Abstract
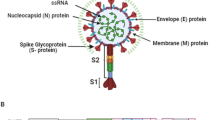
Emerging and re-emerging zoonotic viral diseases continue to significantly impact public health. Of particular interest are enveloped viruses (e.g., SARS-CoV-2, the causative pathogen of COVID-19), which include emerging pathogens of highest concern. Enveloped viruses contain a viral envelope that encapsulates the genetic material and nucleocapsid, providing structural protection and functional bioactivity. The viral envelope is composed of a coordinated network of glycoproteins and lipids. The lipid composition of the envelope consists of lipids preferentially appropriated from host cell membranes. Subsequently, changes to the host cell lipid metabolism and an accounting of what lipids are changed during viral infection provide an opportunity to fingerprint the host cell’s response to the infecting virus. To address this issue, we comprehensively characterized the lipid composition of VeroE6-TMPRSS2 cells infected with SARS-CoV-2. Our approach involved using an innovative solid-phase extraction technique to efficiently extract cellular lipids combined with liquid chromatography coupled to high-resolution tandem mass spectrometry. We identified lipid changes in cells exposed to SARS-CoV-2, of which the ceramide to sphingomyelin ratio was most prominent. The identification of a lipid profile (i.e., lipid fingerprint) that is characteristic of cellular SARS-CoV-2 infection lays the foundation for targeting lipid metabolism pathways to further understand how enveloped viruses infect cells, identifying opportunities to aid antiviral and vaccine development.
Graphical abstract






Similar content being viewed by others
Abbreviations
- WHO:
-
World Health Organization
- HIV:
-
Human immunodeficiency virus
- HSV:
-
Herpes simplex virus
- RSV:
-
Respiratory syncytial virus
- VSV:
-
Vesicular stomatitis virus
- LC HR-MS/MS:
-
Liquid chromatography, high-resolution tandem mass spectrometry
- SPE:
-
Solid-phase extraction
- HPLC:
-
High-pressure liquid chromatography
- MTBE:
-
Tert-Butyl methyl ether
- HRMS:
-
High-resolution mass spectrometry
- Q-TOF:
-
Quadrupole time of flight
- LLE:
-
Liquid-liquid extraction
- PCA:
-
Principal component analysis
- PLS-DSA:
-
Partial least squares discriminate analysis
- CE:
-
Cholesterol esters
- GP:
-
Glycerophospholipids
- DG:
-
Diacylglycerol
- TG:
-
Triacylglycerol
- SP:
-
Sphingolipids
- Cer:
-
Ceramide
- SM:
-
Sphingomyelin
- LPC:
-
Lysophosphocholine
- LPE:
-
Lysophosphoethanolamine
- PC:
-
Glycerophosphoscholine
- ePC:
-
Ether glycerophosphocholine
- PE :
-
Glycerophosphoethanolamine
- PI:
-
Glycerophosphoinositol
- PS:
-
Glycerophosphoserine
- IVA:
-
Influenza A
- VE6-TMPRSS2:
-
Vero E6 cells overexpressing TMPRSS2
- ASM:
-
Acid sphingomyelinase
- FDA:
-
Federal Drug Administration
- FIASMAs:
-
Functional inhibitors of acid sphingomyelinase
- PLA2 :
-
Phospholipase A2
- cPLA2 :
-
Cytosolic PLA2
- SEM:
-
Standard error of the mean
- TIC:
-
Total ion chromatogram
- EIC:
-
Extracted ion chromatogram
- t R :
-
Retention time
- FC:
-
Fold change
References
Åsjö B, Kruse H. Zoonoses in the emergence of human viral diseases. Perspect Med Virol. 2006;16:15–41.
Johns Hopkins University and Medicine CRC. COVID-19 Dashboard by the Center for Systems Science and Engineering (CSSE) at Johns Hopkins University (JHU) 2020 [Available from: https://coronavirus.jhu.edu/map.html. Accessed 22 March 2023.
Sweileh WM. Global research trends of World Health Organization’s top eight emerging pathogens. Global Health. 2017;13(1):9.
Vigant F, Santos NC, Lee B. Broad-spectrum antivirals against viral fusion. Nat Rev Microbiol. 2015;13(7):426–37.
Chan RB, Tanner L, Wenk MR. Implications for lipids during replication of enveloped viruses. Chem Phys Lipid. 2010;163(6):449–59.
Schmitt AP, Lamb RA. Influenza virus assembly and budding at the viral budozone. Adv Virus Res. 2005;64:383–416.
Harrison SC. Viral membrane fusion. Virology. 2015;479–480:498–507.
Lorizate M, Krausslich HG. Role of lipids in virus replication. Cold Spring Harb Perspect Biol. 2011;3(10): a004820.
Gerl MJ, Sampaio JL, Urban S, Kalvodova L, Verbavatz JM, Binnington B, et al. Quantitative analysis of the lipidomes of the influenza virus envelope and MDCK cell apical membrane. J Cell Biol. 2012;196(2):213–21.
Ivanova PT, Myers DS, Milne SB, McClaren JL, Thomas PG, Brown HA. Lipid composition of viral envelope of three strains of influenza virus - not all viruses are created equal. ACS Infect Dis. 2015;1(9):399–452.
Tran A, Monreal IA, Moskovets E, Aguilar HC, Jones JW. Rapid detection of viral envelope lipids using lithium adducts and AP-MALDI high-resolution mass spectrometry. J Am Soc Mass Spectrom. 2021;32(9):2322–33.
Sezgin E, Levental I, Mayor S, Eggeling C. The mystery of membrane organization: composition, regulation and roles of lipid rafts. Nat Rev Mol Cell Biol. 2017;18(6):361–74.
McMahon HT, Boucrot E. Membrane curvature at a glance. J Cell Sci. 2015;128(6):1065–70.
Glaser PE, Gross RW. Plasmenylethanolamine facilitates rapid membrane fusion: a stopped-flow kinetic investigation correlating the propensity of a major plasma membrane constituent to adopt an HII phase with its ability to promote membrane fusion. Biochemistry. 1994;33(19):5805–12.
Meher G, Chakraborty H. Membrane composition modulates fusion by altering membrane properties and fusion peptide structure. J Membr Biol. 2019;252(4–5):261–72.
Schneider-Schaulies J, Schneider-Schaulies S. Sphingolipids in viral infection. Biol Chem. 2015;396(6–7):585–95.
Theken KN, Tang SY, Sengupta S, FitzGerald GA. The roles of lipids in SARS-CoV-2 viral replication and the host immune response. J Lipid Res. 2021;62: 100129.
Farley SE, Kyle JE, Leier HC, Bramer LM, Weinstein JB, Bates TA, et al. A global lipid map reveals host dependency factors conserved across SARS-CoV-2 variants. Nat Commun. 2022;13(1):3487.
Nardacci R, Colavita F, Castilletti C, Lapa D, Matusali G, Meschi S, et al. Evidences for lipid involvement in SARS-CoV-2 cytopathogenesis. Cell Death Dis. 2021;12(3):263.
Vitner EB, Avraham R, Politi B, Melamed S, Israely T. Elevation in sphingolipid upon SARS-CoV-2 infection: possible implications for COVID-19 pathology. Life Sci Alliance. 2022;5(1):e202101168.
Matsuyama S, Nao N, Shirato K, Kawase M, Saito S, Takayama I, et al. Enhanced isolation of SARS-CoV-2 by TMPRSS2-expressing cells. Proc Natl Acad Sci USA. 2020;117(13):7001–3.
Bedford T, Greninger AL, Roychoudhury P, Starita LM, Famulare M, Huang ML, et al. Cryptic transmission of SARS-CoV-2 in Washington state. Science (New York, NY). 2020;370(6516):571–5.
Balish AL, Katz JM, Klimov AI. Influenza: propagation, quantification, and storage. Curr Protoc Microbiol. 2013;29(1):15G.1.1-G.1.24.
Apffel A, Zhao L, Sartain MJ. A novel solid phase extraction sample preparation method for lipidomic analysis of human plasma using liquid chromatography/mass spectrometry. Metabolites. 2021;11(5):294.
Pang Z, Chong J, Zhou G, de Lima Morais DA, Chang L, Barrette M, et al. MetaboAnalyst 5.0: narrowing the gap between raw spectra and functional insights. Nucleic Acids Res. 2021;49(W1):W388-w96.
Koelmel JP, Li X, Stow SM, Sartain MJ, Murali A, Kemperman R, et al. Lipid annotator: towards accurate annotation in non-targeted liquid chromatography high-resolution tandem mass spectrometry (LC-HRMS/MS) lipidomics using a rapid and user-friendly software. Metabolites. 2020;10(3):1–20.
Zhang J, Pekosz A, Lamb RA. Influenza virus assembly and lipid raft microdomains: a role for the cytoplasmic tails of the spike glycoproteins. J Virol. 2000;74(10):4634–44.
Chukkapalli V, Heaton NS, Randall G. Lipids at the interface of virus-host interactions. Curr Opin Microbiol. 2012;15(4):512–8.
Falanga A, Cantisani M, Pedone C, Galdiero S. Membrane fusion and fission: enveloped viruses. Protein Pept Lett. 2009;16(7):751–9.
Saud Z, Tyrrell VJ, Zaragkoulias A, Protty MB, Statkute E, Rubina A, et al. The SARS-CoV2 envelope differs from host cells, exposes procoagulant lipids, and is disrupted in vivo by oral rinses. J Lipid Res. 2022;63(6): 100208.
Casari I, Manfredi M, Metharom P, Falasca M. Dissecting lipid metabolism alterations in SARS-CoV-2. Prog Lipid Res. 2021;82: 101092.
Zhao T, Wang C, Duan B, Yang P, Wu J, Zhang Q. Altered lipid profile in COVID-19 patients and metabolic reprogramming. Front Microbiol. 2022;13: 863802.
Yamate M, Yamashita M, Goto T, Tsuji S, Li YG, Warachit J, et al. Establishment of Vero E6 cell clones persistently infected with severe acute respiratory syndrome coronavirus. Microbes Infect. 2005;7(15):1530–40.
Reis A, Rudnitskaya A, Blackburn GJ, MohdFauzi N, Pitt AR, Spickett CM. A comparison of five lipid extraction solvent systems for lipidomic studies of human LDL. J Lipid Res. 2013;54(7):1812–24.
Sostare J, Di Guida R, Kirwan J, Chalal K, Palmer E, Dunn WB, et al. Comparison of modified Matyash method to conventional solvent systems for polar metabolite and lipid extractions. Anal Chim Acta. 2018;1037:301–15.
Höring M, Stieglmeier C, Schnabel K, Hallmark T, Ekroos K, Burkhardt R, et al. Benchmarking one-phase lipid extractions for plasma lipidomics. Anal Chem. 2022;94(36):12292–6.
Miller ME, Adhikary S, Kolokoltsov AA, Davey RA. Ebolavirus requires acid sphingomyelinase activity and plasma membrane sphingomyelin for infection. J Virol. 2012;86(14):7473–83.
Marchesini N, Hannun YA. Acid and neutral sphingomyelinases: roles and mechanisms of regulation. Biochem Cell Biol = Biochimie et biologie cellulaire. 2004;82(1):27–44.
Hannun YA. Functions of ceramide in coordinating cellular responses to stress. Science (New York, NY). 1996;274(5294):1855–9.
Stancevic B, Kolesnick R. Ceramide-rich platforms in transmembrane signaling. FEBS Lett. 2010;584(9):1728–40.
Hannun YA, Obeid LM. Many ceramides. J Biol Chem. 2011;286(32):27855–62.
Carpinteiro A, Gripp B, Hoffmann M, Pöhlmann S, Hoertel N, Edwards MJ, et al. Inhibition of acid sphingomyelinase by ambroxol prevents SARS-CoV-2 entry into epithelial cells. J Biol Chem. 2021;296: 100701.
Carpinteiro A, Edwards MJ, Hoffmann M, Kochs G, Gripp B, Weigang S, et al. Pharmacological inhibition of acid sphingomyelinase prevents uptake of SARS-CoV-2 by epithelial cells. Cell Rep Med. 2020;1(8): 100142.
Darquennes G, Le Corre P, Le Moine O, Loas G. Association between functional inhibitors of acid sphingomyelinase (FIASMAs) and reduced risk of death in COVID-19 patients: a retrospective cohort study. Pharmaceuticals (Basel, Switzerland). 2021;14(3).
Hoertel N, Sánchez-Rico M, Gulbins E, Kornhuber J, Carpinteiro A, Lenze EJ, et al. Association between FIASMAs and reduced risk of intubation or death in individuals hospitalized for severe COVID-19: an observational multicenter study. Clin Pharmacol Ther. 2021;110(6):1498–511.
Seftel D, Boulware DR. Prospective cohort of fluvoxamine for early treatment of coronavirus disease 19. Open Forum Infect Dis. 2021;8(2):ofab050.
Müller C, Hardt M, Schwudke D, Neuman BW, Pleschka S, Ziebuhr J. Inhibition of cytosolic phospholipase A(2)α impairs an early step of coronavirus replication in cell culture. J Virol. 2018;92(4).
Acknowledgements
The authors acknowledge the University of Maryland School of Pharmacy Faculty Start-up funds (J.W.J.); Agilent Research Gift #4520 (J.W.J.); University of Maryland, Baltimore, Institute for Clinical & Translational Research (ICTR) and the National Center for Advancing Translational Sciences (NCATS) Clinical Translational Science Award (CTSA) grant number 1UL1TR003098 (J.W.J.); University of Maryland School of Pharmacy Mass Spectrometry Center (SOP1841-IQB2014) and Cornell University NIH/NIAID grants R01 IA109022 and R21 AI142377 (H.A.C.); Emergent Ventures at Mercatus Center, George Mason Univ. Fast Grant # 2103 (H.A.C.); and Cornell Seed grants (H.A.C.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
ABC Highlights: authored by Rising Stars and Top Experts.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Abdel-Megied, A.M., Monreal, I.A., Zhao, L. et al. Characterization of the cellular lipid composition during SARS-CoV-2 infection. Anal Bioanal Chem 415, 5269–5279 (2023). https://doi.org/10.1007/s00216-023-04825-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-023-04825-1