Abstract
Most of the largest vertebrate genomes are found in salamanders, a clade of amphibians that includes 686 species. Salamander genomes range in size from 14 to 120 Gb, reflecting the accumulation of large numbers of transposable element (TE) sequences from all three TE classes. Although DNA loss rates are slow in salamanders relative to other vertebrates, high levels of TE insertion are also likely required to explain such high TE loads. Across the Tree of Life, novel TE insertions are suppressed by several pathways involving small RNA molecules. In most known animals, TE activity in the germline is primarily regulated by the Piwi-interacting RNA (piRNA) pathway. In this study, we test the hypothesis that salamanders’ unusually high TE loads reflect the loss of the ancestral piRNA-mediated TE-silencing machinery. We characterized the small RNA pool in the female and male adult gonads, testing for the presence of small RNA molecules that bear the characteristics of TE-targeting piRNAs. We also analyzed the amino acid sequences of piRNA pathway proteins from salamanders and other vertebrates, testing whether the overall patterns of sequence divergence are consistent with conserved pathway function across the vertebrate clade. Our results do not support the hypothesis of piRNA pathway loss; instead, they suggest that the piRNA pathway is expressed in salamanders. Given these results, we propose hypotheses to explain how the extraordinary TE loads in salamander genomes could have accumulated, despite the expression of TE-silencing machinery.


Similar content being viewed by others
References
Adams MD, Celniker SE, Holt RA, Evans CA, Gocayne JD, Amanatides PG, Scherer SE, Li PW, Hoskins RA, Galle RF, George RA, Lewis SE, Richards S, Ashburner M, Henderson SN, Sutton GG, Wortman JR, Yandell MD, Zhang Q, Chen LX, Brandon RC, Rogers Y-HC, Blazej RG, Champe M, Pfeiffer BD, Wan KH, Doyle C, Baxter EG, Helt G, Nelson CR, Gabor GL, Miklos Abril JF, Agbayani A, An H-J, Andrews-Pfannkoch C, Baldwin D, Ballew RM, Basu A, Baxendale J, Bayraktaroglu L, Beasley EM, Beeson KY, Benos PV, Berman BP, Bhandari D, Bolshakov S, Borkova D, Botchan MR, Bouck J, Brokstein P, Brottier P, Burtis KC, Busam DA, Butler H, Cadieu E, Center A, Chandra I, Cherry JM, Cawley S, Dahlke C, Davenport LB, Davies P, Pablos Bd, Delcher A, Deng Z, Mays AD, Dew I, Dietz SM, Dodson K, Doup LE, Downes M, Dugan-Rocha S, Dunkov BC, Dunn P, Durbin KJ, Evangelista CC, Ferraz C, Ferriera S, Fleischmann W, Fosler C, Gabrielian AE, Garg NS, Gelbart WM, Glasser K, Glodek A, Gong F, Gorrell JH, Gu Z, Guan P, Harris M, Harris NL, Harvey D, Heiman TJ, Hernandez JR, Houck J, Hostin D, Houston KA, Howland TJ, Wei M-H, Ibegwam C, Jalali M, Kalush F, Karpen GH, Ke Z, Kennison JA, Ketchum KA, Kimmel BE, Kodira CD, Kraft C, Kravitz S, Kulp D, Lai Z, Lasko P, Lei Y, Levitsky AA, Li J, Li Z, Liang Y, Lin X, Liu X, Mattei B, McIntosh TC, McLeod MP, McPherson D, Merkulov G, Milshina NV, Mobarry C, Morris J, Moshrefi A, Mount SM, Moy M, Murphy B, Murphy L, Muzny DM, Nelson DL, Nelson DR, Nelson KA, Nixon K, Nusskern DR, Pacleb JM, Palazzolo M, Pittman GS, Pan S, Pollard J, Puri V, Reese MG, Reinert K, Remington K, Saunders RDC, Scheeler F, Shen H, Shue BC, Sidén-Kiamos I, Simpson M, Skupski MP, Smith T, Spier E, Spradling AC, Stapleton M, Strong R, Sun E, Svirskas R, Tector C, Turner R, Venter E, Wang AH, Wang X, Wang Z-Y, Wassarman DA, Weinstock GM, Weissenbach J, Williams SM, Woodage T, Worley KC, Wu D, Yang S, Yao QA, Ye J, Yeh R-F, Zaveri JS, Zhan M, Zhang G, Zhao Q, Zheng L, Zheng XH, Zhong FN, Zhong W, Zhou X, Zhu S, Zhu X, Smith HO, Gibbs RA, Myers EW, Rubin GM, Venter JC (2000) The genome sequence of Drosophila melanogaster. Science 287:2185–2195
AmphibiaWeb (2015) Information on amphibian biology and conservation. Berkeley. http://amphibiaweb.org/
Aravin AA, Sachidanandam R, Girard A, Fejes-Toth K, Hannon GJ (2007) Developmentally regulated piRNA clusters implicate MILI in transposon control. Science 316:744–747
Aravin AA, Sachidanandam R, Bourc’his D, Schaefer C, Pezic D, Toth KF, Bestor T, Hannon GJ (2008) A piRNA pathway primed by individual transposons is linked to de novo DNA methylation in mice. Mol Cell 31:785–799
Armisen J, Gilchrist MJ, Wilczynska A, Standart N, Miska EA (2009) Abundant and dynamically expressed miRNAs, piRNAs, and other small RNAs in the vertebrate Xenopus tropicalis. Genome Res 19:1766–1775
Barrón MG, Fiston-Lavier A-S, Petrov DA, González J (2014) Population genomics of transposable elements in Drosophila. Ann Rev Genet 48:561–581
Blumenstiel JP (2011) Evolutionary dynamics of transposable elements in a small RNA world. Trend Genet 27:23–31
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120
Brennecke J, Aravin AA, Stark A, Dus M, Kellis M, Sachidanandam R, Hannon GJ (2007) Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128:1089–1103
Castel SE, Martienssen RA (2013) RNA interference in the nucleus: roles for small RNAs in transcription, epigenetics and beyond. Nat Rev Gen 14:100–112
Clark JP, Lau NC (2014) Piwi proteins and piRNAs step onto the systems biology stage. In: Yeo GW (ed) Systems biology of RNA binding proteins. Springer, New York, pp 159–197
dos Santos G, Schroeder AJ, Goodman JL, Strelets VB, Crosby MA, Thurmond J, Emmert DB, Gelbart WM, Flybase Consortium (2015) FlyBase: introduction of the Drosophila melanogaster Release 6 reference genome assembly and large-scale migration of genome annotations. Nucleic Acids Res 43:D690–D697
Dumesic PA, Madhani HD (2014) Recognizing the enemy within: licensing RNA-guided genome defense. Trend Biochem Sci 39:25–34
Frahry MB, Sun C, Chong R, Mueller RL (2015) Low levels of LTR retrotransposon deletion by ectopic recombination in the gigantic genomes of salamanders. J Mol Evol 80:120–129
Girard A, Sachidanandam R, Hannon GJ, Carmell MA (2006) A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 442:199–202
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q, Chen Z, Mauceli E, Hacohen N, Gnirke A, Rhind N, di Palma F, Birren BW, Nusbaum C, Lindblad-Toh K, Friedman N, Regev A (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29:644–652
Gregory TR (2004) Insertion-deletion biases and the evolution of genome size. Gene 324:15–34
Gregory TR (2016) Animal Genome Size Database. http://www.genomesize.com.
Grimson A, Srivastava M, Fahey B, Woodcroft BJ, Chiang HR, King N, Degnan BM, Rokhsar DS, Bartel DP (2008) Early origins and evolution of microRNAs and Piwi-interacting RNAs in animals. Nature 455:1193–1197
Gunawardane LS, Saito K, Nishika KM, Kawamura Y, Nagami T, Siomi H, Siomi MC (2007) A slicer-mediated mechanism for repeat-associated siRNA 5′ end formation in Drosophila. Science 315:1587–1590
G-w Chirn, Rahman R, Sytnikova YA, Matts JA, Zeng M, Gerlach D, Yu M, Berger B, Naramura M, Kile BT, Lau NC (2015) Conserved piRNA expression from a distinct set of piRNA cluster loci in eutherian mammals. PLoS Genet 11:e1005652
Han BW, Wang W, Li C, Weng Z, Zamore PD (2015) piRNA-guided transposon cleavage initiates Zucchini-dependent, phased piRNA production. Science 348:817–821
Houwing S, Kamminga LM, Berezikov E, Cronembold D, Girard A, van den Elst H, Filippov DV, Blaser H, Raz E, Moens CB, Plasterk RHA, Hannon GJ, Draper BW, Ketting RF (2007) A role for Piwi and piRNAs in germ gell maintenance and transposon silencing in zebrafish. Cell 129:69–82
Huang Xiao A, Yin H, Sweeney S, Raha D, Snyder M, Lin H (2013) A major epigenetic programming mechanism guided by piRNAs. Dev Cell 24:502–516
Iwasaki YW, Siomi MC, Siomi H (2015) PIWI-interacting RNA: its biogenesis and functions. Annu Rev Biochem 84:405–433
Jockusch EL (1997) An evolutionary correlate of genome size change in plethodontid salamanders. Proc R Soc Lond B 264:597
Keinath MC, Timoshevskiy VA, Timoshevskaya NY, Tsonis PA, Voss SR, Smith JJ (2015) Initial characterization of the large genome of the salamander Ambystoma mexicanum using shotgun and laser capture chromosome sequencing. Sci Rep 5:16413
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:R25
Lau NC, Seto AG, Kim J, Kuramochi-Miyagawa S, Nakano T, Bartel DP, Kingston RE (2006) Characterization of the piRNA complex from rat testes. Science 313:363–367
Lau NC, Ohsumi T, Borowsky M, Kingston RE, Blower MD (2009) Systematic and single cell analysis of Xenopus Piwi-interacting RNAs and Xiwi. EMBO J 28:2945–2958
Laurin M, Canoville A, Struble M, Organ C, de Buffrénil V (2016) Early genome size increase in urodeles. CR Palevol 15(1–2):74–82
Le Thomas A, Rogers AK, Webster A, Marinov GK, Liao SE, Perkins EM, Hur JK, Aravin AA, Tóth KF (2013) Piwi induces piRNA-guided transcriptional silencing and establishment of a repressive chromatin state. Genes Dev 27:390–399
Li C, Vagin VV, Lee S, Xu J, Ma S, Xi H, Seitz H, Horwich MD, Syrzycka M, Honda BM, Kittler ELW, Zapp ML, Klattenhoff C, Schulz N, Theurkauf WE, Weng Z, Zamore PD (2009) Collapse of germline piRNAs in the absence of Argonaute3 reveals somatic piRNAs in flies. Cell 137:509–521
Lu J, Clark AG (2010) Population dynamics of PIWI-interacting RNAs (piRNAs) and their targets in Drosophila. Genome Res 20:212–227
Lynch M (2007) The origins of genome architecture. Sinauer Associates Inc, Sunderland
Lynch M, Conery JS (2003) The origins of genome complexity. Science 302:1401–1404
Malone CD, Hannon GJ (2009) Small RNAs as guardians of the genome. Cell 136:656–668
Malone CD, Brennecke J, Dus M, Stark A, McCombie WR, Sachidanandam R, Hannon GJ (2009) Specialized piRNA pathways act in germline and somatic tissues of the Drosophila ovary. Cell 137:522–535
Marjanović D, Laurin M (2007) Fossils, molecules, divergence times, and the origin of lissamphibians. Syst Biol 56:369–388
Mohlhenrich E, Mueller RL (2016) Genetic drift, mutational hazard, and the evolution of genomic gigantism in salamanders. Evolution. doi:10.1111/evo.13084
Mohn F, Handler D, Brennecke J (2015) piRNA-guided slicing specifies transcripts for Zucchini-dependent, phased piRNA biogenesis. Science 348:812–817
Mueller RL (2006) Evolutionary rates, divergence dates, and the performance of mitochondrial genes in Bayesian phylogenetic analysis. Syst Biol 55:289
Mueller RL, Gregory TR, Gregory SM, Hsieh A, Boore JL (2008) Genome size, cell size, and the evolution of enucleated erythrocytes in attenuate salamanders. Zoology 111:218–230
Notredame C, Higgins D, Heringa J (2000) T-Coffee: a novel method for fast and accurate multiple sequence alignment. J Mol Biol 302:205–217
Organ CL, Canoville A, Reisz RR, Laurin M (2011) Paleogenomic data suggest mammal-like genome size in the ancestral amniote and derived large genome size in amphibians. J Evol Biol 24:372–380
Petrov DA (2002) Mutational equilibrium model of genome size evolution. Theor Popul Biol 61:533–546
Quinlan AR, Hall IM (2010) BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26:841–842
Robine N, Lau NC, Balla S, Jin Z, Okamura K, Kuramochi-Miyagawa S, Blower MD, Lai EC (2009) A broadly conserved pathway generates 3′ UTR-directed primary piRNAs. Curr Biol 19:2066–2076
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542
Roovers EF, Rosenkranz D, Mahdipour M, Han C-T, He N, de Sousa Lopes SMC, van der Westerlaken LA, Zischler H, Butter F, Roelen BA (2015) Piwi proteins and piRNAs in mammalian oocytes and early embryos. Cell Rep 10:2069–2082
Roth G, Nishikawa KC, Wake DB (1997) Genome size, secondary simplification, and the evolution of the brain in salamanders. Brain Behav Evol 50:50–59
Rozhkov NV, Hammell M, Hannon GJ (2013) Multiple roles for Piwi in silencing Drosophila transposons. Genes Dev 27:400–412
Senti K-A, Jurczak D, Sachidanandam R, Brennecke J (2015) piRNA-guided slicing of transposon transcripts enforces their transcriptional silencing via specifying the nuclear piRNA repertoire. Genes Dev 29:1747–1762
Sessions SK (2008) Evolutionary cytogenetics in salamanders. Chromosome Res 16:183–201
Sessions SK, Kezer J (1991) Evolutionary cytogenetics of bolitoglossine salamanders (family Plethodontidae). In: Green DM, Sessions SK (eds) Amphibian cytogenetics and evolution. Academic Press, San Diego, pp 89–130
Sessions SK, Larson A (1987) Developmental correlates of genome size in plethodontid salamanders and their implications for genome evolution. Evolution. doi:10.2307/2409090
Sienski G, Dönertas D, Brennecke J (2012) Transcriptional silencing of transposons by Piwi and maelstrom and its impact on chromatin state and gene expression. Cell 151:964–980
Siomi MC, Sato K, Pezic D, Aravin AA (2011) PIWI-interacting small RNAs: the vanguard of genome defence. Nat Rev Mol Cell Biol 12:246–258
Sun C, Mueller RL (2014) Hellbender genome sequences shed light on genome expansion at the base of crown salamanders. Gen Biol Evol 6:1818–1829
Sun C, Arriaza JRL, Mueller RL (2012a) Slow DNA loss in the gigantic genomes of salamanders. Gen Biol Evol 4:1340–1348
Sun C, Shepard DB, Chong RA, Arriaza JL, Hall K, Castoe TA, Feschotte C, Pollock DD, Mueller RL (2012b) LTR retrotransposons contribute to genomic gigantism in plethodontid salamanders. Gen Biol Evol 4:168–183
Sytnikova YA, Rahman R, G-w Chirn, Clark JP, Lau NC (2014) Transposable element dynamics and PIWI regulation impacts lncRNA and gene expression diversity in Drosophila ovarian cell cultures. Genome Res 24:1977–1990
Szarski H (1983) Cell size and the concept of wasteful and frugal evolutionary strategies. J Theor Biol 105:201–209
Vagin VV, Sigova A, Li C, Seitz H, Gvozdev V, Zamore PD (2006) A distinct small RNA pathway silences selfish genetic elements in the germline. Science 313:320–324
Wang W, Yoshikawa M, Han Bo W, Izumi N, Tomari Y, Weng Z, Zamore Phillip D (2014) The initial uridine of primary piRNAs does not create the tenth adenine that is the hallmark of secondary piRNAs. Mol Cell 56:708–716
Wang W, Han Bo W, Tipping C, Ge Daniel T, Zhang Z, Weng Z, Zamore Phillip D (2015) Slicing and binding by Ago3 or Aub trigger Piwi-bound piRNA production by distinct mechanisms. Mol Cell 59:819–830
Yamanaka S, Siomi M, Siomi H (2014) piRNA clusters and open chromatin structure. Mob DNA 5:22
Yi M, Chen F, Luo M, Cheng Y, Zhao H, Cheng H, Zhou R (2014) Rapid evolution of piRNA pathway in the teleost fish: implication for an adaption to transposon diversity. Gen Biol Evol 6:1393–1407
Zhang Z, Xu J, Koppetsch Birgit S, Wang J, Tipping C, Ma S, Weng Z, Theurkauf William E, Zamore Phillip D (2011) Heterotypic piRNA ping-pong requires qin, a protein with both E3 ligase and Tudor domains. Mol Cell 44:572–584
Zhu W, Kuo D, Nathanson J, Satoh A, Pao GM, Yeo GW, Bryant SV, Voss SR, Gardiner DM, Hunter T (2012) Retrotransposon long interspersed nucleotide element-1 (LINE-1) is activated during salamander limb regeneration. Dev Growth Differ 54:673–685
Acknowledgments
This research was supported by NSF–DBI 1103746 to MM-V and NSF-DEB 1021489 to RLM. NCL was supported by the Searle Scholars Foundation and the NIH (R00HD057298). J. Krakovil and D. Weisrock (U Kentucky) provided the tissues and access to unpublished Cryptobranchus alleganiensis transcriptome data; funding for collecting trips was provided by Highlands Biological Station and the University of Kentucky Department of Biology to J. Krakovil. D. New at IBEST provided critical technical expertise. Suggestions from anonymous reviewers improved the quality of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
M. J. Madison-Villar and Cheng Sun have contributed equally.
An erratum to this article is available at http://dx.doi.org/10.1007/s00239-016-9769-1.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary material 1 (PDF 483 kb)
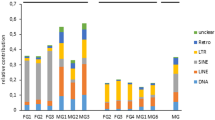
Transposable element families targeted by piRNAs and siRNAs in the female and male samples. Results were obtained from mapping each small RNA dataset to its respective reference transcriptome. TE families are ranked by density of mapped small RNA reads
Supplementary material 2 (PDF 1550 kb)
Phylogenies estimated for eleven piRNA pathway protein sequences obtained from salamanders and other vertebrates
Rights and permissions
About this article
Cite this article
Madison-Villar, M.J., Sun, C., Lau, N.C. et al. Small RNAs from a Big Genome: The piRNA Pathway and Transposable Elements in the Salamander Species Desmognathus fuscus . J Mol Evol 83, 126–136 (2016). https://doi.org/10.1007/s00239-016-9759-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-016-9759-3