Abstract
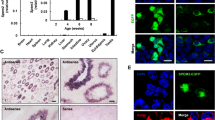
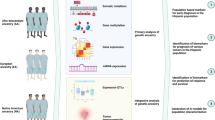
NYD-SP12 is a recently identified spermatogenesis-related gene with a pivotal role in human testis development. In this study, we analyzed between-species divergence and within-species variation of NYD-SP12 in seven representative primate species, four worldwide human populations, and 124 human clinical subjects. Our results indicate that NYD-SP12 evolves rapidly in both the human and the chimpanzee lineages, which is likely caused by Darwinian positive selection and/or sexual selection. We observed significant interpopulation divergence among human populations, which might be due to the varied demographic histories. In the association analysis, we demonstrated significant frequency discrepancy of a synonymous sequence polymorphism among the clinical groups with different sperm traits.


Similar content being viewed by others
References
Altshuler D, Brooks LD, Chakravarti A, Collins FS, Daly MJ, Donnelly P (2005) A haplotype map of the human genome. Nature 437:1299–1320
Anderson MJ, Dixson AF (2002) Sperm competition: motility and the midpiece in primates. Nature 416:496
Cavalli-Sforza LL (1966) Population structure and human evolution. Proc R Soc Lond B Biol Sci 164:362–379
Cavalli-Sforza L, Menozzi P, Piazza A (1994) The history and geography of human genes. Princeton University Press, Princeton, NJ
Civetta A, Singh RS (1995) High divergence of reproductive tract proteins and their association with postzygotic reproductive isolation in Drosophila melanogaster and Drosophila virilis group species. J Mol Evol 41:1085–1095
Comeron JM (1995) A method for estimating the numbers of synonymous and nonsynonymous substitutions per site. J Mol Evol 41:1152–1159
Comeron JM (1999) K-Estimator: calculation of the number of nucleotide substitutions per site and the confidence intervals. Bioinformatics 15(9):763–764
Dorus S, Evans PD, Wyckoff GJ, Choi SS, Lahn BT (2004) Rate of molecular evolution of the seminal protein gene SEMG2 correlates with levels of female promiscuity. Nat Genet 36:1326–1329
Fay JC, Wu CI (2000) Hitchhiking under positive Darwinian selection. Genetics 155:1405–1413
Fu YX, Li WH (1993) Statistical tests of neutrality of mutations. Genetics 133:693–709
Fullerton SM, Bartoszewicz A, Ybazeta G, Horikawa Y, Bell GI, Kidd KK, Cox NJ, Hudson RR, Di Rienzo A (2002) Geographic and haplotype structure of candidate type 2 diabetes susceptibility variants at the calpain-10 locus. Am J Hum Genet 70:1096–1106
Goodman M, Porter CA, Czelusniak J, Page SL, Schneider H, Shoshani J, Gunnell G, Groves CP (1998) Toward a phylogenetic classification of primates based on DNA evidence complemented by fossil evidence. Mol Phylogenet Evol 9:585–598
Harcourt AH, Purvis A, Liles L (1995) Sperm competition: mating system, not breeding season, affects testes size of primates. Funct Ecol 9:468–476
Hughes AL, Nei M (1988) Pattern of nucleotide substitution at major histocompatibility complex class I loci reveals overdominant selection. Nature 335:167–170
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for Molecular Evolutionary Genetics Analysis and sequence alignment. Brief Bioinform 5:150–163
Lewontin RC, Krakauer J (1973) Distribution of gene frequency as a test of the theory of the selective neutrality of polymorphisms. Genetics 74:175–195
Lu L, Lin M, Xu M, Zhou ZM, Sha JH (2006) Gene functional research using polyethylenimine–mediated in vivo gene transfection into mouse spermatogenic cells. Asian J Androl 8:53–59
Majewski J, Cohan FM (1999) Adapt globally, act locally: the effect of selective sweeps on bacterial sequence diversity. Genetics 152:1459–1474
McDonald JH, Kreitman M (1991) Adaptive protein evolution at the Adh locus in Drosophila. Nature 351:652–654
Messier W, Stewart CB (1997) Episodic adaptive evolution of primate lysozymes. Nature 385:151–154
Nielsen R (2005) Molecular signatures of natural selection. Annu Rev Genet 39:197–218
Nielsen R, Bustamante C, Clark AG, Glanowski S, Sackton TB, Hubisz MJ, Fledel-Alon A, Tanenbaum DM, Civello D, White TJ, Adams MD, Cargill M (2005) A scan for positively selected genes in the genomes of humans and chimpanzees. PLoS Biol 3:e170
Page SL, Goodman M (2001) Catarrhine phylogeny: noncoding DNA evidence for a diphyletic origin of the mangabeys and for a human-chimpanzee clade. Mol Phylogenet Evol 18:14–25
Podlaha O, Zhang J (2003) Positive selection on protein-length in the evolution of a primate sperm ion channel. Proc Natl Acad Sci USA 100:12241–12246
Rooney AP, Zhang J (1999) Rapid evolution of a primate sperm protein: Relaxation of functional constraint or positive Darwinian selection? Mol Biol Evol 16:706–710
Rozas J, Rozas R (1999) DnaSP version 3:an integrated program for molecular population genetics and molecular evolution analysis. Bioinformatics 15:174–175
Slatkin M, Wiehe T (1998) Genetic hitch-hiking in a subdivided population. Genet Res 71:155–160
Swanson WJ, Vacquier VD (2002) The rapid evolution of reproductive proteins. Nat Rev Genet 3:137–144
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Voight BF, Kudaravalli S, Wen X, Pritchard JK (2006) A map of recent positive selection in the human genome. PLoS Biol 4:e72
Wang X, Zhang J (2004) Rapid evolution of mammalian X-linked testis-expressed homeobox genes. Genetics 167:879–888
World Health Organization (1999) WHO laboratory manual for the examination of human semen and sperm-cervical mucus interaction. Cambridge University Press, Cambridge
Wright S (1950) Genetical structure of populations. Nature 166:247–249
Wyckoff GJ, Wang W, Wu CI (2000) Rapid evolution of male reproductive genes in the descent of man. Nature 403:304–309
Xu M, Xiao J, Chen J, Li J, Yin L, Zhu H, Zhou Z, Sha J (2003) Identification and characterization of a novel human testis-specific Golgi protein, NYD-SP12. Mol Hum Reprod 9:9–17
Yang Z (1997) PAML: a program package for phylogenetic analysis by maximum likelihood. Comput Appl Biosci 13:555–556
Yang Z (1998) Likelihood ratio tests for detecting positive selection and application to primate lysozyme evolution. Mol Biol Evol 15:568–573
Yang Z, Nielsen R (2002) Codon-substitution models for detecting molecular adaptation at individual sites along specific lineages. Mol Biol Evol 19:908–917
Yang Z, Swanson WJ (2002) Codon-substitution models to detect adaptive evolution that account for heterogeneous selective pressures among site classes. Mol Biol Evol 19:49–57
Yang Z, Wong WS, Nielsen R (2005) Bayes empirical Bayes inference of amino acid sites under positive selection. Mol Biol Evol 22:1107–1118
Zhang J, Nielsen R, Yang Z (2005) Evaluation of an improved branch-site likelihood method for detecting positive selection at the molecular level. Mol Biol Evol 22:2472–2479
Acknowledgments
This study was supported by grants from the Chinese Academy of Sciences (KSCX1-YW-R-34), the National Natural Science Foundation of China (30370755, 30525028, 30630013), the Natural Science Foundation of Yunnan Province of China, and the National 973 Project of China (2006CB701506). We thank Hui Zhang and Yi-chuan Yu for technical help.
Author information
Authors and Affiliations
Corresponding author
Additional information
[Reviewing Editor: Dr. Manyuan Long]
Rights and permissions
About this article
Cite this article
Zhang, Q., Zhang, F., Chen, Xh. et al. Rapid Evolution, Genetic Variations, and Functional Association of the Human Spermatogenesis-Related Gene NYD-SP12 . J Mol Evol 65, 154–161 (2007). https://doi.org/10.1007/s00239-006-0127-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-006-0127-6