Abstract
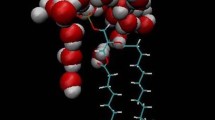
Lipid membrane interfaces are complex environments that host essential cellular processes such as binding of proteins or drug molecules. A key open question is how water molecules at the interface of membranes with anionic lipids participate in protein binding. To address this question, we studied the dynamics of water hydrogen bonding at the interface of membranes composed of phosphatidylcholine and phosphatidylglycerol, and implemented an algorithm to identify hydrogen-bonded networks at the interface of a lipid membrane, and to characterize their dynamics and linear connections. We find that the membrane interface is characterized by a rich network of hydrogen-bonded water chains that bridge lipid headgroups, some of which form transient lipid clusters. Water-mediated bridges between with lipid phosphate groups are dynamic, with residence lifetimes on the order of picoseconds. These clusters of water/lipid headgroup hydrogen bonds could provide a platform for the binding of proteins or of drug molecules with cationic groups.








Similar content being viewed by others
References
Agmon N, Gutman M (2011) Proton fronts on membranes. Nat Chem 3:840–842
Alami M, Dalal K, Lelj-Garolla B, Sligar SG, Duong F (2007) Nanodiscs reveal the interaction between the SecYEG channel and its cytosolic partner SecA. EMBO J 26:1995–2004
Bankaitis VA, Mousley CJ, Schaaf G (2009) The Sec14 superfamiliy and mechanisms for crosstalk between lipid metabolism and lipid signaling. Trends Biochem Sci 35:150–160
Berkowitz ML, Bostik DL, Pandit S (2006) Aqueous solution next to phospholipid membrane interfaces: Insights from simulations. Chem Rev 106:1527–1539
Bhide SY, Berkowitz ML (2005) Structure and dynamics of water at the interface with phospholipid bilayers. J Chem Phys 123:224702
Böckmann RA, Grubmüller H (2004) Multistep binding of divalent cations to phospholipid bilayers: a molecular dynamics study. Angew Chem Int Ed 43:1021–1024
Bondar A-N (2016) Biophysical mechanism of rhomboid proteolysis: setting a foundation for therapeutics. Seminars Cell Dev Biol 60:46–51
Bondar A-N, White SH (2012) Hydrogen bond dynamics in membrane protein function. Biochim Biophys Acta 1818:942–950
Brooks BR, Bruccoleri RE, Olafson BD, States DJ, Swaminathan S, Karplus M (1983) CHARMM: a program for macromolecular energy, minimization, and dynamics calculations. J Comput Chem 4:187–217
Connor J, Pak CC, Schroit AJ (1994) Exposure of phosphatidylserine in the outer leaflet of human red blood cells. J Biol Chem 269:2399–2404
Cormen TH, Leiserson CE, Rivest RL, Sten C (2009). Introduction to algorithms, 3rd edn. Massachusetts Institute of Technology
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N x log(N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092
Dicko A, Bourque H, Pézolet M (1998) Study by infrared spectroscopy of the conformation of dipalmitoylphospatidylglycerol monolayers at the air-water interface and transferred on solid substrates. Chem Phys Lipids 96:125–139
Ebbinghaus S, Kim SJ, Heyden M, Yu X, Heugen U, Gruebele M, Leitner DM, Havenith M (2007) An extended dynamical hydration shell around proteins. Proc Natl Acad Sci USA 104:20479–20752
Essmann U, Perera L, Berkowitz ML, Darden T, Lee H, Pedersen LG (1995) A smooth particle mesh Ewald method. J Chem Phys 103:8577–8593
Feller SE, MacKerell AD Jr (2000) An improved empirical potential energy function for molecular simulations of phospholipids. J Phys Chem B 104:7510–7515
Feller SE, Zhang Y, Pastor RW, Brooks B (1995) Constant pressure molecular dynamics simulation: the Langevin piston method. J Chem Phys 103:4613–4621
Hübner W, Blume A (1998) Interactions at the lipid-water interface. Chem Phys Lipid 96:99–123
Humphrey W, Dalke W, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14:33–38
Jo S, Kim T, Iyer VG, Im W (2008) CHARMM-GUI: a web-based graphical user interface for CHARMM. J Comput Chem 29:1859–1865
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926–935
Kalé L, Skeel R, Bhandarkar M, Brunner R, Gursoy A, Krawetz N, Phillips J, Shinozaki A, Varadarajan K, Schulten K (1999) NAMD2: greater scalability for parallel molecular dynamics. J Comput Phys 151:283–312
Klauda JB, Venable RM, Freites JA, O’Connor JW, Tobias DJ, Mondragon-Ramirez C, Votrobyov I, MacKerell AD Jr, Pastor RW (2010) Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J Phys Chem B 114:7830–7843
Lopez CF, Nielsen SO, Klein ML, Moore PB (2004) Hydrogen binding structure and dynamics of water at the dimyristoylphosphatidylcholine lipid bilayer surface from a molecular dynamics simulation. J Phys Chem B 108:6603–6610
Lorch S, Capponi S, Pieront F, Bondar A-N (2015) Dynamic carboxylate/water networks on the surface of the PsbO subunit of photosystem II. J Phys Chem B 119:12172–12181
Makarov VA, Feig M, Andrews BK, Pettitt MB (1998) Diffusion of solvent around biomolecular solutes: a molecular dynamics simulation study. Biophys J 75:150–158
Marrink SJ, Berkowitz ML, Berendsen HJC (1993) Molecular dynamics simulation of a membrane/water interface: the ordering of water and its relation to the hydration force. Langmuir 9:3122–3131
Milenkovic S, Bondar A-N (2016) Mechanism of conformational coupling in SecA: key role of hydrogen-bonding networks and water interactions. Biochim Biophys Acta 1858:374–385
Milenkovic S, Bondar A-N (2018) Motions of the SecA protein motor bound to signal peptide: Insights from molecular dynamics simulations. Biochim Biophys Acta 1860:416–427
Muegge I, Knapp E-W (1995) Residence times and lateral diffusion of water at protein surfaces: application to BPTI. J Phys Chem 99:1371–1374
Mukhopadhyay P, Monticelli L, Tieleman DP (2004) Molecular dynamics simulations of a palmitoyl-oleoyl phosphatidylserine bilayer with Na+ counterions and NaCl. Biophys J 86:1601–1609
Murzyn K, Róg T, Pasenkiewicz-Gierula M (2005) Phosphatidylethanolamine-phosphatidylglycerol bilayer as a model of the inner bacterial membrane. Biophys J 88:1091–1103
Nagle JF, Tristram-Nagle S (2000) Structure of lipid bilayers. Biochim Biophys Acta 1429:159–195
Noble JM, Thomas TH, Ford GA (1999). Effect of age on plasma membrane asymmetry and membrane fluidity in human leokocytes and platelets. J Gerontol: Med Sci 54:M60–M606
Pandit KR, Klauda JB (2012) Membrane models of E. coli containing cyclic moieties in the aliphatic lipid chain. Biochim Biophys Acta 1818:1205–1210
Pandit SA, Bostick D, Berkowitz ML (2003) Mixed bilayer containing dipalmitoylphosphatidylcholine and dipalmitoylphosphatidylserine: lipid compensation, ion binding, and electrostatics. Biophys J 85:3120–3131
Pasenkiewicz-Gierula M, Baczynski K, Markiewicz M, Murzyn K (2016) Computer modelling studies of the bilayer/water interface. Biochim Biophys Acta 1858:2305–2321
Pasenkiewicz-Gierula M, Takaoka Y, Miyagawa H, Kitamura K, Kusumi A (1997) Hydrogen bonding of water to phosphatidylcholine in the membrane as studied by a molecular dynamics simulation: location, geometry, and lipid-bridging via hydrogen-bonded water. J Phys Chem A 101:3677–3691
Petrache HI, Tristam-Nagle S, Gawrisch K, Harries D, Parsegian VA, Nagle JF (2004) Structure and fluctuations of charged phosphatidylserine bilayers in the absence of salt. Biophys J 86:1574–1586
Phillips JC, Braun B, Wang W, Gumbart J, Takjkhorshid E, Villa E, Chipot C, Skeel RD, Kale L, Schulten K (2005) Scalable molecular dynamics with NAMD. J Comput Chem 26:1781–1802
Prats M, Tocanne JF, Teissie J (1987) Lateral proton conduction at a lipid/water interface. Effect of lipid nature and ionic content of the aqueous phase. Eur J Biochem 162:379–385
Ran S, Thorpe PE (2002) Phosphatidylserine is a marker of tumor vasuculature and a potential target for cancer imaging and therapy. Int J Radioation Oncology Biol Phys 54:1479–1484
Riedl S, Kinner B, Asslaber M, Schaider H, Walzer S, Novak A, Lohner K, Zweytick D (2011) In search of a novel target: phosphatidylserine exposed by non-apoptotic tumor cells and metastases of malignacies with poor treatment efficacy. Biochim Biophys Acta 1808:2638–2645
Róg T, Mutzyn K, Milhaud J, Karttunen M, Pasenkiewicz-Gierula M (2009) Water isotope effect on the phosphatidylcholine bilayer properties: a molecular dynamics simulation study. J Phys Chem B 113:2378–2387
Ryckaert J-P, Ciccotti G, Berendsen HJC (1977) Numerical integration of the Cartesian equations of motion of a system with constraints. Molecular dynamics of n-alkanes. J Comput Phys 23:327–341
Sengupta N, Jaud S, Tobias DJ (2008) Hydration dynamics in a partially denatured ensemble of the globular protein human a-lactalbumin investigated with molecular dynamics simulations. Biophys J 95:5257–5267
Smondyrev AM, Berkowitz ML (1999) Structure of dipalmidoylphosphatidylcholine/cholesterol bilayer at low and high cholesterol concentrations: molecular dynamics simulation. Biophys J 77:2075–2089
Springer A, Hagen V, Cherepanov DA, Antonenko YN, Pohl P (2011) Protons migrate along interfacial water without significant contributions from jumps between ionizable groups on the membrane surface. Proc Natl Acad Sci 108:14461–14466
Sterpone F, Stirnemann G, Hynes JT, Laage D (2010) Water hydrogen-bond dynamics around amino acids: the key role of hydrophilic hydrogen-bond acceptor groups. J Phys Chem B 114:2083–2089
Sushko O, Dubrovka R, Donnan RS (2013) Terahertz spectral domain computational analysis of hydration shell of proteins with increasingly complex tertiary structure. J Phys Chem B 117:16486–16492
The MathWorks, I (2017). MATLAB. Natick, Massachusetts
Tielrooij KJ, Paparo D, Piatkowski L, Bakker HJ, Bonn M (2009) Dielectroc relaxation dynamics of water in model membranes probed by terahertz spectroscopy. Biophys J 97:2484–2492
Tuckermann M, Berne BJ, Martyna GJ (1992) Reversible multiple time scale molecular dynamics. J Chem Phys 97:1990–2001
Urban S, Lee JR, Freeman M (2001) Drosphila rhomboid-1 defines a family of putative intramembrane serine proteases. Cell 107:173–182
Urban S, Wolfe MS (2005) Reconstitution of intramembrane proteolysis in vitro reveals that pure rhomboid is sufficient for catalysis and specificity. Proc Natl Acad Sci USA 102:1883–1888
van Klompenburg W, Nilsson I, von Heijne G, de Kruijff B (1997) Anionic phospholipids are determinants of membrane protein topology. EMBO J 16:4261–4266
Volkov VV, Palmer DJ, Righini R (2007) Heterogeneity of water at the phospholipid membrane interface. J Phys Chem B 111:1377–1383
Wiener MC, White SH (1992) Structure of a fluid dioleoylphosphatidylcholine bilayer determined by joint refinement of X-ray and neutron diffraction data. III. Complete structure. Biophys J 61:434–447
Wolf MG, Grubmüller H, Groenhof G (2014) Anomalous surface diffusion of protons on lipid membranes. Biophys J 107:76–87
Wu EL, Cheng X, Jo S, Rui H, Song KC, Dávila-Contreras EM, Qi Y, Lee J, Monje-Galvan V, Venable RM, Klauda JB, Im W (2014) CHARMM-GUI membrane builder toward realistic biological membrane simulations. J Comput Chem 35:1997–2004
Yeung T, Gilbert GE, Shi J, Silvius J, Kapus A, Grinstein S (2008) Membrane phosphatidylserine regulates surface charge and protein localization. Science 319:210–213
Zhao W, Róg T, Gurtovenko AA, Vattulainen I, Karttunen M (2007) Atomic-scale structure and electrostatics of anionic palmitoyloleoylphosphatidylglycerol lipid bilayers with Na+ counterions. Biophys J 92:1114–1124
Zwaal RFA, Comfurius P, Bevers EM (2005) Surface exposure of phosphatidylserine in pathological cells. Cell Mol Life Sci 62:971–988
Zwaal RFA, Schroit AJ (1997) Pathophysiological implications of membrane phospholipd asymmetry in blood cells. J Am Soc Hemathol 89:1121–1132
Acknowledgements
The project was supported by the Freie Universität Berlin within the Excellence Initiative of the German Research Foundation and by an allocation of computing time from HLRN, the North-German Supercomputing Alliance (bec00063, to A-NB).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors declare no conflicts of interest.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Karathanou, K., Bondar, AN. Dynamic Water Hydrogen-Bond Networks at the Interface of a Lipid Membrane Containing Palmitoyl-Oleoyl Phosphatidylglycerol. J Membrane Biol 251, 461–473 (2018). https://doi.org/10.1007/s00232-018-0023-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-018-0023-1