Abstract
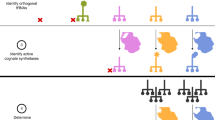
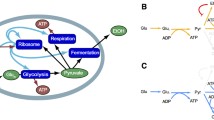
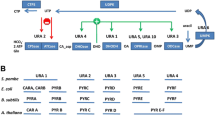
The “cognate bias hypothesis” states that early in evolutionary history the biosynthetic enzymes for amino acid x gradually lost residues of x, thereby reducing the threshold for deleterious effects of x scarcity. The resulting reduction in cognate amino acid composition of the enzymes comprising a particular amino acid biosynthetic pathway is predicted to confer a selective growth advantage on cells. Bioinformatic evidence from protein-sequence data of two bacterial species previously demonstrated reduced cognate bias in amino acid biosynthetic pathways. Here we show that cognate bias in amino acid biosynthesis is present in the other domains of life—Archaebacteria and Eukaryota. We also observe evolutionarily conserved underrepresentations (e.g., glycine in methionine biosynthesis) and overrepresentations (e.g., tryptophan in asparagine biosynthesis) of amino acids in noncognate biosynthetic pathways, which can be explained by secondary amino acid metabolism. Additionally, we experimentally validate the cognate bias hypothesis using the yeast Saccharomyces cerevisiae. Specifically, we show that the degree to which growth declines following amino acid deprivation is negatively correlated with the degree to which an amino acid is underrepresented in the enzymes that comprise its cognate biosynthetic pathway. Moreover, we demonstrate that cognate fold representation is more predictive of growth advantage than a host of other potential growth-limiting factors, including an amino acid’s metabolic cost or its intracellular concentration and compartmental distribution.





Similar content being viewed by others
References
Akashi H, Gojobori T (2002) Metabolic efficiency and amino acid composition in the proteomes of Escherichia coli and Bacillus subtilis. Proc Natl Acad Sci USA 99:3695–3700
Albanesi D, Mansilla MC, Schujman GE, de Mendoza D (2005) Bacillus subtilis cysteine synthetase is a global regulator of the expression of genes involved in sulfur assimilation. J Bacteriol 187:7631–7638
Alves R, Savageau MA (2005) Evidence of selection for low cognate amino acid bias in amino acid biosynthetic enzymes. Mol Microbiol 56:1017–1034
Baudouin-Cornu P, Surdin-Kerjan Y, Marliere P, Thomas D (2001) Molecular evolution of protein atomic composition. Science 293:297–300
Bhattacharjee JK (1985) Alpha-aminoadipate pathway for the biosynthesis of lysine in lower eukaryotes. Crit Rev Microbiol 12:131–151
Bogatyreva NS, Finkelstein AV, Galzitskaya OV (2006) Trend of amino acid composition of proteins of different taxa. J Bioinform Comput Biol 4:597–608
Bono H, Ogata H, Goto S, Kanehisa M (1998) Reconstruction of amino acid biosynthesis pathways from the complete genome sequence. Genome Res 8:203–210
Chasin LA, Magasanik B (1968) Induction and repression of the histidine-degrading enzymes of Bacillus subtilis. J Biol Chem 243:5165–5178
Christensen KE, MacKenzie RE (2006) Mitochondrial one-carbon metabolism is adapted to the specific needs of yeast, plants and mammals. Bioessays 28:595–605
Dufton MJ (1997) Genetic code synonym quotas and amino acid complexity: Cutting the cost of proteins? J Theor Biol 187:165–173
Gene Ontology Consortium (2001) Creating the gene ontology resource: design and implementation. Genome Res 11:1425–1433
Gerstein M, Hegyi H (1998) Comparing genomes in terms of protein structure: surveys of a finite parts list. FEMS Microbiol Rev 22:277–304
Harris MA, et al. (2004) The Gene Ontology (GO) database and informatics resource. Nucleic Acids Res 32[Database Issue]:D258–D261
Heizer EM, Raiford DW, Raymer ML, Doom TE, Miller RV, Krane DE (2006) Amino acid cost and codon-usage biases in 6 prokaryotic genomes: a whole-genome analysis. Mol Biol Evol 23:1670–1680
Hinnebusch AG (2005) Translational regulation of GCN4 and the general amino acid control of yeast. Annu Rev Microbiol 59:407–450
Jain V, Kumar M, Chatterji D (2006) ppGpp: stringent response and survival. J Microbiol 44:1–10
Jones EW, Fink GR (1982) The molecular biology of the yeast Saccharomyces. Metabolism and gene expression. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Karlin S, Bucher P (1992) Correlation analysis of amino acid usage in protein classes. Proc Natl Acad Sci USA 89:12165–12169
Kitamoto K, Yoshizawa K, Ohsumi Y, Anraku Y (1988) Dynamic aspects of vacuolar and cytosolic amino acid pools of Saccharomyces cerevisiae. J Bacteriol 170:2683–2686
Kovach JS, Berberich MA, Venetianer P, Goldberger RF (1969) Repression of the histidine operon: effect of the first enzyme on the kinetics of repression. J Bacteriol 97:1283–1290
Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157:105–132
Mazel D, Marliere P (1989) Adaptive eradication of methionine and cysteine from cyanobacterial light-harvesting proteins. Nature 341:245–248
Messenguy F, Colin D, ten Have JP (1980) Regulation of compartmentation of amino acid pools in Saccharomyces cerevisiae and its effects on metabolic control. Eur J Biochem 108:439–447
Messenguy F, Cooper TG (1977) Evidence that specific and “general” control of ornithine carbamoyltransferease production occurs at the level of transcription in Saccharomyces cerevisiae. J Bacteriol 130:1253–1261
Nelson DL, Cox MM (2000) Lehninger principles of biochemistry, 3rd ed. Worth, New York
Newman EB, Magasanik B (1963) The relation of serine-glycine metabolism to the formation of single-carbon units. Biochim Biophys Acta 78:437–448
Pascal G, Medique C, Danchin A (2006) Persistent biases in the amino acid composition of prokaryotic proteins. Bioessays 28:726–738
Piper MD, Hong SP, Ball GE, Dawes IW (2000) Regulation of the balance of one-carbon metabolism in Saccharomyces cerevisiae. J Biol Chem 275:30987–30995
Seligmann H (2003) Cost-minimization of amino acid usage. J Mol Evol 56:151–161
Stouthamer AH (1973) A theoretical study on the amount of ATP required for synthesis of microbial cell material. Antonie Van Leeuwenhoek 39:545–565
Szentirmai A, Horvath I (1976) Regulation of branched-chain amino acid biosynthesis. Acta Microbiol Acad Sci Hung 23:137–149
Thomas D, Surdin-Kerjan Y (1997) Metabolism of sulfur amino acids in Saccharomyces cerevisiae. Microbiol Mol Biol Rev 61:503–532
Umbarger HE (1978) Amino acid biosynthesis and its regulation. Annu Rev Biochem 47:532–606
Velasco JA, Cansado J, Pena MC, Kawakami T, Laborda J, Notario V (1993) Cloning of the dihydroxyacid dehydratase-encoding gene (ILV3) from Saccharomyces cerevisiae. Gene 137:179–185
Acknowledgments
We are indebted to Drs. Boris Magasanik and Finny Kuruvilla for their constructive discussions. This work was supported by a Merck-Wiley Fellowship (B.L.d.) and by the National Institute of General Medicine Sciences (S. L. S.). S. L. S. is an Investigator at the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Additional information
Reviewing Editor: Dr. Niles Lehman
Ethan O. Perlstein and Benjamin L. de Bivort contributed equally to this work.
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Perlstein, E.O., de Bivort, B.L., Kunes, S. et al. Evolutionarily Conserved Optimization of Amino Acid Biosynthesis. J Mol Evol 65, 186–196 (2007). https://doi.org/10.1007/s00239-007-0013-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-007-0013-x