Abstract
Background
An automated method for identifying the anatomical region of an image independent of metadata labels could improve radiologist workflow (e.g., automated hanging protocols) and help facilitate the automated curation of large medical imaging data sets for machine learning purposes. Deep learning is a potential tool for this purpose.
Objective
To develop and test the performance of deep convolutional neural networks (DCNN) for the automated classification of pediatric musculoskeletal radiographs by anatomical area.
Materials and methods
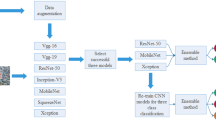
We utilized a database of 250 pediatric bone radiographs (50 each of the shoulder, elbow, hand, pelvis and knee) to train 5 DCNNs, one to detect each anatomical region amongst the others, based on ResNet-18 pretrained on ImageNet (transfer learning). For each DCNN, the radiographs were randomly split into training (64%), validation (12%) and test (24%) data sets. The training and validation data sets were augmented 30 times using standard preprocessing methods. We also tested our DCNNs on a separate test set of 100 radiographs from a single institution. Receiver operating characteristics (ROC) with area under the curve (AUC) were used to evaluate DCNN performances.
Results
All five DCNN trained for classification of the radiographs into anatomical region achieved ROC AUC of 1, respectively, for both test sets. Classification of the test radiographs occurred at a rate of 33 radiographs per s.
Conclusion
DCNNs trained on a small set of images with 30 times augmentation through standard processing techniques are able to automatically classify pediatric musculoskeletal radiographs into anatomical region with near-perfect to perfect accuracy at superhuman speeds. This concept may apply to other body parts and radiographic views with the potential to create an all-encompassing semantic-labeling DCNN.


Similar content being viewed by others
References
Rajkomar A, Lingam S, Taylor AG et al (2017) High-throughput classification of radiographs using deep convolutional neural networks. J Digit Imaging 30:95–101
Lakhani P (2017) Deep convolutional neural networks for endotracheal tube position and X-ray image classification: challenges and opportunities. J Digit Imaging 30:460–468
Lakhani P, Sundaram B (2017) Deep learning at chest radiography: automated classification of pulmonary tuberculosis by using convolutional neural networks. Radiology 284:574–582
LeCun Y, Bengio Y, Hinton G (2015) Deep learning. Nature 521:436–444
Cheng PM, Malhi HS (2017) Transfer learning with convolutional neural networks for classification of abdominal ultrasound images. J Digit Imaging 30:234–243
Zhou B, Khosla A, Lapedriza A et al (2016) Learning deep features for discriminative localization. In: 2016 IEEE Conf Comput Vis Pattern Recognit, pp 2921–2929
Ting DSW, Cheung CY, Lim G et al (2017) Development and validation of a deep learning system for diabetic retinopathy and related eye diseases using retinal images from multiethnic populations with diabetes. JAMA 318:2211–2223
Kim DH, MacKinnon T (2018) Artificial intelligence in fracture detection: transfer learning from deep convolutional neural networks. Clin Radiol 73:439–445
Acknowledgments
The authors acknowledge Tae Soo Kim, MSE, for technical advising.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
None
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yi, P.H., Kim, T.K., Wei, J. et al. Automated semantic labeling of pediatric musculoskeletal radiographs using deep learning. Pediatr Radiol 49, 1066–1070 (2019). https://doi.org/10.1007/s00247-019-04408-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00247-019-04408-2