Abstract
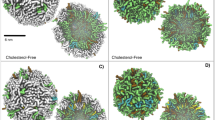
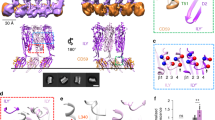
Pore-forming toxins are proteins expressed by bacteria to primarily cause infections in the host cell. Cholesterol-dependent cytolysins (CDCs) are a class of proteins whose pore-forming ability requires the presence of cholesterol in the membrane. Upon binding to the target cell, cholesterol-recognizing residues in the membrane binding D4 subdomain assist in stabilizing both the pre-pore and pore states which occur during protein oligomerization on the cell membrane. Super resolution-stimulated emission depletion (STED) microscopy experiments (Sarangi et al. in Langmuir, 32:9649–9657, 2016) on supported lipid bilayers have shown that listeriolysin (LLO), a CDC expressed by Listeria monocytogenes, a food-borne pathogen, induces both spatial and dynamic heterogeneity in bilayer membranes. Here, we use all-atom molecular dynamics simulations to explore molecular details of the induced membrane reorganization by considering two distinct states of the oligomerized LLO protein in a 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC): cholesterol membrane. In the membrane bound (MB) state, four D4 subunits are placed at the bilayer interface in a pre-pore configuration and the membrane-inserted (MI) state consists of a tetrameric arc-like pore configuration. By analyzing lipid-order parameters, mobilities, and diffusion coefficients, we examine the induced spatial heterogeneity that occurs in both the MB and MI states. This heterogeneity is primarily driven by the local density enhancement of cholesterol in the vicinity of the MB, D4 subunits leading to distinct differences in lipid and cholesterol mobility across the two leaflets as well as enhanced lipid mobilities in regions where cholesterol is depleted. The leaflet-induced heterogeneity is greater for the MB state when compared with the MI state and the dynamic variations are more pronounced in the extracellular leaflet when compared with the cytosolic leaflet. Our study provides molecular-level insights into the inhomogeneity and perturbation induced in bilayer membranes upon LLO binding and pore formation and is expected to represent trends across PFTs in the broad CDC subclass of proteins.
Graphic Abstract










Similar content being viewed by others
References
Adzhubei AA, Sternberg MJ, Makarov AA (2013) Polyproline-II helix in proteins: structure and function. J Mol Biol 425(12):2100–2132
Alwarawrah M, Dai J, Huang J (2010) A molecular view of the cholesterol condensing effect in DOPC lipid bilayers. J Phys Chem B 114(22):7516–7523
Andersen HC (1983) Rattle: a “velocity” version of the shake algorithm for molecular dynamics calculations. J Comput Phys 52(1):24–34
Bavdek A, Gekara NO, Priselac D, Gutiérrez Aguirre I, Darji A, Chakraborty T, Macek P, Lakey JH, Weiss S, Anderluh G (2007) Sterol and pH interdependence in the binding, oligomerization, and pore formation of Listeriolysin O. Biochemistry 46(14):4425–4437
Bavdek A, Kostanjšek R, Antonini V, Lakey JH, Dalla Serra M, Gilbert RJ, Anderluh G (2012) pH dependence of listeriolysin O aggregation and pore-forming ability. FEBS J 279(1):126–141
Benke S, Roderer D, Wunderlich B, Nettels D, Glockshuber R, Schuler B (2015) The assembly dynamics of the cytolytic pore toxin ClyA. Nat Commun 6:6198
Bhakdi S, Tranum-Jensen J, Sziegoleit A (1985) Mechanism of membrane damage by streptolysin O. Infect Immun 47(1):52–60
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an n log (n) method for Ewald sums in large systems. J Chem Phys 98(12):10089–10092
De S, Olson R (2011) Crystal structure of the vibrio cholerae cytolysin heptamer reveals common features among disparate pore-forming toxins. Proc Natl Acad Sci 108(18):7385–7390
Desikan R, Maiti PK, Ayappa KG (2017) Assessing the structure and stability of transmembrane oligomeric intermediates of an \(\alpha\)-helical toxin. Langmuir 33(42):11496–11510
Desikan R, Padmanabhan P, Ayappa KG (2020) Opening of smaller toxin pores by lipid micelle formation. Proc Natl Acad Sci 117(10):5107–5108
Duncan AL, Corey RA, Sansom MSP (2020) Defining how multiple lipid species interact with inward rectifier potassium (Kir2) channels. Proceedings of the National Academy of Sciences of the United States of America
Faraj BHA, Collard L, Cliffe R, Blount LA, Lonnen R, Wallis R, Andrew PW, Hudson AJ (2020) Formation of pre-pore complexes of pneumolysin is accompanied by a decrease in short-range order of lipid molecules throughout vesicle bilayers. Sci Rep 10(1):4585
Farrand AJ, LaChapelle S, Hotze EM, Johnson AE, Tweten RK (2010) Only two amino acids are essential for cytolytic toxin recognition of cholesterol at the membrane surface. Proc Natl Acad Sci 107(9):4341–4346. https://doi.org/10.1073/pnas.0911581107
Flenner E, Das J, Rheinstädter MC, Kosztin I (2009) Subdiffusion and lateral diffusion coefficient of lipid atoms and molecules in phospholipid bilayers. Phys Rev E 79:011907
Gekara N, Weiss S (2004) Lipid rafts clustering and signalling by listeriolysin O. Biochem Soc Trans 32(5):712–714
Gekara NO, Jacobs T, Chakraborty T, Weiss S (2005) The cholesterol-dependent cytolysin listeriolysin O aggregates rafts via oligomerization. Cell Microbiol 7(9):1345–1356
Gerber C, Lang HP (2006) How the doors to the nanoworld were opened. Nat Nanotechnol 1(1):3–5
Gonzalez MR, Bischofberger M, Pernot L, van der Goot FG, Freche B (2008) Bacterial pore-forming toxins: the (w) hole story? Cell Mol Life Sci 65(3):493–507
Gouaux E (1997) Channel-forming toxins: tales of transformation. Curr. Opin. Struct. Biol. 7(4):566–573
Guex N, Peitsch MC, Schwede T (2009) Automated comparative protein structure modeling with SWISS-MODEL and Swiss-PdbViewer: a historical perspective. Electrophoresis 30(S1):S162–S173
Heuck AP, Moe PC, Johnson BB (2010) The cholesterol-dependent cytolysin family of gram-positive bacterial toxins. In: Cholesterol Binding and cholesterol transport proteins, pp. 551–577. Springer
Heuck AP, Tweten RK, Johnson AE (2003) Assembly and topography of the prepore complex in cholesterol-dependent cytolysins. J Biol Chem 278(33):31218–31225
Hoover WG (1985) Canonical dynamics: equilibrium phase-space distributions. Phys Rev A 31(3):1695–1697
Hotze EM, Tweten RK (2012) Membrane assembly of the cholesterol-dependent cytolysin pore complex. Biochim Biophys Acta (Biomembranes) 1818(4):1028–1038
Husmann M, Beckmann E, Boller K, Kloft N, Tenzer S, Bobkiewicz W, Neukirch C, Bayley H, Bhakdi S (2009) Elimination of a bacterial pore-forming toxin by sequential endocytosis and exocytosis. FEBS Lett 583:337–344
Iacovache I, van der Goot FG, Pernot L (2008) Pore formation: an ancient yet complex form of attack. Biochim Biophys Acta 1778(7–8):1611–1623
Ponmalar Ilanila I, Cheerla R, Ayappa KG, Basu JK (2019) Correlated protein conformational states and membrane dynamics during attack by pore-forming toxins. Proc Natl Acad Sci USA 116(26):12839–12844
Jämbeck JP, Lyubartsev AP (2012) Derivation and systematic validation of a refined all-atom force field for phosphatidylcholine lipids. J Phys Chem B 116(10):3164–3179
Jo S, Kim T, Iyer VG, Im W (2008) Charmm-gui: a web-based graphical user interface for charmm. J Comput Chem 29(11):1859–1865
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79(2):926–935
Tilley JS, Orlova EV, Gilbert RJC, Andrew PW, Saibil HR (2005) Structural basis of pore formation by the bacterial toxin pneumolysin. Cell 121(2):247–256
Keyel P, Loultcheva L, Roth R, Salter R, Watkins S, Yokoyama W, Heuser J (2011) Streptolysin O clearance through sequestration into blebs that bud passively from the plasma membrane. J Cell Sci 124:2414–2423
Kiefer F, Arnold K, Künzli M, Bordoli L, Schwede T (2008) The SWISS-MODEL repository and associated resources. Nucleic Acids Res 37:D387–D392
Kneller GR, Baczynski K, Pasenkiewicz-Gierula M (2011) Communication: consistent picture of lateral subdiffusion in lipid bilayers: molecular dynamics simulation and exact results. J Chem Phys 135:141105
Köster S, Van Pee K, Hudel M, Leustik M, Rhinow D, Kühlbrandt W, Chakraborty T, Yildiz Ö (2014) Crystal structure of listeriolysin O reveals molecular details of oligomerization and pore formation. Nat Commun 5:3690
Lacovache L, Bischofberger M, van der Goot FG (2010) Structure and assembly of pore-forming proteins. Curr Opin Struct Biol 20(2):241–246
Lesieur C, Vécsey-Semjén B, Abrami L, Fivaz M, van der Goot FG (1997) Membrane insertion: the strategies of toxins. Mol Membrane Biol 14(2):45–64
Leung C, Dudkina NV, Lukoyanova N, Hodel AW, Farabella I, Pandurangan AP, Jahan N, Damaso MP, Osmanović D, Reboul CF et al (2014) Stepwise visualization of membrane pore formation by suilysin, a bacterial cholesterol-dependent cytolysin. Elife 3:e04247
Los FC, Randis TM, Aroian RV, Ratner AJ (2013) Role of pore-forming toxins in bacterial infectious diseases. Microbiol Mol Biol Rev 77(2):173–207
Mondal AK, Chattopadhyay K (2019) Taking toll on membranes: curious cases of bacterial \(\beta\)-barrel pore-forming toxins. Biochemistry 59(2):163–170
Morton CJ, Sani MA, Parker MW, Separovic F (2019) Cholesterol-dependent cytolysins: membrane and protein structural requirements for pore formation: Focus review. Chem Rev 119(13):7721–7736
Mueller M, Grauschopf U, Maier T, Glockshuber R, Ban N (2009) The structure of a cytolytic \(\alpha\)-helical toxin pore reveals its assembly mechanism. Nature 459(7247):726
Mulvihill E, van Pee K, Mari SA, Müller DJ, Yildiz Ö (2015) Directly observing the lipid-dependent self-assembly and pore-forming mechanism of the cytolytic toxin listeriolysin O. Nano Lett 15(10):6965–6973
Nosé S (1984) A unified formulation of the constant temperature molecular dynamics methods. J Chem Phys 81(1):511–519
Palmer M, Harris R, Freytag C, Kehoe M, Tranum-Jensen J, Bhakdi S (1998) Assembly mechanism of the oligomeric streptolysin o pore: the early membrane lesion is lined by a free edge of the lipid membrane and is extended gradually during oligomerization. EMBO J 17(6):1598–1605
Parrinello M, Rahman A (1981) Polymorphic transitions in single crystals: a new molecular dynamics method. J App. Phys 52(12):7182–7190
van Pee K, Alexander N, Edoardo D, Deryck JM, Kühlbrandt Yildiz, Ö (2017) CryoEM structures of membrane pore and prepore complex reveal cytolytic mechanism of pneumolysin. Elife 6:e23644
Peraro MD, Goot FGVD (2016) Pore-forming toxins: ancient, but never really out of fashion. Nat Rev Microbiol 14(2):77–92
Podobnik M, Marchioretto M, Bavdek A, Kisovec M, Cajnko M, Lunelli L, Dalla Serra M, Anderluh G (2015) Plasticity of listeriolysin O pores revealed by mutagenesis of a unique histidine. Sci Rep 5:9623–9633
Podobnik M, Marchioretto M, Zanetti M, Bavdek A, Kisovec M, Cajnko MM, Lunelli L, Dalla Serra M, Anderluh G (2015) Plasticity of listeriolysin O pores and its regulation by ph and unique histidine. Sci Rep 5:9623
Prasanna X, Jafurulla M, Sengupta D, Chattopadhyay A (2016) The ganglioside GM1 interacts with the serotonin1A receptor via the sphingolipid binding domain. Biochim Biophys Act Biomembr. 1858:2818–2826
Pronk S, Páll S, Schulz R, Larsson P, Bjelkmar P, Apostolov R, Shirts MR, Smith JC, Kasson PM, Van Der Spoel D et al (2013) Gromacs 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29(7):845–854
Ramachandran R, Tweten RK, Johnson AE (2005) The domains of a cholesterol-dependent cytolysin undergo a major FRET-detected rearrangement during pore formation. Proc Natl Acad Sci USA 102(20):7139–7144
Rao HG, Desikan R, Ayappa KG, Gosavi S (2016) Capturing the membrane-triggered conformational transition of an \(\alpha\)-helical pore-forming toxin. J Phys Chem B 120(47):12064–12078
Repp H, Pamukçi Z, Koschinski A, Domann E, Darji A, Birringer J, Brockmeier D, Chakraborty T, Dreyer F (2002) Listeriolysin of listeria monocytogenes forms Ca\(^{2+}\)-permeable pores leading to intracellular Ca\(^{2+}\) oscillations. Cell Microbiol 4(8):483–491
Ruan Y, Rezelj S, Zavec AB, Anderluh G, Scheuring S (2016) Listeriolysin O membrane damaging activity involves arc formation and lineaction-implication for listeria monocytogenes escape from phagocytic vacuole. PLoS Pathog 12(4):e1005597
Sarangi NK, Ayappa KG, Visweswariah SS, Basu JK (2016) Super-resolution stimulated emission depletion-fluorescence correlation spectroscopy reveals nanoscale membrane reorganization induced by pore-forming proteins. Langmuir 32(37):9649–9657
Sarangi NK, Basu JK (2018) Pathways for creation and annihilation of nanoscale biomembrane domains reveal \(\alpha\) and \(\beta\)-toxin nanopore formation processes. Phys Chem Chem Phys 20(46):29116–29130
Sarangi NK, Roobala C, Basu JK (2018) Unraveling complex nanoscale lipid dynamics in simple model biomembranes: Insights from fluorescence correlation spectroscopy in super-resolution stimulated emission depletion mode. Methods 140–141:198–211
Sathyanarayana P, Desikan R, Ayappa KG, Visweswariah SS (2016) The solvent-exposed C-terminus of the cytolysin a pore-forming toxin directs pore formation and channel function in membranes. Biochemistry 55(42):5952–5961
Sathyanarayana P, Maurya S, Behera A, Ravichandran M, Visweswariah SS, Ayappa KG, Roy R (2018) Cholesterol promotes cytolysin a activity by stabilizing the intermediates during pore formation. Proc Natl Acad Sci USA 115(31):E7323–E7330
Savinov SN, Heuck AP (2017) Interaction of cholesterol with perfringolysin O: what have we learned from functional analysis? Toxins 9(12):381
Schnupf P, Portnoy DA, Decatur AL (2006) Phosphorylation, ubiquitination and degradation of listeriolysin O in mammalian cells: role of the pest-like sequence. Cell Microbiol 8(2):353–364
Schuerch DW, Wilson-Kubalek EM, Tweten RK (2005) Molecular basis of listeriolysin O pH dependence. Proc Natl Acad Sci USA 102(35):12537–12542
Shatursky O, Heuck AP, Shepard LA, Rossjohn J, Parker MW, Johnson AE, Tweten RK (1999) The mechanism of membrane insertion for a cholesterol-dependent cytolysin: a novel paradigm for pore-forming toxins. Cell 99(3):293–299
Shepard LA, Shatursky O, Johnson AE, Tweten RK (2000) The mechanism of pore assembly for a cholesterol-dependent cytolysin: formation of a large prepore complex precedes the insertion of the transmembrane \(\beta\)-hairpins. Biochemistry 39(33):1028410293
Shepard LA, Shatursky O, Johnson AE, Tweten RK (2000) The mechanism of pore assembly for a cholesterol-dependent cytolysin: formation of a large prepore complex precedes the insertion of the transmembrane \(\beta\)-hairpins. Biochemistry 39(33):10284–10293
Soltani CE, Hotze EM, Johnson AE, Tweten RK (2007) Specific protein-membrane contacts are required for prepore and pore assembly by a cholesterol-dependent cytolysin. J Biol Chem 282(21):15709–15716
Sonnen AFP, Plitzko JM, Gilbert RJ (2014) Incomplete pneumolysin oligomers form membrane pores. Open Biol 4(4):140044
Sun R, Liu Y (2013) Listeriolysin O as a strong immunogenic molecule for the development of new anti-tumor vaccines. Hum Vaccin Immunother 9(5):1058–1068
Tilley SJ, Saibil HR (2006) The mechanism of pore formation by bacterial toxins. Curr Opin Struct Biol 16(2):230–236
Tweten RK (2005) Cholesterol-dependent cytolysins, a family of versatile pore-forming toxins. Infect Immun 73(10):6199–6209
Tweten RK, Hotze EM, Wade KR (2015) The unique molecular choreography of giant pore formation by the cholesterol-dependent cytolysins of gram-positive bacteria. Annu Rev Microbiol 69:323–340
Tweten RK, Parker MW, Johnson AE (2001) The cholesterol-dependent cytolysins. In: Pore-forming toxins, pp. 15–33. Springer
Vadia S, Arnett E, Haghighat AC, Wilson-Kubalek EM, Tweten RK, Seveau S (2011) The pore-forming toxin listeriolysin O mediates a novel entry pathway of L. monocytogenes into human hepatocytes. PLOS Pathog 7(11):e1002356
Varadarajan V, Dasgupta C, Ayappa KG (2018) Influence of surface commensurability on the structure and relaxation dynamics of a confined monatomic fluid. J Chem Phys 149(6):064503
Varadarajan V, Desikan R, Ayappa KG (2020) Assessing the extent of the structural and dynamic modulation of membrane lipids due to pore forming toxins: insights from molecular dynamics simulations. Soft Matter 16(20):4840–4857
Vögele M, Bhaskara RM, Mulvihill E, van Pee K, Yildiz Ö, Kühlbrandt W, Müller DJ, Hummer G (2019) Membrane perforation by the pore-forming toxin pneumolysin. Proc Natl Acad Sci 116(27):13352–13357
Vögele M, Köfinger J, Hummer G (2018) Hydrodynamics of diffusion in lipid membrane simulations. Phys Rev Lett 120(26):268104
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) Development and testing of a general AMBER force field. J Comput Chem 25(9):1157–1174
Waterhouse A, Bertoni M, Bienert S, Studer G, Tauriello G, Gumienny R, Heer FT, de Beer TAP, Rempfer C, Bordoli L et al (2018) SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res 46(W1):W296–W303
Wolfmeier H, Radecke J, Schoenauer R, Koeffel R, Babiychuk VS, Drücker P, Hathaway LJ, Mitchell TJ, Zuber B, Draeger A et al (2016) Active release of pneumolysin prepores and pores by mammalian cells undergoing a streptococcus pneumoniae attack. Biochim Biophys Acta (General Subj) 1860(11):2498–2509
Acknowledgements
This work was supported by the Department of Science and Technology, Science and Engineering Research Board. We thank the Supercomputer Education and Research Center, Indian Institute of Science for computational facilities. R.C. acknowledges University Grants Commission for the Dr. D. S. Kothari postdoctoral fellowship. We also thank Ilanila I. P. and Jaydeep Basu for several useful discussions during the course of this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Rights and permissions
About this article
Cite this article
Cheerla, R., Ayappa, K.G. Molecular Dynamics Study of Lipid and Cholesterol Reorganization Due to Membrane Binding and Pore Formation by Listeriolysin O. J Membrane Biol 253, 535–550 (2020). https://doi.org/10.1007/s00232-020-00148-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-020-00148-9