacterial microbiota reveals functional compartmentation of the gut in wood-feeding higher termites (Nasutitermes spp.). Appl Environ Microbiol 78(13):4691–4701
Warnecke F, Luginbuhl P, Ivanova N, Ghassemian M, Richardson TH, Stege JT, Cayouette M, McHardy AC, Djordjevic G, Aboushadi N, Sorek R, Tringe SG, Podar M, Martin HG, Kunin V, Dalevi D, Madejska J, Kirton E, Platt D, Szeto E, Salamov A, Barry K, Mikhailova N, Kyrpides NC, Matson EG, Ottesen EA, Zhang X, Hernandez M, Murillo C, Acosta LG, Rigoutsos I, Tamayo G, Green BD, Chang C, Rubin EM, Mathur EJ, Robertson DE, Hugenholtz P, Leadbetter JR (2007) Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nature 450(7169):560–565
Lawson PA, Falsen E, Inganas E, Weyant RS, Collins MD (2002) Dysgonomonas mossii sp. nov., from human sources. Syst Appl Microbiol 25(2):194–197
Martens EC, Koropatkin NM, Smith TJ, Gordon JI (2009) Complex glycan catabolism by the human gut microbiota: the Bacteroidetes Sus-like paradigm. J Biol Chem 284(37):24673–24677
Dodd D, Mackie RI, Cann IK (2011) Xylan degradation, a metabolic property shared by rumen and human colonic Bacteroidetes. Mol Microbiol 79(2):292–304
Mackenzie AK, Pope PB, Pedersen HL, Gupta R, Morrison M, Willats WG, Eijsink VG (2012) Two SusD-like proteins encoded within a polysaccharide utilization locus of an uncultured ruminant Bacteroidetes phylotype bind strongly to cellulose. Appl Environ Microbiol 78(16):5935–5937
Carpenter KJ, Keeling PJ (2007) Morphology and phylogenetic position of Eucomonympha imla (Parabasalia: Hypermastigida). J Eukaryot Microbiol 54(4):325–332
Hansen PS, Jensen TG, Gahrn-Hansen B (2005) Dysgonomonas capnocytophagoides bacteraemia in a neutropenic patient treated for acute myeloid leukaemia. APMIS 113(3):229–231
Hironaga M, Yamane K, Inaba M, Haga Y, Arakawa Y (2008) Characterization and antimicrobial susceptibility of Dysgonomonas capnocytophagoides isolated from human blood sample. Jpn J Infect Dis 61(3):212–213
Hofstad T, Olsen I, Eribe ER, Falsen E, Collins MD, Lawson PA (2000) Dysgonomonas gen. nov. to accommodate Dysgonomonas gadei sp. nov., an organism isolated from a human gall bladder, and Dysgonomonas capnocytophagoides (formerly CDC group DF-3). Int J Syst Evol Microbiol 50(6):2189–2195
Lawson PA, Carlson P, Wernersson S, Moore ER, Falsen E (2010) Dysgonomonas hofstadii sp. nov., isolated from a human clinical source. Anaerobe 16(2):161–164
Matsumoto T, Kawakami Y, Oana K, Honda T, Yamauchi K, Okimura Y, Shiohara M, Kasuga E (2006) First isolation of Dysgonomonas mossii from intestinal juice of a patient with pancreatic cancer. Arch Med Res 37(7):914–916
Collins MD, Lawson PA, Labrenz M, Tindall BJ, Weiss N, Hirsch P (2002) Nesterenkonia lacusekhoensis sp. nov., isolated from hypersaline Ekho Lake, East Antarctica, and emended description of the genus Nesterenkonia. Int J Syst Evol Microbiol 52(Pt 4):1145–1150
Martin MM, Matrtin JS (1979) The distribution and origins of the cellulolytic enzymes of the higher termite, Macrotermes natalensis. Physiological Zoology 52(1):11–21
Bhatia Y, Mishra S, Bisaria VS (2002) Microbial beta-glucosidases: cloning, properties, and applications. Crit Rev Biotechnol 22:375–407
Suto M, Tomita F (2001) Induction and catabolite repression mechanisms of cellulase in fungi. J Biosci Bioeng 92(4):305–311
Percival Zhang YH, Himmel ME, Mielenz JR (2006) Outlook for cellulase improvement: screening and selection strategies. Biotechnol Adv 24(5):452–481
Feng Y, Duan CJ, Liu L, Tang JL, Feng JX (2009) Properties of a metagenome-derived beta-glucosidase from the contents of rabbit cecum. Biosci Biotechnol Biochem 73(7):1470–1473
Jiang C, Ma G, Li S, Hu T, Che Z, Shen P, Yan B, Wu B (2009) Characterization of a novel beta-glucosidase-like activity from a soil metagenome. J Microbiol 47(5):542–548
Kataeva IA, Seidel RD 3rd, Shah A, West LT, Li XL, Ljungdahl LG (2002) The fibronectin type 3-like repeat from the Clostridium thermocellum cellobiohydrolase CbhA promotes hydrolysis of cellulose by modifying its surface. Appl Environ Microbiol 68(9):4292–4300
Brulc JM, Antonopoulos DA, Miller ME, Wilson MK, Yannarell AC, Dinsdale EA, Edwards RE, Frank ED, Emerson JB, Wacklin P, Coutinho PM, Henrissat B, Nelson KE, White BA (2009) Gene-centric metagenomics of the fiber-adherent bovine rumen microbiome reveals forage specific glycoside hydrolases. Proc Natl Acad Sci U S A 106(6):1948–1953
Acknowledgments
This work was supported by grants of the National Natural Science Foundation of China (31070098). We appreciate Professor Shubiao Wu from the University of New England for his critical reading and kind suggestions. We also give our thanks to Prashanth Singanallur and Lauren Christine Radlinski from the University of Illinois at Urbana-Champaign for proofreading. We appreciated Dr. Haokui Zhou from Department of Microbiology, The Chinese University of Hongkong and Lei zhang from Logic Informatics Co.,Ltd., for their kind help in data analysis.
Conflict of Interest
The authors have no conflict of interest to declare.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PPTX 48413 kb)
Rights and permissions
About this article
Cite this article
Zhang, M., Liu, N., Qian, C. et al. Phylogenetic and Functional Analysis of Gut Microbiota of a Fungus-Growing Higher Termite: Bacteroidetes from Higher Termites Are a Rich Source of β-Glucosidase Genes. Microb Ecol 68, 416–425 (2014). https://doi.org/10.1007/s00248-014-0388-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-014-0388-3