Abstract
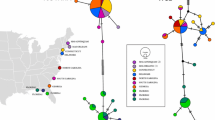
Comparing the genetic structure of host populations with that of their parasites can shed light on the efficiency and independence of their respective dispersal mechanisms. The degree of congruence between host and parasite genetic structure should reflect to what extent they share dispersal mechanisms. Here, we contrast the genetic structure of the acanthocephalan parasite Profilicollis novaezelandensis with that of its intermediate host, the hairy-handed shore crab Hemigrapsus crenulatus, along the east coast of New Zealand’s South Island. We expected no congruence in their phylogeographic patterns because of the very different modes of dispersal used by the crabs (planktonic drift) and the acanthocephalans (bird-mediated dispersal). Based on analysis of cytochrome oxidase subunit I gene sequences, we found no significant genetic structure among isolated populations of the crab and those of their parasite, along a roughly 600 km stretch of coastline. Surprisingly, based on a distance-based co-evolutionary analysis statistical tool (PACo), we observed an overall significant level of congruence between host and parasite population-level phylogenies. The most parsimonious interpretation is that statistical significance does not translate into biological significance, with the result likely due to chance, possibly because bird movements that disperse parasites coincidentally match patterns of crab dispersal by ocean currents in parts of our study area. In this system, the connectivity among different localities and the apparent genetic mixing among populations may have implications for host–parasite co-evolution.





Similar content being viewed by others
References
Balbuena JA, Miguez-Lozano R, Blasco-Costa I (2013) PACo: a novel procrustes application to cophylogenetic analysis. PLoS One 8:e61048
Blasco-Costa I, Poulin R (2013) Host traits explain the genetic structure of parasites: a meta-analysis. Parasitology 140:1316–1322
Brockerhoff AM, Smales LR (2002) Profilicollis novaezelandensis n. sp (Polymorphidae) and two other acanthocephalan parasites from shore birds (Haematopodidae and Scolopacidae) in New Zealand, with records of two species in intertidal crabs (Decapoda: Grapsidae and Ocypodidae). Syst Parasitol 52:55–65
Bryan-Walker K, Leung TLF, Poulin R (2007) Local adaptation of immunity against a trematode parasite in marine amphipod populations. Mar Biol 152:687–695
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1659
Cornet S, Biard C, Moret Y (2009) Variation in immune defence among populations of Gammarus pulex (Crustacea: Amphipoda). Oecologia 159:257–269
Criscione CD, Blouin MS (2004) Life cycles shape parasite evolution: comparative population genetics of salmon trematodes. Evolution 58:198–202
de Vienne DM, Aguileta G, Ollier S (2011) Euclidean nature of phylogenetic distance matrices. Syst Biol 60:826–832
Devlin CM, Diamond AW, Saunders GW (2004) Sexing arctic terns in the field and laboratory. Waterbirds 27:314–320
Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol 7:214
Dybdahl M, Lively CM (1996) The geography of coevolution: comparative population structures for a snail and its trematode parasite. Evolution 50:2264–2275
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Fordham R (1968) Dispersion and dispersal of the Dominican gull in Wellington, New Zealand. Proc N Z Ecol Soc 15:40–50
Garcia-Varela M, Pérez-Ponce de Léon G, Aznar FJ, Nadler SA (2013) Phylogenetic relationship among genera of Polymorphidae (Acanthocephala), inferred from nuclear and mitochondrial gene sequences. Mol Phylogenet Evol 68:176–184
Geller J, Meyer C, Parker M, Hawk H (2013) Redesign of PCR primers for mitochondrial cytochrome c oxidase subunit I for marine invertebrates and application in all-taxa biotic surveys. Mol Ecol Resour 13:851–861
Gilg MR, Hilbish TJ (2003) The geography of marine larval dispersal: coupling genetics with fine-scale physical oceanography. Ecology 84:2989–2998
Goulding TC, Cohen CS (2014) Phylogeography of a marine acanthocephalan: lack of cryptic diversity in a cosmopolitan parasite of mole crabs. J Biogeogr 41:965–976
Groner ML, Maynard J, Breyta R et al (2016) Managing marine disease emergencies in an era of rapid change. Phil Trans R Soc B 371:20150364
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321
Harper JT, Saunders GW (2001) The application of sequences of the ribosomal cistron to the systematics and classification of the florideophyte red algae (Florideophyceae, Rhodophyta). Cah Biol Mar 42:25–38
Jombart T (2008) adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 11:94
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Kearse M et al (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinform 28:1647–1649
Keeney DB, King TM, Rowe DL, Poulin R (2009) Contrasting mtDNA diversity and population structure in a direct-developing marine gastropod and its trematode parasites. Mol Ecol 18:4591–4603
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Lagrue C, Joannes A, Poulin R, Blasco-Costa I (2016) Genetic structure and host–parasite co-divergence: evidence for trait-specific local adaptation. Biol J Linn Soc 118:344–358
Lanfear R, Calcott B, Ho SYW, Guindon S (2012) PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol Biol Evol 29:1695–1701
Lanfear R, Frandsen PB, Wright AM, Senfeld T, Calcott B (2016) PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol Biol Evol 34:772–773
Latham ADM, Poulin R (2002a) Field evidence of the impact of two acanthocephalan parasites on the mortality of three species of New Zealand shore crabs (Brachyura). Mar Biol 141:1131–1139
Latham ADM, Poulin R (2002b) Effect of acanthocephalan parasites on hiding behaviour in two species of shore crabs. J Helminthol 76:323–326
Latham ADM, Poulin R (2003) Spatiotemporal heterogeneity in recruitment of larval parasites to shore crab intermediate hosts: the influence of shorebird definitive hosts. Can J Zool 81:1282–1291
Leigh JW, Bryant D (2015) POPART: full-feature software for haplotype network construction. Methods Ecol Evol 6:1110–1116
Martinez-Aquino A, Reyna-Fabian ME, Rosas-Valdez R, Razo-Mendivil U, Pérez-Ponce de León G, Garcia-Varela M (2009) Detecting a complex of cryptic species within Neoechinorhynchus golvani (Acanthocephala: Neoechinorhynchidae) inferred from ITSs and LSU rDNA gene sequences. J Parasitol 95:1040–1047
Mazé-Guilmo E, Blanchet S, McCoy KD, Loot G (2016) Host dispersal as the driver of parasite genetic structure: a paradigm lost? Ecol Lett 19:336–347
McLay CL (1988) Brachyura and crab-like Anomura of New Zealand. University of Auckland Marine Laboratory, Auckland
Miller MA, Pfeiffer W, Schwartz T (2010) Creating the CIPRES science gateway for inference of large phylogenetic trees. In: Proceedings of the gateway computing environments workshop, New Orleans, LA, pp 1–8
Morjan CL, Rieseberg LH (2004) How species evolve collectively: implications of gene flow and selection for the spread of advantageous alleles. Mol Ecol 13:1341–1356
Paradis E, Claude J, Strimmer K (2004) APE: analyses of phylogenetics and evolution in R language. Bioinform 20:289–290
Prugnolle F, Théron A, Pointier JP, Jabbour-Zahab R, Jarne P, Durand P, de Meeûs T (2005) Dispersal in a parasitic worm and its two hosts: consequences for local adaptation. Evolution 59:296–303
Rodriguez SM, D’Elia G (2017) Pan-American marine coastal distribution of the acanthocephalan Profilicollis altmani based on morphometric and phylogenetic analyses of cystacanths from the mole crab Emerita brasiliensis. J Helminthol 91:371–375
Ronquist F, Huelsenbeck JP (2003) MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinform 19:1572–1574
Ross PM, Hogg ID, Pilditch CA, Lundquist CJ (2009) Phylogeography of New Zealand’s coastal benthos. N Z J Mar Freshw Res 43:1009–1027
Rowe L (2013) Dispersal of southern black-backed gulls (Larus dominicanus dominicanus) banded in Canterbury, New Zealand, 1959–1993. Notornis 60:134–142
Steinauer ML, Nickol BB, Orti G (2007) Cryptic speciation and patterns of phenotypic variation of a highly variable acanthocephalan parasite. Mol Ecol 16:4097–4109
Sutton PJH (2003) The Southland current: a subantarctic current. N Z J Mar Freshw Res 37:645–652
Tamura K, Nei M (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial-DNA in humans and chimpanzees. Mol Biol Evol 10:512–526
Tobias ZJC, Jorge F, Poulin R (2017) Life at the beach: comparative phylogeography of a sandhopper and its nematode parasite reveals extreme lack of parasite mtDNA variation. Biol J Linn Soc 122:113–132
Wallis GP, Trewick SA (2009) New Zealand phylogeography: evolution on a small continent. Mol Ecol 18:3548–3580
Ward JR, Lafferty KD (2004) The elusive baseline of marine disease: are diseases in ocean ecosystems increasing? PLoS Biol 2:e120
Wear RG (1970) Life-history studies on New Zealand Brachyura. N Z J Mar Freshw Res 4:3–35
Zhang Z, Schwartz S, Wagner L, Miller W (2000) A greedy algorithm for aligning DNA sequences. J Comput Biol 7:203–214
Acknowledgements
We thank Lynda Hay, Lance Hay, Jahmaine Hay and Kirby McKenzie for assistance with crab collection in the field, and Dr. Bronwen Presswell for providing an adult acanthocephalan specimen.
Funding
This research was funded internally by the Department of Zoology, University of Otago, and received no external funding from commercial or non-profit sectors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare having no conflict of interest.
Ethical statement
Collection and euthanasia of crabs were approved by the Otago University Animal Ethics Committee (Application no. ET 2/17).
Additional information
Responsible Editor: T. Reusch.
Reviewed by Undisclosed experts.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hay, E., Jorge, F. & Poulin, R. The comparative phylogeography of shore crabs and their acanthocephalan parasites. Mar Biol 165, 69 (2018). https://doi.org/10.1007/s00227-018-3326-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00227-018-3326-y