Abstract
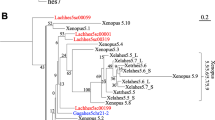
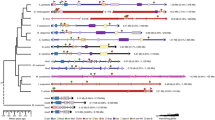
The statistical analysis of phylogenetic footprints in the two known horn shark Hox clusters and the four mammalian clusters shows that the shark HoxN cluster is HoxD-like. This finding implies that the most recent common ancestor of jawed vertebrates had at least four Hox clusters, including those which are orthologous to the four mammalian Hox clusters.



Similar content being viewed by others
References
SF Altschul W Gish W Miller EW Myers DJ Lipman (1990) ArticleTitleBasic local alignment search tool. J Mol Biol 215 403–410 Occurrence Handle10.1006/jmbi.1990.9999 Occurrence Handle1:CAS:528:DyaK3MXitVGmsA%3D%3D Occurrence Handle2231712
A Amores A Force YL Yan et al. (1998) ArticleTitleZebrafish hox clusters and vertebrate genome evolution. Science 282 1711–1714 Occurrence Handle1:CAS:528:DyaK1cXnslGgtrY%3D Occurrence Handle9831563
S Aparicio J Chapman E Stupka et al. (2002) ArticleTitleWhole-genome shotgun assembly and analysis of the genome of Fugu rubripes. Science 297 1301–1310 Occurrence Handle10.1126/science.1072104 Occurrence Handle1:CAS:528:DC%2BD38Xms1ejtr8%3D Occurrence Handle12142439
WJ Bailey J Kim G Wagner FH Ruddle (1997) ArticleTitlePhylogenetic reconstruction of vertebrate Hox cluster duplications. Mol Biol Evol 14 843–853 Occurrence Handle1:CAS:528:DyaK2sXltVeisbw%3D Occurrence Handle9254922
HJ Bandelt AWM Dress (1992) ArticleTitleA canonical decomposition theory for metrics on a finite set. Adv Math 92 47
HJ Bandelt AWM Dress (1993) A relational approach to split decomposition. O Opitz B Lausen R Klar (Eds) Information and classification. Springer-Verlag Berlin 123–131
P Buneman (1971) The recovery of trees from measures of dissimilarity. FR Hodson DG Kendall P Tauto (Eds) Mathematics and the archeological and historical sciences. Edinburgh University Press Edinburgh, UK 387–395
Ch Chiu C Amemiya K Dewar CB Kim FH Ruddle GP Wagner (2002) ArticleTitleMolecular evolution of the HoxA cluster in the three major gnathostome lineages. Proc Natl Acad Sci USA 99 5492–5497 Occurrence Handle10.1073/pnas.052709899 Occurrence Handle1:CAS:528:DC%2BD38XjtFKltrw%3D
DOE Joint Genome Institute (2002) Fugu genome database. version 2.0: http://genome.jgi-psf.org/fugu3/fugu3.home.html ; version 3.0: http://genome.jgi-psf.org/fugu6/fugu6.home.html
L Duret P Bucher (1997) ArticleTitleSearching for regulatory elements in human noncoding sequences. Curr Opin Struct Biol 7 399–406
J Felsenstein (1989) ArticleTitlePHYLIP—Phylogeny inference package (version 3.2). Cladistics 5 164–166
J Garcia-Fernández PW Holland (1994) ArticleTitleArchetypal organization of the amphioxus hox gene cluster. Nature 370 563–566 Occurrence Handle1:CAS:528:DyaK2cXmslensLY%3D Occurrence Handle7914353
PW Holland J Garcia-Fernández (1999) ArticleTitleHox genes and chordate evolution. Dev Biol 173 382–395 Occurrence Handle10.1006/dbio.1996.0034
PWH Holland J Garcia-Fernández NA Williams A Sidow (1994) ArticleTitleGene duplication and the origins of vertebrate development. Development . IssueID(Suppl) 125–133
DH Huson (1998) ArticleTitleSplitstree: Analyzing and visualizing evolutionary data. Bioinformatics 14 68–73
C Kappen K Schughart FJH Ruddle (1989) ArticleTitleTwo steps in the evolution of antennapedia-class vertebrate homeobox genes. Proc Natl Acad Sci USA 86 5459–5463 Occurrence Handle1:CAS:528:DyaL1MXkvFWmu74%3D Occurrence Handle2568634
CB Kim C Amemiya W Bailey K Kawasaki J Mezey W Miller S Minosima N Shimizu GP Wagner GP Ruddle (2000) ArticleTitleHox cluster genomics in the horn shark, Heterodontus francisci. Proc Nat Acad Sci USA 97 1655–1660 Occurrence Handle10.1073/pnas.030539697 Occurrence Handle1:CAS:528:DC%2BD3cXhsFCrsbg%3D Occurrence Handle10677514
DR Maddison WP Maddison (2000) MacClade 4: Analysis of phylogeny and character evolution. Sinauer Associates Sunderland, MA (e-book and computer program)
E Málaga-Trillo A Meyer (2001) ArticleTitleGenome duplications and accelerated evolution of Hox genes and cluster architecture in teleost fishes. Am Zool 41 6767–686
W McGinnis R Krumlauf (1992) ArticleTitleHomeobox genes and axial patterning. Cell 68 283–302 Occurrence Handle1:CAS:528:DyaK38XhtVShs7o%3D Occurrence Handle1346368
B Morgenstern (1999) ArticleTitleDIALIGN 2: Improvement of the segment-to-segment approach to multiple sequence alignment. Bioinformatics 15 211–218 Occurrence Handle10.1093/bioinformatics/15.3.211 Occurrence Handle1:CAS:528:DyaK1MXjs1artrc%3D Occurrence Handle10222408
Prohaska S, Fried C, Flamm C, Wagner G, Stadler PF (2003) Surveying phylogenetic footprints, in large gene clusters: Applications to Hox cluster duplications. Mol Phylog Evol (in press)
N Saitou M Nei (1987) ArticleTitleThe neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol Biol Evol 4 406–425 Occurrence Handle1:STN:280:BieC1cbgtVY%3D Occurrence Handle3447015
C Semple M Steel (2003) Phylogenetics. Oxford University Press Oxford
K Sumiyama C Kim FH Ruddle (2001) ArticleTitleAn efficient cis-element discovery method using multiple sequence comparisons based on evolutionary relationships. Genomics 71 260–262 Occurrence Handle10.1006/geno.2000.6422 Occurrence Handle1:CAS:528:DC%2BD3MXptleqtw%3D%3D Occurrence Handle11161821
DA Tagle BF Koop M Goodman JL Slightom DL Hess RT Jones (1988) ArticleTitleEmbryonic epsilon and gamma globin genes of a prosimian primate (galago crassicaudatus). Nucleotide and amino acid sequences, developmental regulation and phylogenetic footprints. J Mol Biol 203 439–455 Occurrence Handle1:CAS:528:DyaL1MXktlCnsrs%3D Occurrence Handle3199442
JD Thompson DG Higgs TJ Gibson (1994) ArticleTitleCLUSTALW: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position specific gap penalties, and weight matrix choice. Nucleic Acids Res 22 4673–4680 Occurrence Handle7984417
Acknowledgements
Funding for this research is gratefully acknowledged: DFG Bioinformatics Initiative BIZ-6/1-2 to S.J.P., C.F., and P.F.S., NSF IBN-9905408 to F.R. and IBN-0321470 to G.P.W.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Prohaska, S.J., Fried, C., Amemiya, C.T. et al. The Shark HoxN Cluster Is Homologous to the Human HoxD Cluster . J Mol Evol 58, 212–217 (2004). https://doi.org/10.1007/s00239-003-2545-z
Issue Date:
DOI: https://doi.org/10.1007/s00239-003-2545-z