Abstract
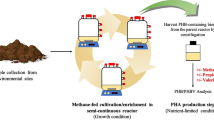
Methanotrophs are known to produce poly-3-hydroxybutyrate (PHB), but there is conflicting evidence in the literature as to which genera produce the polymer. We screened type I and II proteobacterial methanotrophs that use the ribulose monophosphate and serine pathways for carbon assimilation, respectively, for both phaC, which encodes for PHB synthase, and the ability to produce PHB under nitrogen-limited conditions. Twelve strains from six different genera were evaluated. All type I strains tested negative for phaC and PHB production; all Type II strains tested positive for phaC and PHB production. In order to identify conditions that favor PHB production, we also evaluated a range of selection conditions using a diverse activated sludge inoculum. Use of medium typically recommended for methanotroph enrichment led to enrichments dominated by type I methanotrophs. Conditions that were selected for enrichments dominated by PHB-producing Type II methanotrophs were: (1) use of nitrogen gas as the sole nitrogen source in the absence of copper, (2) use of a dilute mineral salts media in the absence of copper, and (3) use of media prepared at pH values of 4–5.
Similar content being viewed by others
References
Amaral JA, Archambault C, Richards SR, Knowles R (1995) Denitrificationassociated with groups I and II methanotrophs in a gradient enrichment system. FEMS Microbiol Ecol 18:289–298
Anderson AJ, Dawes EA (1990) Occurrence, metabolism, metabolic role, and industrial uses of bacterial polyhydroxyalkanoates. Microbiol Rev 54:450–472
Asenjo JA, Suk J (1986) Microbial conversion of methaneintopoly-beta-hydroxybutrate (PHB)-growth and intracellular product accumulation in a type-II methanotroph. J Ferment Technol 64:271–278
Ashby RD, Solaiman DKY, Foglia TA (2004) Bacterialpoly (hydroxyalkanoate) polymer production from the bio diesel co-product stream. J Polym Environ 12:105–112
Auman AJ, Speake CC, Lidstrom ME (2001) nifH sequences and nitrogen fixation in type I and type II methanotrophs. Appl Environ Microbiol 67:4009–4016
Babel W (1992) Pecularities of methylotrophs concerning over flow metabolism, especially the synthesis of polyhydroxyalkanoates. FEMS Microbiol Rev 103:141–148
Bowman JP (2001) Family I. Methylococcaceae and Family V. Methylocystaceae. In: Bergey’s manual of systematic bacteriology. Williams & Wilkins, Baltimore, pp 256–270, pp. 411–420
Bowman JP (2006) The methanotrophs-the families Methylococcacceae and Methylocystaceae. In: The prokaryotes a: handbook on the biology of bacteria. Springer, NewYork, pp 266–289
Bowman JP, Sly LI, Nichols PD, Hayward A (1993) Revised Taxonomy of the Methanotrophs: Description of Methylobacter gen.nov., emendation of Methylococcus, validation of Methylosinus and Methylocystis species, and a proposal that the family Methylococcaceae includes only the group I methanotrophs. Int J Syst Bacteriol 43:735–753
Braunegg G, Lefebvre G, Genser KF (1999) Polyhydroxyalkanoates, biopolyesters from renewable resources: physiological and engineering aspects. J Biotechnol 65:127–161
Braunegg G, Sonnleitner B, Lafferty R (1978) Arapid gas chromatographic method for the determination of poly-β-hydroxybutyric acid in microbial biomass. Eur J Appl Microbiol Biotechnol 6:29–37
Budwill K, Fedorak PM, Page WJ (1992) Methanogenic degradation of poly (3- hydroxyalkanoates). Appl Environ Microbiol 58:1398–1401
Bussman I, Pester M, Brune A, Schink B (2004) Preferential cultivation of type II methanotrophic bacteria from littoral sediments (Lake Constance). FEMS Microbiol Ecol 47:179–189
Cebron A, Bodrossy L, Stralis-Pavese N, Singer AC, Thompson IP, Prosser JI, Murrell JC (2007) Nutrient amendments in soil DNA stable isotope probing experiments reduce the observed methanotroph diversity. Appl Environ Microbiol 73:798–807
Choi J, Lee SY (1997) Process analysis and economic evaluation for Poly (3- hydroxybutyrate) production by fermentation. Bioprocess Eng 17:335–342
Costello AM, Lidstrom ME (1999) Molecular characterization of functional and phylogenetic genes from natural populations of methanotrophs in lake sediments. Appl Environ Microbiol 65:5066–5074
Dedysh SN (2002) Methanotrophic bacteria of acids phagnum bogs. Mikrobiologiia 71:741–754
Dedysh SN, Khmelenina VN, Suzina NE, Trotsenko YA, Semrau JD, Liesack W, Tiedje JM (2002) Methylocapsa acidiphila gen. nov., sp. nov., a novel methane-oxidizing and dinitrogen-fixing acidophilic bacterium from sphagnum bog. Int J Syst Evol Microbiol 52:251–261
Dedysh SN, Liesack W, Khmelenina VN, Suzina NE, Trotsenko YA, Semrau JD, Bares AM, Panikov NS, Tiedje JM (2000) Methylocellapalustrisgen.nov., sp.nov., a new methane-oxidizing acid ophilic bacterium from peat bogs, representing a novel subtype of serine-pathway methanotrophs. Int J Syst Evol Microbiol 50:955–969
Du G, Chen LXL, Yu J (2004) High-efficiency production of bioplastics from biodegradable organic solids. J Polym Environ 12:89–94
Dunfield PF, Khmelenina VN, Suzina NE, Trotsenko YA, Dedysh SN (2003) Methylocella silvestrissp.nov.,a novel methanotroph isolated from an acidic forest cambisol. Int J Syst Evol Microbiol 53:1231–1239
Dunfield PF, Yuryev A, Senin P, Smirnova AV, Stott MB, Hou S, Ly B, Saw JH, Zhou Z, Ren Y, Wang J, Mountain BW, Crowe MA, Weatherby TM, Bodelier PL, Liesack W, Feng L, Wang L, Alam M (2007) Methane oxidation by an extremely acid ophilic bacterium of the phylum Verrucomicrobia. Nature 450:879–882
Follner CG, Babel W, Valentin HE, Steinbuchel A (1993) Expression of polyhydroxy alkanoic-acid-biosynthesis genes in methylotrophic bacteria relying on the ribulose monophosphate pathway. Appl Microbiol Biotechnol 40:284–291
Graham DW, Chaudhary JA, Hanson RS, Arnold RG (1993) Factors affecting competition between type I and type II methanotrophs in two-organism, continuous-flowreactors. Microb Ecol 25:1–17
Hanson RS (1998) Ecology of methylotrophic bacteria. In: Techniques in microbial ecology. Oxford University Press, London, pp 137–162
Hanson RS, Hanson TE (1996) Methanotrophic bacteria. Microbiol Rev 60:439–471
Harrington AA, Kallio RE (1960) Oxidation of methanol and formal dehyde by Pseudomonas methanica. Can J Microbiol 6:1–7
Helm J, Wendlandt KD, Jechorek M, Stottmeister U (2008) Potassium deficiency results in accumulation of ultra-high molecular weight poly-beta-hydroxybutyrate in a methane-utilizing mixed culture. J Appl Microbiol 105:1054–1061
Helm J, Wendlandt KD, Rogge G, Kappelmeyer U (2006) Characterizing a stable methane-utilizing mixed culture used in the synthesis of a high-quality biopolymer in an open system. J Appl Microbiol 101:387–395
Heyer J, Berger U, Hardt M, Dunfield PF (2005) Methylohalobiuscrimeensisgen.nov., sp.nov., a moderately halophilic, methanotrophic bacterium isolated from hyper-saline lakes of Crimea. Int J Syst Evol Microbiol 55:1817–1826
Higgins IJ, Best DJ, Hammond RC, Scott D (1981) Methane-oxidizing microorganisms. Microbiol Rev 45:556–590
Islam T, Jensen S, Reigstad LJ, Larsen O, Birkeland NK (2008) Methane oxidation at 55 degrees C and pH2 by a thermo acid ophilic bacterium belonging to the Verrucomicrobia phylum. Proc Natl Acad Sci USA 105:300–304
Kallio RE, Harrington AA (1960) Sudanophilic granules and lipid of Pseudomonas methanica. J Bacteriol 80:321–324
Koller M, Bona R, Braunegg G, Hermann C, Horvat P, Kroutil M, Martinz J, Neto J, Pereira L, Varila P (2005) Production of polyhydroxyalkanoates from agricultural waste and surplus materials. Biomacro Molecules 6:561–565
Korotkova N, Lidstrom ME (2001) Connection between poly-beta-hydroxybutyrate biosynthesis and growth on C (1) and C (2) compounds in the methylotroph Methylobacterium extorquens AM1. J Bacteriol 183:1038–1046
Lee SY, Park SJ, Park JP, Lee Y, Lee SH (2005) Economic aspects of biopolymer production, vol 2. WILEY-VCH, Weinheim
Lidstrom ME (2006) Aerobic methylotrophic prokaryotes. In: Prokaryotes. pp. 618–634
Madison LL, Huisman GW (1999) Metabolic engineering of poly (3-hydroxyalkanoates): from DNA to plastic. Microbiol Mol Biol Rev 63:21–53
Morse M, Liao Q, Criddle CS, Frank CW (2011) An aerobic biodegradation of the microbial copolymer poly (3-hydroxybutyrate-co-3-hydroxyhexanoate): Effects of comonomer content, processing history, and semi-crystalline morphology. Polymer 52:547–555
Murrell J, Dalton H (1983) Nitrogen-fixation in obligate methanotrophs. J Gen Microbiol 129:3481–3486
Nguyen HH, Elliott SJ, Yip JH, Chan SI (1998) The particulate methane monooxygenase from M. capsulatus (Bath) is a novel copper-containing three-subunit enzyme. J Biol Chem 273:7957–7966
Oakley C, Murrell J (1988) Nifh genes in the obligate methane oxidizing bacteria. FEMS Microbiol Lett 49:53–57
HJ Op denCamp, Islam T, Stott MB, Harhangi HR, Hynes A, Schouten S, Jetten MS, Birkeland NK, Pol A, Dunfield PF (2009) Environmental, genomic and taxonomic perspectives on methanotrophic Verrucomicrobia. Environ Microbiol Rep 1:293–306
Pol A, Heijmans K, Harhangi HR, Tedesco D, Jetten MS, OpdenCamp HJ (2007) Methanotrophy below pH1 by a new Verrucomicrobia species. Nature 450:874–878
Povolo S, Casella S (2003) Bacterial production of PHA from lactose and cheese whey permeate. Macromol Symp 197:1–9
Shah NN, Hanna ML, Jackson KJ, Taylor RT (1996) Batch cultivation of Methylosinus trichosporium OB3B: IV production of hydrogen-driven soluble or particulate methane monooxygenase activity. Biotechnol Bioeng 45:229–238
Sheu DS, Wang YT, Lee CY (2000) Rapid detection of polyhydroxyalkanoate-accumulating bacteria isolated from the environment by colony PCR. Microbiology 146:2019–2025
Stocks PK, Mc Cleskey CS (1964) Identity of the pink-pigmented methanol- oxidizing bacteria as Vibrio extorquens. J Bacteriol 88:1065–1070
U.S.EnvironmentalProtectionAgency. Methane: sources and emissions. http://www.epa.gov/outreach/sources.html. Accessed April21, 2011
Vecherskaya M, Dijkema C, Stams AJ (2001) Intracellular PHB conversion in a type II methanotroph studied by13CNMR. J Ind Microbiol Biotechnol 26:15–21
Vorob’ev AV, Dedysh SN (2008) Use of enrichment cultures for assessing the structure of methanotrophic communities in peat soil: the problem of representativity of the results. Mikrobiologiia 77:566–569
Ward N, Larsen O, Sakwa J, Bruseth L, Khouri H, Durkin AS, Dimitrov G, Jiang L, Scanlan D, Kang KH, Lewis M, Nelson KE, Methe B, Wu M, Heidelberg JF, Paulsen IT, Fouts D, Ravel J, Tettelin H, Ren Q, Read T, DeBoy RT, Seshadri R, Salzberg SL, Jensen HB, Birkeland NK, Nelson WC, Dodson RJ, Grindhaug SH, Holt I, Eidhammer I, Jonasen I, Vanaken S, Utterback T, Feldblyum TV, Fraser CM, Lillehaug JR, Eisen JA (2004) Genomic insights into methanotrophy: the complete genome sequence of M. capsulatus (Bath). PLoS Biol 2:e303
Wendlandt KD, Geyer W, Mirschel G, Al-HajHemidi F (2005) Possibilities for controlling a PHB accumulation process using various analytical methods. J Biotechnol 117:119–129
Wendlandt KD, Jechorek M, Helm J, Stottmeister U (2001) Producing poly-3-hydroxybutyrate with a high molecular mass from methane. J Biotechnol 86:127–133
Wise MG, McArthur JV, Shimkets LJ (1999) Methanotroph diversity in land fill soil: isolation of novel type I and type II methanotrophs whose presence was suggested by culture-independent 16S ribosomal DNA analysis. Appl Environ Microbiol 65:4887–4897
Xin JY, Zhang YX, Zhang S, Xia CG, Li SB (2007) Methanol production from CO2by resting cells of the methanotrophic bacterium Methylosinus trichosporium IMV3011. J Basic Microbiol 47:426–435
Zahn JA, DiSpirito AA (1996) Membrane-associated methane monooxygenase from M. capsulatus (Bath). J Bacteriol 178:1018–1029
Zhang Y, Xin J, Chen L, Song H, Xia C (2008) Biosynthesis of poly-3-hydroxybutyrate with a high molecular weight by methanotroph from methane and methanol. J Nat Gas Chem 17:103–109
Acknowledgments
We thank Dr. Lisa Stein and the Carbon-one Oxidation Network for the phaC gene sequence for M. trichosporium OB3b. We also thank Holly Sewell for her help in the laboratory.
This work was supported by a Graduate Research Fellowship from the National Science Foundation to AJP and by the California Environmental Protection Agency, Department of Toxic Substances Control under contract 07T3451.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pieja, A.J., Rostkowski, K.H. & Criddle, C.S. Distribution and Selection of Poly-3-Hydroxybutyrate Production Capacity in Methanotrophic Proteobacteria. Microb Ecol 62, 564–573 (2011). https://doi.org/10.1007/s00248-011-9873-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-011-9873-0