Abstract
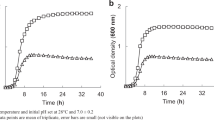
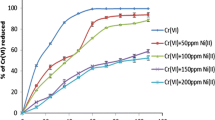
Cupriavidus metallidurans CH34 and Pseudomonas putida mt2 were used as cadmium (Cd)-resistant and -sensitive bacteria, respectively, to study Cd uptake, sorption, intracellular accumulation, metallothionein (MT) induction, and bioremediation potential of both isolates. According to this research work, Cd had a stimulatory effect on the growth of CH34 cells (OD578 = 1.43) compared with mt2 cells (OD578 = 0.8). Addition of N,N′-dicyclohexylcarbodiimide (DCCD) and 2,4-dinitrophenol (DNP) along with Cd resulted in more cell growth in mt2 (OD578 = 0.71) compared with CH34 (OD578 = 0.34). DCCD and DNP inhibited this active uptake only in CH34 but not in mt2. Greater Cd interaction with the cell surface was observed in mt2 cells compared with CH34 cells. Intracellular Cd accumulation was interrupted by DCCD and DNP in CH34 (only 1.81 ± 0.04 μg L−1 at 5 h) but not in mt2 (24.41 ± 0.01 μg L−1 at 5 h). Intracellular Cd uptake was observed in even killed mt2 cells (7.11 ± 0.05 μg L−1 at 5 h) compared with CH34 cells (2.50 ± 0.08 μg L−1 at 5 h). This result showed that the Cd accumulation mechanism in CH34 is ATPase-dependent, whereas in mt2 uptake mechanism is not ATPase-dependent because mt2 ATPase was not inhibited by DCCD and DNP. CH34 removed 93 mg L−1 of Cd after 8 days from original industrial effluent, which was more than Cd removal by CH34 from distilled water (i.e. 90 mg L−1 after 8 days). mt2 was able to remove 80 mg L−1 of Cd after 8 days from original industrial effluent, which was more than Cd removal by mt2 from distilled water (i.e. 77 mg L−1 after 8 days). Cd did not induce any MT in CH34, but it did so in mt2 (14 kDa), which was thought to be a Cd-resistance mechanism operative in mt2.




Similar content being viewed by others
References
Anton A, Grosse C, Reißmann J, Pribyl T et al (1999) CzcD is a heavy metal ion transporter involved in regulation of heavy metal resistance in Ralstonia sp. strain CH34. J Bacteriol 181:6876–6881
Argüello JM (2003) Identification of ion-selectivity determinants in heavy-metal transport P1B-type ATPases. J Membrane Biol 195:93–108
Blindauer CA, Harrison MD, Robinson AK, Parkinson JA et al (2002) Multiple bacteria encode metallothioneins and SmtA-like zinc fingers. Mol Microbiol 45:1421–1432
Bradford M (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Canovas D, Cases I, de Lorenzo V (2003) Heavy metal tolerance and metal homeostasis in Pseudomonas putida as revealed by complete genome analysis. Environ Microbiol 5:1242–1256
Carrasquer G, Li M, Dinno MA (2000) Effect of dicyclohexylcarbodiimide (DCCD) on transport parameters in the frog cornea epithelium. J Membrane Biol 174(2):97–103
Cavet JS, Borrelly GPM, Robinson NJ (2003) Zn, Cu and Co in cyanobacteria: selective control of metal availability. FEMS Microbiol Rev 27:165–181
Das N, Vimala R, Karthika P (2008) Biosorption of heavy metals: an overview. Indian J Biotechnol 7:159–169
Enshaei M, Khanafari A, Sepahey AA (2010) Metallothionein induction in two species of Pseudomonas exposed to cadmium and copper contamination. Iran J Environ Health 7(4):287–298
Gadd GM (1990) Heavy metal accumulation by bacteria and other microorganisms. Experientia 46(8):834–840
Goyal N, Jain SC, Banerjee UC (2003) Comparative studies on the adsorption of heavy metals. Adv Environ Res 7:311–319
Grass G, Wong MD, Rosen BP, Smith RL et al (2002) ZupT is a Zn (II) uptake system in Escherichia coli. J Bacteriol 184:864–866
Hassan MT, van der Lelie D, Springael D, Römling U et al (1999) Identification of a gene cluster, czr, involved in cadmium and zinc resistance in Pseudomonas aeruginosa. Gene 238:417–425
Hu N, Zhao B (2007) Key genes involved in heavy-metal resistance in Pseudomonas putida CD2. FEMS Microbiol Lett 267:17–22
Hu P, Brodie EL, Suzuki Y, McAdams HH et al (2005) Whole-genome transcriptional analysis of heavy metal stresses in Caulobacter crescentus. J Bacteriol 187(24):8437–8449
Hynninen A, Tõnismann K, Virta M (2010) Improving the sensitivity of bacterial bioreporters for heavy metals. Bioeng Bugs 1(2):1–7
Khleifat KM, Abboud MM, Al-Mustafa AH (2006) Effect of Vitreoscilla hemoglobin gene (vgb) and metabolic inhibitors on cadmium uptake by the heterologous host Enterobacter aerogenes. Process Biochem 41:930–934
Korkmaz K, Kara SM, Ozkutlu F, Gul V (2010) Monitoring of heavy metals and selected micronutrients in hempseeds from north-western Turkey. Afr J Agric Res 5(6):463–467
Koutinas M, Lam MC, Kiparissides A, Silva-Rocha R et al (2010) The regulatory logic of m-xylene biodegradation by Pseudomonas putida mt-2 exposed by dynamic modelling of the principal node Ps/Pr of the TOL plasmid. Environ Microbiol 12(6):1705–1718
Kunito T, Kusano T, Oyaizu H, Senoo K et al (1996) Cloning and sequence analysis of czc genes in Alcaligenes sp. strain CT14. Biosci Biotechnol Biochem 60:699–704
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Leedjärv A, Ivask A, Virta M (2008) Interplay of different transporters in the mediation of divalent heavy metal resistance in Pseudomonas putida KT2440. J Bacteriol 190:2680–2689
Legatzki A, Grass G, Anton A, Rensing C et al (2003) Interplay of the Czc system and two P-type ATPases in conferring metal resistance to Ralstonia metallidurans. J Bacteriol 185(15):4354–4361
Li Z, Yuan H, Hu X (2008) Cadmium-resistance in growing Rhodotorula sp. Y11. Bioresour Technol 99:1339–1344
Liu CQ, Khunajakr N, Chia LG, Deng YM et al (1997) Genetic analysis of regions involved in replication and cadmium resistance of the plasmid pND302 from Lactococcus lactis. Plasmid 38:79–90
Mahvi AH, Diels L (2004) Biological removal of cadmium by Alcaligenes eutrophus CH34. Int J Environ Sci Technol 1(3):199–204
Malekzadeh F, Latifi AM, Shahamat M, Levin M et al (2002) Effects of selected physical and chemicals parameters on uranium uptake by the bacterium Chryseomonas MGF-48. World J Microbiol Biotechnol 18:599–602
Matsumoto Y, Hayama K, Sakakihara S, Nishino K et al (2011) Evaluation of multidrug efflux pump inhibitors by a new method using microfluidic channels. PLoS One 6(4):1–12
Miller CD, Pettee B, Zhang C, Pabst M et al (2009) Copper and cadmium: responses in Pseudomonas putida KT2440. Lett Appl Microbiol 49:775–783
Monchy S, Vallaeys T, Bossus A, Mergeay M (2006) Metal transport ATPase genes from Cupriavidus metallidurans CH34: a transcriptomic approach. Int J Environ Anal Chem 86:677–692
Muñoz R, Hernández M, Segura A, Gouveia J et al (2009) Continuous cultures of Pseudomonas putida mt-2 overcome catabolic function loss under real case operating conditions. Appl Microbiol Biot 83:189–198
Murakami S, Nakashima R, Yamashita E, Yamaquchi A (2002) Crystal structure of bacterial multidrug efflux transporter AcrB. Nature 419:587–593
Murthy S, Bali G, Sarangi SK (2011) Effect of lead on metallothionein concentration in lead resistant bacteria Bacillus cereus isolated from industrial effluent. Afr J Biotechnol 10(71):15966–15972
Naik MM, Shamim K, Dubey SK (2012) Biological characterization of lead-resistant bacteria to explore role of bacterial metallothionein in lead resistance. Curr Sci India 103(45):426–429
Neumann G, Veeranagouda Y, Karegoudar TB, Sahin O et al (2005) Cells of Pseudomonas putida and Enterobacter sp. adapt to toxic organic compounds by increasing their size. Extremophiles 9:163–168
Nies DH (1992) Resistance to cadmium, cobalt, zinc, nickel in microbes. Plasmid 27:17–28
Nies DH (1999) Microbial heavy metal resistance. Appl Microbiol Biotechnol 51:730–750
Nies DH (2003) Efflux-mediated heavy metal resistance in prokaryotes. FEMS Microbiol Rev 27:313–339
Nies DH, Silver S (1989) Plasmid-determined inducible efflux is responsible for resistance to cadmium, zinc, and cobalt in Alcaligenes eutrophus. J Bacteriol 171(2):896–900
Nies DH, Silver S (1995) Ion efflux systems involved in bacterial metal resistances. J Indust Microbiol 14(2):186–199
Nies D, Mergeay M, Friedrich B, Schlegel HG (1987) Cloning of plasmid genes encoding resistance to cadmium, zinc and cobalt in Alcaligenes eutrophus CH34. J Bacteriol 169:4865–4868
Nikaido H, Takatsuka Y (2009) Mechanisms of RND multidrug efflux pumps. Biochem Biophys Acta 1794(5):769–781
Patel SP, Sullivan PG, Pandya JD, Rabchevsky AG (2009) Differential effects of the mitochondrial uncoupling agent 2,4-dinitrophenol, or the nitroxide antioxidant, Tempol, on synaptic or nonsynaptic mitochondria after spinal cord injury. J Neurosci Res 87(1):130–140
Pieper DH, Santos VAP, Golyshin PN (2004) Genomic and mechanistic insights into the biodegradation of organic pollutants. Curr Opin Biotech 15:215–224
Rani MJ, Hemambika B, Hemapriya J, Kannan VR (2010) Comparative assessment of heavy metal removal by immobilized and dead bacterial cells: a biosorption approach. Afr J Environ Sci Technol 4(2):77–83
Rehman A, Anjum MS, Hasnain S (2010) Cadmium biosorption by yeast Candida tropicalis CBL-1, isolated from industrial wastewater. J Gen Appl Microbiol 56:359–368
Ryvolova M, Krizkova S, Adam V, Beklova M et al (2011) Analytical methods for metallothionein detection. Curr Anal Chem 7:243–261
Scherer J, Nies DH (2009) CzcP is a novel efflux system contributing to transition metal resistance in Cupriavidus metallidurans CH34. Mol Microbiol 73(4):601–621
Shamim S, Rehman A (2012) Cadmium resistance and accumulation potential of Klebsiella pneumoniae strain CBL-1 isolated from industrial wastewater. Pak J Zool 44(1):203–208
Shamim S, Rehman A (2013) Antioxidative enzyme profiling and biosorption ability of Cupriavidus metallidurans CH34 and Pseudomonas putida mt2 under cadmium stress. J Basic Microbiol 53:1–8
Shaw JP, Harayama S (1992) Purification and characterization of the NADH-acceptor reductase component of xylene monooxygenase encoded by the TOL plasmid pWW0 of Pseudomonas putida mt2. Eur J Biochem 209:51–61
Siegel A, Sigel H, Sigel RKO (2009) Metal ions in life sciences, vol 5. RSC Publishing, Cambridge, pp 51–58
Acknowledgments
The corresponding author greatly acknowledges DAAD (Deutscher Akademischer Austauschdienst–German Academic Exchange Program). The author conveys special thanks to Sibylle Mothes for providing AAS and ICP-AES facilities and Lukas Y. Wick for providing P. putida mt2 isolate and laboratory facilities at the Department of Environmental Microbiology–UMB, Helmholtz Center for Environmental Research–UFZ, Leipzig, Germany. The technical assistance of Jana Reichenbach (UMB), Ines Volkmann, and Jürgen Steffen at the Department of Analytical Chemistry, Helmholtz Center for Environmental Research–UFZ, are also gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shamim, S., Rehman, A. & Qazi, M.H. Cadmium-Resistance Mechanism in the Bacteria Cupriavidus metallidurans CH34 and Pseudomonas putida mt2. Arch Environ Contam Toxicol 67, 149–157 (2014). https://doi.org/10.1007/s00244-014-0009-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00244-014-0009-7