Abstract
Determining the quantity of active virus is the most important basis to judge the risk of virus infection, which usually relies on the virus median tissue culture infectious dose (TCID50) assay performed in a biosafety level 3 laboratory within 5–7 days. We have developed a culture-free method for rapid and accurate quantification of active severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) by targeting subgenomic RNA (sgRNA) based on reverse transcription digital PCR (RT-dPCR). The dynamic range of quantitative assays for sgRNA-N and sgRNA-E by RT-dPCR was investigated, and the result showed that the limits of detection (LoD) and quantification (LoQ) were 2 copies/reaction and 10 copies/reaction, respectively. The delta strain (NMDC60042793) of SARS-CoV-2 was cultured at an average titer of 106.13 TCID50/mL and used to evaluate the developed quantification method. Copy number concentrations of the cultured SARS-CoV-2 sgRNA and genomic RNA (gRNA) gave excellent linearity (R2 = 0.9999) with SARS-CoV-2 titers in the range from 500 to 105 TCID50/mL. Validation of 63 positive clinical samples further proves that the quantification of sgRNA-N by RT-dPCR is more sensitive for active virus quantitative detection. It is notable that we can infer the active virus titer through quantification of SARS-CoV-2 sgRNA based on the linear relationship in a biosafety level 2 laboratory within 3 h. It can be used to timely and effectively identify infectious patients and reduce unnecessary isolation especially when a large number of COVID-19 infected people impose a burden on medical resources.
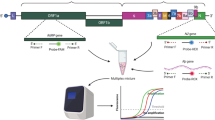
Graphical abstract






Similar content being viewed by others
References
Boger B, Fachi MM, Vilhena RO, Cobre AF, Tonin FS, Pontarolo R. Systematic review with meta-analysis of the accuracy of diagnostic tests for COVID-19. Am J Infect Control. 2021;49(1):21–9. https://doi.org/10.1016/j.ajic.2020.07.011.
Sule WF, Oluwayelu DO. Real-time RT-PCR for COVID-19 diagnosis: challenges and prospects. Pan Afr Med J, 2020, 35(Suppl 2): 121. 10.11604/pamj.supp.2020.35.24258.
Bustin S, Kirvell S, Huggett JF, Nolan T. RT-qPCR diagnostics: the “Drosten” SARS-CoV-2 Assay Paradigm. Int J Mol Sci. 2021;22(16) https://doi.org/10.3390/ijms22168702.
Wolfel R, Corman VM, Guggemos W, Seilmaier M, Zange S, Muller MA, et al. Virological assessment of hospitalized patients with COVID-2019. Nature. 2020;581(7809):465–9. https://doi.org/10.1038/s41586-020-2196-x.
Dagotto G, Mercado NB, Martinez DR, Hou YJ, Nkolola JP, Carnahan RH, et al. Comparison of subgenomic and total RNA in SARS-CoV-2 challenged rhesus macaques. J Virol. 2021;95(8) https://doi.org/10.1128/JVI.02370-20.
Rayms-Keller A, Powers AM, Higgs S, Olson KE, Kamrud KI, Carlson JO, et al. Replication and expression of a recombinant Sindbis virus in mosquitoes. Insect Mol Biol. 1995;4(4):245–51. https://doi.org/10.1111/j.1365-2583.1995.tb00030.x.
Smither SJ, Lear-Rooney C, Biggins J, Pettitt J, Lever MS, Olinger GG Jr. Comparison of the plaque assay and 50% tissue culture infectious dose assay as methods for measuring filovirus infectivity. J Virol Methods. 2013;193(2):565–71. https://doi.org/10.1016/j.jviromet.2013.05.015.
Santos Bravo M, Berengua C, Marín P, Esteban M, Rodriguez C, Del Cuerpo M, et al. Viral culture confirmed SARS-CoV-2 subgenomic RNA value as a good surrogate marker of infectivity. J Clin Microbiol. 2022;60(1):e0160921. https://doi.org/10.1128/jcm.01609-21.
Sawicki SG, Sawicki DL, Siddell SG. A contemporary view of coronavirus transcription. J Virol. 2007;81(1):20–9. https://doi.org/10.1128/jvi.01358-06.
Sola I, Almazan F, Zuniga S, Enjuanes L. Continuous and discontinuous RNA synthesis in coronaviruses. Annu Rev Virol. 2015;2(1):265–88. https://doi.org/10.1146/annurev-virology-100114-055218.
Kim D, Lee JY, Yang JS, Kim JW, Kim VN, Chang H. The architecture of SARS-CoV-2 transcriptome. Cell. 2020;181(4):914–21.e10. https://doi.org/10.1016/j.cell.2020.04.011.
Davidson AD, Williamson MK, Lewis S, Shoemark D, Carroll MW, Heesom KJ, et al. Characterisation of the transcriptome and proteome of SARS-CoV-2 reveals a cell passage induced in-frame deletion of the furin-like cleavage site from the spike glycoprotein. Genome Med. 2020;12(1):68. https://doi.org/10.1186/s13073-020-00763-0.
Finkel Y, Mizrahi O, Nachshon A, Weingarten-Gabbay S, Morgenstern D, Yahalom-Ronen Y, et al. The coding capacity of SARS-CoV-2. Nature. 2021;589(7840):125–30. https://doi.org/10.1038/s41586-020-2739-1.
Taiaroa G, Rawlinson D, Featherstone L, Pitt M, Caly L, Druce J, Purcell D, Harty L, Tran T, Roberts J, Scott N, Catton M, Williamson D, Coin L. Direct RNA sequencing and early evolution of SARS-CoV-2. bioRxiv. 2020. https://doi.org/10.1101/2020.03.05.976167.
Xu J, Kirtek T, Xu Y, Zheng H, Yao H, Ostman E, et al. Digital droplet PCR for SARS-CoV-2 resolves borderline cases. Am J Clin Pathol. 2021;155(6):815–22. https://doi.org/10.1093/ajcp/aqab041.
Nyaruaba R, Li C, Mwaliko C, Mwau M, Odiwuor N, Muturi E, et al. Developing multiplex ddPCR assays for SARS-CoV-2 detection based on probe mix and amplitude based multiplexing. Expert Rev Mol Diagn. 2021;21(1):119–29. https://doi.org/10.1080/14737159.2021.1865807.
Liu X, Feng J, Zhang Q, Guo D, Zhang L, Suo T, et al. Analytical comparisons of SARS-COV-2 detection by qRT-PCR and ddPCR with multiple primer/probe sets. Emerg Microbes Infect. 2020;9(1):1175–9. https://doi.org/10.1080/22221751.2020.1772679.
Nyaruaba R, Mwaliko C, Dobnik D, Neuzil P, Amoth P, Mwau M, et al. Digital PCR Applications in the SARS-CoV-2/COVID-19 era: a roadmap for future outbreaks. Clin Microbiol Rev. 2022;35(3):e0016821. https://doi.org/10.1128/cmr.00168-21.
Dong L, Zhou J, Niu C, Wang Q, Pan Y, Sheng S, et al. Highly accurate and sensitive diagnostic detection of SARS-CoV-2 by digital PCR. Talanta. 2021;224:121726. https://doi.org/10.1016/j.talanta.2020.121726.
Németh B, Fasseeh A, Molnár A, Bitter I, Horváth M, Kóczián K, et al. A systematic review of health economic models and utility estimation methods in schizophrenia. Expert Rev Pharmacoecon Outcomes Res. 2018;18(3):267–75. https://doi.org/10.1080/14737167.2018.1430571.
Casagrande M, Fitzek A, Spitzer M, Puschel K, Glatzel M, Krasemann S, et al. Detection of SARS-CoV-2 genomic and subgenomic RNA in retina and optic nerve of patients with COVID-19. Br J Ophthalmol. 2022;106(9):1313–7. https://doi.org/10.1136/bjophthalmol-2020-318618.
Niu C, Wang X, Gao Y, Qiao X, Xie J, Zhang Y, et al. Accurate quantification of SARS-CoV-2 RNA by isotope dilution mass spectrometry and providing a correction of reverse transcription efficiency in droplet digital PCR. Anal Bioanal Chem. 2022;414(23):6771–7. https://doi.org/10.1007/s00216-022-04238-6.
Huggett JF, Foy CA, Benes V, Emslie K, Garson JA, Haynes R, et al. The digital MIQE guidelines: minimum information for publication of quantitative digital PCR experiments. Clinical chemistry. 2013;59(6):892–902. https://doi.org/10.1373/clinchem.2013.206375.
Weaver S, Dube S, Mir A, Qin J, Sun G, Ramakrishnan R. Taking qPCR to a higher level: analysis of CNV reveals the power of high throughput qPCR to enhance quantitative resolution. Methods (San Diego, Calif). 2010;50:271–6.
Daniel WT, Kristian L, Marina K, David AA, Patricia EG, Robert LJ, Martin HK, Rudolf ML, Thomas JP, Scassellati GA, Heinz S, Jane T. EP17-A—Protocols for determination of limits of detection and limits of quantitation; Approved Guideline.
Yang M, Cao S, Liu Y, Zhang Z, Zheng R, Li Y, et al. Performance verification of five commercial RT-qPCR diagnostic kits for SARS-CoV-2. Clin Chim Acta. 2022;525:46–53. https://doi.org/10.1016/j.cca.2021.12.004.
Vierbaum L, Wojtalewicz N, Grunert HP, Lindig V, Duehring U, Drosten C, et al. RNA reference materials with defined viral RNA loads of SARS-CoV-2-A useful tool towards a better PCR assay harmonization. PLoS One. 2022;17(1):e0262656. https://doi.org/10.1371/journal.pone.0262656.
Niu C, Wang X, Zhang Y, Lu L, Wang D, Gao Y, et al. Interlaboratory assessment of quantification of SARS-CoV-2 RNA by reverse transcription digital PCR. Anal Bioanal Chem. 2021;413(29):7195–204. https://doi.org/10.1007/s00216-021-03680-2.
Tan SYH, Kwek SYM, Low H, Pang YLJ. Absolute quantification of SARS-CoV-2 with Clarity Plus™ digital PCR. Methods (San Diego, Calif). 2022;201:26–33. https://doi.org/10.1016/j.ymeth.2021.07.005.
Hou YJ, Okuda K, Edwards CE, Martinez DR, Asakura T, Dinnon KH 3rd, et al. SARS-CoV-2 reverse genetics reveals a variable infection gradient in the respiratory tract. Cell. 2020;182(2):429–46.e14. https://doi.org/10.1016/j.cell.2020.05.042.
Telwatte S, Martin HA, Marczak R, Fozouni P, Vallejo-Gracia A, Kumar GR, et al. Novel RT-ddPCR assays for measuring the levels of subgenomic and genomic SARS-CoV-2 transcripts. Methods. 2022;201:15–25. https://doi.org/10.1016/j.ymeth.2021.04.011.
Dimcheff DE, Valesano AL, Rumfelt KE, Fitzsimmons WJ, Blair C, Mirabelli C, et al. Severe Acute respiratory syndrome coronavirus 2 total and subgenomic RNA viral load in hospitalized patients. J Infect Dis. 2021;224(8):1287–93. https://doi.org/10.1093/infdis/jiab215.
Moron-Lopez S, Riveira-Munoz E, Urrea V, Gutierrez-Chamorro L, Avila-Nieto C, Noguera-Julian M, et al. Comparison of reverse transcription (RT)-quantitative PCR and RT-droplet digital PCR for detection of genomic and subgenomic SARS-CoV-2 RNA. Microbiol Spectr. 2023;11(2):e0415922. https://doi.org/10.1128/spectrum.04159-22.
Osborn LJ, Chen PY, Flores-Vazquez J, Mestas J, Salas E, Glucoft M, et al. Clinical utility of SARS-CoV-2 subgenomic RT-PCR in a pediatric quaternary care setting. J Clin Virol. 2023;164:105494. https://doi.org/10.1016/j.jcv.2023.105494.
Yang J, Kidd M, Nordquist AR, Smith SD, Hurth C, Modlin IM, et al. A sensitive, portable microfluidic device for SARS-CoV-2 detection from self-collected Saliva. Infect Dis Rep. 2021;13(4):1061–77. https://doi.org/10.3390/idr13040097.
Singanayagam A, Patel M, Charlett A, Lopez Bernal J, Saliba V, Ellis J, et al. Duration of infectiousness and correlation with RT-PCR cycle threshold values in cases of COVID-19, England, January to May 2020. Euro Surveill., 2020, 25(32). https://doi.org/10.2807/1560-7917.ES.2020.25.32.2001483.
Funding
This work was supported by Basic scientific research project of National institute of metrology (AKYZZ2126, AKYYJ2009/ZYZJ2001) and the Major Science and Technique Programs in Yunnan Province (202102AA310055).
Author information
Authors and Affiliations
Contributions
Conceptualization and methodology: Lianhua Dong, Xinhua Dai, Xiang Fang, and Minghua Li. Methodology: Lianhua Dong, Xia Wang, Chunyan Niu, and Xiaoli Feng. Software and formal analysis: Lianhua Dong, Yi Yang, and Tao Peng. Investigation and data curation: Yi Yang, Yang Pan, Xiaoli Feng, Wang Qu, and Qingcui Zou. Resources: Lianhua Dong, Xinhua Dai, and Minghua Li. Writing—original draft preparation: Lianhua Dong, Yi Yang, Xia Wang, and Tao Peng. Writing—review and editing: Lianhua Dong, Yang Pan, and Tao Peng. Supervision: Lianhua Dong, Xinhua Dai, and Xiang Fang. All authors have read and agreed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This study was approved by the Ethics Committee of the Beijing Center for Disease Prevention and Control. The analysis was performed on existing samples collected during standard diagnostic tests, posing no extra burden to patients.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(DOCX 1349 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, Y., Feng, X., Pan, Y. et al. A culture-free method for rapidly and accurately quantifying active SARS-CoV-2. Anal Bioanal Chem 415, 5745–5753 (2023). https://doi.org/10.1007/s00216-023-04855-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-023-04855-9